Abstract
Autophagy protects cells from harmful substances such as protein aggregates, damaged mitochondria and intracellular pathogens, and has been implicated in a variety of diseases. Selectivity of autophagic processes is mediated by cargo receptors that link cargo to Atg8 family proteins on the developing autophagosomal membrane. To avoid collateral degradation during constitutive autophagic pathways, the autophagic machinery must not only select cargo but also exclude non-cargo material. Here we show that cargo directly activates the cargo receptor Atg19 by exposing multiple Atg8 binding sites. Furthermore, Atg19 mediates tight apposition of the cargo and Atg8-coated membranes in a fully reconstituted system. These properties are essential for the function of Atg19 during selective autophagy in vivo. Our results suggest that cargo receptors contribute to tight membrane bending of the isolation membrane around the cargo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Tooze, S. A. & Yoshimori, T. The origin of the autophagosomal membrane. Nat. Cell Biol. 12, 831–835 (2010).
Weidberg, H., Shvets, E. & Elazar, Z. Biogenesis and cargo selectivity of autophagosomes. Annu. Rev. Biochem. 80, 125–156 (2011).
Nakatogawa, H., Suzuki, K., Kamada, Y. & Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: Lessons from yeast. Nat. Rev. Mol. Cell Biol. 10, 458–467 (2009).
Orsi, A., Polson, H. E. J. & Tooze, S. A. Membrane trafficking events that partake in autophagy. Curr. Opin. Cell Biol. 22, 150–156 (2010).
Xie, Z. & Klionsky, D. J. Autophagosome formation: core machinery and adaptations. Nat. Cell Biol. 9, 1102–1109 (2007).
Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075 (2008).
Egan, D. F. et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331, 456–461 (2011).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42 (2008).
Youle, R. J. & van der Bliek, A. M. Mitochondrial fission, fusion, and stress. Science 337, 1062–1065 (2012).
Johansen, T. & Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 7, 279–296 (2011).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell 34, 259–269 (2009).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Noda, N. N. et al. Structural basis of target recognition by Atg8/LC3 during selective autophagy. Genes Cells 13, 1211–1218 (2008).
Noda, N. N., Ohsumi, Y. & Inagaki, F. Atg8-family interacting motif crucial for selective autophagy. FEBS Lett. 584, 1379–1385 (2010).
Ichimura, Y. et al. Structural basis for sorting mechanism of p62 in selective autophagy. J. Biol. Chem. 283, 22847–22857 (2008).
Kraft, C., Peter, M. & Hofmann, K. Selective autophagy: Ubiquitin-mediated recognition and beyond. Nat. Cell Biol. 12, 836–841 (2010).
Scott, S. V., Guan, J., Hutchins, M. U., Kim, J. & Klionsky, D. J. Cvt19 Is a receptor for the cytoplasm-to-vacuole targeting pathway. Mol. Cell 7, 1131–1141 (2001).
Leber, R., Silles, E., Sandoval, I. V. & Maz’on, M. a. J. Yol082p, a Novel CVT protein involved in the selective targeting of aminopeptidase I to the yeast vacuole. J. Biol. Chem. 276, 29210–29217 (2001).
Hutchins, M. U. & Klionsky, D. J. Vacuolar localisation of oligomeric α-mannosidase requires the cytoplasm to vacuole targeting and autophagy pathway components in Saccharomyces cerevisiae. J. Biol. Chem. 276, 20491–20498 (2001).
Yuga, M., Gomi, K., Klionsky, D. J. & Shintani, T. Aspartyl aminopeptidase is imported from the cytoplasm to the vacuole by selective autophagy in Saccharomyces cerevisiae. J. Biol. Chem. 286, 13704–13713 (2011).
Suzuki, K., Morimoto, M., Kondo, C. & Ohsumi, Y. Selective autophagy regulates insertional mutagenesis by the Ty1 retrotransposon in Saccharomyces cerevisiae. Dev. Cell 21, 358–365 (2011).
Baba, M., Osumi, M., Scott, S. V., Klionsky, D. J. & Ohsumi, Y. Two distinct pathways for targeting proteins from the cytoplasm to the vacuole/lysosome. J. Cell Biol. 139, 1687–1695 (1997).
Shintani, T., Huang, W. P., Stromhaug, P. E. & Klionsky, D. J. Mechanism of cargo selection in the cytoplasm to vacuole targeting pathway. Dev. Cell 3, 825–837 (2002).
Suzuki, K., Kondo, C., Morimoto, M. & Ohsumi, Y. Selective transport of α-mannosidase by autophagic pathways: identification of a novel receptor, Atg34p. J. Biol. Chem. 285, 30019–30025 (2010).
Rozenknop, A. et al. Characterization of the Interaction of GABARAPL-1 with the LIR Motif of NBR1. J. Mol. Biol. 410, 477–487 (2011).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Geng, J., Baba, M., Nair, U. & Klionsky, D. J. Quantitative analysis of autophagy-related protein stoichiometry by fluorescence microscopy. J. Cell Biol. 182, 129–140 (2008).
Romanov, J. et al. Mechanism and functions of membrane binding by the Atg5–Atg12/Atg16 complex during autophagosome formation. EMBO J. 31, 4304–4317 (2012).
Kamada, Y. et al. Tor-mediated induction of autophagy via an Apg1 protein kinase complex. J. Cell Biol. 150, 1507–1513 (2000).
Shintani, T. & Klionsky, D. J. Cargo proteins facilitate the formation of transport vesicles in the cytoplasm to vacuole targeting pathway. J. Biol. Chem. 279, 29889–29894 (2004).
Alemu, E. A. et al. ATG8 family proteins act as scaffolds for assembly of the ULK complex: Sequence requirements for LC3-interacting region (LIR) Motifs. J. Biol. Chem. 287, 39275–39290 (2012).
Von Muhlinen, N. et al. LC3C, bound selectively by a noncanonical LIR Motif in NDP52, Is required for antibacterial autophagy. Mol. Cell 48, 329–342 (2012).
Farre, J. C., Burkenroad, A., Burnett, S. F. & Subramani, S. Phosphorylation of mitophagy and pexophagy receptors coordinates their interaction with Atg8 and Atg11. EMBO Rep. 14, 441–449 (2013).
Suzuki, H. et al. Structural basis of the autophagy-related LC3/Atg13 LIR complex: Recognition and interaction mechanism. 22, 47–58 (2014).
Kaufmann, A., Beier, V., Franquelim, Henri G. & Wollert, T. Molecular mechanism of autophagic membrane-scaffold assembly and disassembly. Cell 156, 469–481 (2014).
Kirkin, V. et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol. Cell 33, 505–516 (2009).
Tokuyasu, K.T. A technique for ultracryotomy of cell suspensions and tissues. J. Cell Biol. 57, 551–565 (1973).
Acknowledgements
We thank C. Kraft and T. Brach for providing reagents and G. Warren for critical comments on the manuscript. Funding by the University of Vienna is gratefully acknowledged. The research leading to these results has received funding from the European Research Council under the European Community’s Seventh Framework Programme (FP7/2007-2013)/ERC grant agreement number 260304, from the FWF Austrian Science Fund (grant number P25546-B20) and by the EMBO Young Investigator Program to S.M. We also acknowledge funding by the VIPS Program of the Austrian Federal Ministry of Science and Research and the City of Vienna to J.S-M. and by the Austrian FFG to C.A.
Author information
Authors and Affiliations
Contributions
J.S-M., C.A, J.R., B.Z., I.I. and S.M. carried out experiments, J.S-M., C.A. and S.M. planned experiments and S.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Interaction of Atg19 with Atg8 is weak but stimulated by cargo and depends on multiple interaction sites.
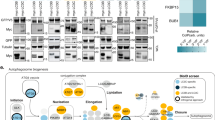
(a) Profile of an isothermal titration calorimetry experiment with Atg8 as ligand injected into the cell containing Atg19. The calculated KD is 35 μM. (b) Anti-Atg8 western blot of a pull down experiment with the indicated proteins. (c) Alignment of the C-termini of S. cerevisiae Atg19 and Atg34. The position of the Atg11 binding sites and the LIR motifs is indicated. (d) Coomassie stained gel showing the amounts of the indicated proteins used for the pull down experiment shown in Fig. 2d. (e) Coomassie stained gel showing the amounts of the indicated proteins used for the pull down experiment shown in Fig. 3a. (f) Coomassie stained gel showing the amounts of the indicated proteins used for the pull down experiment shown in Fig. 3b. (g) Anti-Atg8 western blot (upper part) using the indicated GST fusion proteins shown on the left as bait to pull down Atg8. The lower part shows the amount of Atg8 in the inputs on a Coomassie stained gel. Abbreviations: In: input, B: bound. The experiments shown in (a, b, g) have been conducted 2 times.
Supplementary Figure 2 The C-terminus of Atg19 interacts in a cargo-dependent manner with Atg8 via multiple sites.
(a, b) Anti-Atg8 western blots with the indicated GST fusion proteins as bait to pull down Atg8 are shown on the left. On the right the Coomassie-stained gels of the respective inputs are shown. (c) Coomassie stained gel showing the amounts of the indicated proteins used for the pull down experiment shown in Fig. 3f. Abbreviations: In: input, B: bound. The experiments shown in (a, b) have been conducted two times.
Supplementary Figure 3 Recruitment of Atg19 and Atg34 to Atg8 coated GUV and reconstitution of membrane bending during selective autophagy.
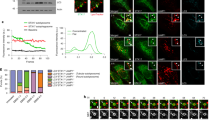
(a, b) GUVs containing 5% Nickel lipids were incubated with Atg8–His or GFP–His. Subsequently, different concentrations of mCherry–Atg19 or mCherry–Atg34 were added to the GUVs. For better visibility the mCherry signal is shown in pseudo-colour. Scale bars: 20 μm. (c) Atg19-dependent envelopment of prApe1-coated beads with Atg8 positive membranes Scale bars: 5M. The experimental setup for the experiment shown in (c) is shown in (d). Abbreviations: 3: Atg3; 5: Atg5; 7: Atg7; 8: Atg8; 12: Atg12; 16: Atg16; 19: Atg19. The experiments shown in (a, b) have been conducted two times; the experiment in (c) two times.
Supplementary Figure 4 Efficient pull down of Atg19 by GST–Atg8.
(a) Coomassie stained gel of a pull down experiment using GST–Atg8 as bait to pull down the indicated Atg19 proteins. (b) Coomassie stained gel of a pull down experiment using GST–Atg8 as bait to pull down the indicated Atg19 proteins. The quantification above the gel is based 3 independent experiments (N = 3), one of which is shown below the graph. Shown are the averages and standard deviations. The experiment shown in (a) has been conducted two times, the experiment in (b) three times.
Supplementary Figure 5 The Atg19–34 chimera localises to prApe1 oligomers.
(a, b) Representative electron micrographs of Ypt7, Atg19 yeast cells grown under Cvt conditions expressing the indicated proteins labelled with an anti-Myc primary antibody and a secondary anti-mouse antibody coupled to 10 nm gold particles. The boxes indicate the cropped regions shown in Fig. 6b. (c, d) Electron micrographs of Ypt7, Atg19 yeast cells grown under Cvt conditions expressing the indicated proteins co-labelled with a mouse anti-Myc antibody and a rabbit anti-prApe1 antiserum followed by incubation with secondary anti-mouse antibody coupled to 10 nm gold particles (black arrow) and anti-rabbit antibody coupled to 5 nm gold particles (white arrow). Scale bars: 200 nm. All experiments have conducted three times.
Supplementary Figure 6 Expression of Atg19 variants in yeast cells and function of Atg19 mutants during selective and bulk autophagy.
(a) Anti-Myc and anti-Pgk1 western blots of the same lysates shown in Fig. 6d. (b) Quantification of the percent of processed prApe1 compared to total prApe1. The graphs are based on three independent experiments (N = 3), one of which is shown in Fig. 6d. Shown are averages and standard deviations. (c) Anti-Ape1, anti-Myc and anti-Pgk1 western blots of atg17atg19 cells transformed with the indicated expression constructs. The lower mApe1 band indicates prApe1 processing and thus its delivery into the vacuole. The experiment in (a) has been conducted 2 times, the experiment in (c) two times.
Supplementary information
Supplementary Information
Supplementary Information (PDF 632 kb)
Supplementary Table 1
Supplementary Information (XLSX 41 kb)
Supplementary Table 2
Supplementary Information (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Sawa-Makarska, J., Abert, C., Romanov, J. et al. Cargo binding to Atg19 unmasks additional Atg8 binding sites to mediate membrane–cargo apposition during selective autophagy. Nat Cell Biol 16, 425–433 (2014). https://doi.org/10.1038/ncb2935
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2935
This article is cited by
-
Multifaceted membrane interactions of human Atg3 promote LC3-phosphatidylethanolamine conjugation during autophagy
Nature Communications (2023)
-
The caspase-6–p62 axis modulates p62 droplets based autophagy in a dominant-negative manner
Cell Death & Differentiation (2022)
-
Spatial control of avidity regulates initiation and progression of selective autophagy
Nature Communications (2021)
-
An N-terminal conserved region in human Atg3 couples membrane curvature sensitivity to conjugase activity during autophagy
Nature Communications (2021)
-
NMR resonance assignments of human Atg3 in aqueous solution and bicelles
Biomolecular NMR Assignments (2021)