Abstract
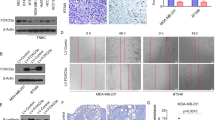
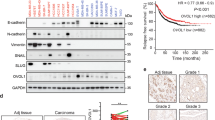
Despite advances in our understanding of breast cancer, patients with metastatic disease have poor prognoses. GATA3 is a transcription factor that specifies and maintains mammary luminal epithelial cell fate, and its expression is lost in breast cancer, correlating with a worse prognosis in human patients. Here, we show that GATA3 promotes differentiation, suppresses metastasis and alters the tumour microenvironment in breast cancer by inducing microRNA-29b (miR-29b) expression. Accordingly, miR-29b is enriched in luminal breast cancers and loss of miR-29b, even in GATA3-expressing cells, increases metastasis and promotes a mesenchymal phenotype. Mechanistically, miR-29b inhibits metastasis by targeting a network of pro-metastatic regulators involved in angiogenesis, collagen remodelling and proteolysis, including VEGFA, ANGPTL4, PDGF, LOX and MMP9, and targeting ITGA6, ITGB1 and TGFB, thereby indirectly affecting differentiation and epithelial plasticity. The discovery that a GATA3-miR-29b axis regulates the tumour microenvironment and inhibits metastasis opens up possibilities for therapeutic intervention in breast cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
ArrayExpress
Gene Expression Omnibus
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Valastyan, S. & Weinberg, R. A. Tumor metastasis: molecular insights and evolving paradigms. Cell 147, 275–292 (2011).
Hanahan, D. & Coussens, L. M. Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21, 309–322 (2012).
Lu, P., Weaver, V. M. & Werb, Z. The extracellular matrix: a dynamic niche in cancer progression. J. Cell Biol. 196, 395–406 (2012).
Yang, J. & Weinberg, R. A. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev. Cell 14, 818–829 (2008).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715 (2008).
Chou, J., Provot, S. & Werb, Z. GATA3 in development and cancer differentiation: cells GATA have it! J. Cell Physiol. 222, 42–49 (2010).
Asselin-Labat, M. L. et al. Gata-3 is an essential regulator of mammary-gland morphogenesis and luminal-cell differentiation. Nat. Cell Biol. 9, 201–209 (2007).
Kouros-Mehr, H., Slorach, E. M., Sternlicht, M. D. & Werb, Z. GATA-3 maintains the differentiation of the luminal cell fate in the mammary gland. Cell 127, 1041–1055 (2006).
Kouros-Mehr, H. et al. GATA-3 links tumor differentiation and dissemination in a luminal breast cancer model. Cancer Cell 13, 141–152 (2008).
Yoon, N. K. et al. Higher levels of GATA3 predict better survival in women with breast cancer. Hum. Pathol. 41, 1794–1801 (2010).
Mehra, R. et al. Identification of GATA3 as a breast cancer prognostic marker by global gene expression meta-analysis. Cancer Res. 65, 11259–11264 (2005).
Jacquemier, J. et al. Association of GATA3, P53, Ki67 status and vascular peritumoral invasion are strongly prognostic in luminal breast cancer. Breast Cancer Res. 11, R23 (2009).
Usary, J. et al. Mutation of GATA3 in human breast tumors. Oncogene 23, 7669–7678 (2004).
Network, C. G. A. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Asselin-Labat, M. L. et al. Gata-3 negatively regulates the tumor-initiating capacity of mammary luminal progenitor cells and targets the putative tumor suppressor caspase-14. Mol. Cell Biol. 31, 4609–4622 (2011).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Ma, L., Teruya-Feldstein, J. & Weinberg, R. A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449, 682–688 (2007).
Tavazoie, S. F. et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451, 147–152 (2008).
Valastyan, S. et al. A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell 137, 1032–1046 (2009).
Dore, L. C. et al. A GATA-1-regulated microRNA locus essential for erythropoiesis. Proc. Natl Acad. Sci. USA 105, 3333–3338 (2008).
Ibarra, I., Erlich, Y., Muthuswamy, S. K., Sachidanandam, R. & Hannon, G. J. A role for microRNAs in maintenance of mouse mammary epithelial progenitor cells. Genes Dev. 21, 3238–3243 (2007).
Yu, F. et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131, 1109–1123 (2007).
Sorlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001).
Foulkes, W. D., Smith, I. E. & Reis-Filho, J. S. Triple-negative breast cancer. New Engl. J. Med. 363, 1938–1948 (2010).
Neve, R. M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006).
Debnath, J. & Brugge, J. S. Modelling glandular epithelial cancers in three-dimensional cultures. Nat. Rev. Cancer 5, 675–688 (2005).
Lee, G. Y., Kenny, P. A., Lee, E. H. & Bissell, M. J. Three-dimensional culturemodels of normal and malignant breast epithelial cells. Nat. Methods 4, 359–365 (2007).
Visvader, J. E. Keeping abreast of the mammary epithelial hierarchy and breast tumorigenesis. Genes Dev. 23, 2563–2577 (2009).
Mott, J. L. et al. Transcriptional suppression of mir-29b-1/mir-29a promoter by c-Myc, hedgehog, and NF-κB. J. Cell Biochem. 110, 1155–1164 (2010).
Winbanks, C. E. et al. TGF-β regulates miR-206 and miR-29 to control myogenic differentiation through regulation of HDAC4. J. Biol. Chem. 286, 13805–13814 (2011).
Wang, H. et al. NF-κB-YY1-miR-29 regulatory circuitry in skeletal myogenesis and rhabdomyosarcoma. Cancer Cell 14, 369–381 (2008).
Blenkiron, C. et al. MicroRNA expression profiling of human breast cancer identifies new markers of tumor subtype. Genome Biol. 8, R214 (2007).
Buffa, F. M. et al. microRNA-associated progression pathways and potential therapeutic targets identified by integrated mRNA and microRNA expression profiling in breast cancer. Cancer Res. 71, 5635–5645 (2011).
Enerly, E. et al. miRNA-mRNA integrated analysis reveals roles for miRNAs in primary breast tumors. PLoS One 6, e16915 (2011).
Zhu, M. et al. Integrated miRNA and mRNA expression profiling of mouse mammary tumor models identifies miRNA signatures associated with mammary tumor lineage. Genome Biol. 12, R77 (2011).
Aslakson, C. J. & Miller, F. R. Selective events in the metastatic process defined by analysis of the sequential dissemination of subpopulations of a mouse mammary tumor. Cancer Res. 52, 1399–1405 (1992).
Dykxhoorn, D. M. et al. miR-200 enhances mouse breast cancer cell colonization to form distant metastases. PLoS One 4, e7181 (2009).
Lin, E. Y. et al. Progression to malignancy in the polyoma middle T oncoprotein mouse breast cancer model provides a reliable model for human diseases. Am. J. Pathol. 163, 2113–2126 (2003).
Ewald, A. J., Brenot, A., Duong, M., Chan, B. S. & Werb, Z. Collective epithelial migration and cell rearrangements drive mammary branching morphogenesis. Dev. Cell 14, 570–581 (2008).
Lewis, B. P., Burge, C. B. & Bartel, D. P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Betel, D., Wilson, M., Gabow, A., Marks, D. S. & Sander, C. The microRNA.org resource: targets and expression. Nucl. Acids Res. 36, D149–D153 (2008).
Krek, A. et al. Combinatorial microRNA target predictions. Nat. Genet. 37, 495–500 (2005).
Yu, K. R. et al. CD49f enhances multipotency and maintains stemness through the direct regulation of OCT4 and SOX2. Stem Cells 30, 876–887 (2012).
Padua, D. et al. TGFβ primes breast tumors for lung metastasis seeding through angiopoietin-like 4. Cell 133, 66–77 (2008).
Yan, W., Cao, Q. J., Arenas, R. B., Bentley, B. & Shao, R. GATA3 inhibits breast cancer metastasis through the reversal of epithelial-mesenchymal transition. J. Biol. Chem. 285, 14042–14051 (2010).
Chu, I. M. et al. GATA3 inhibits lysyl oxidase-mediated metastases of human basal triple-negative breast cancer cells. Oncogene 31, 2017–2027 (2012).
Levental, K. R. et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139, 891–906 (2009).
Du, R. et al. HIF1α induces the recruitment of bone marrow-derived vascular modulatory cells to regulate tumor angiogenesis and invasion. Cancer Cell 13, 206–220 (2008).
van Rooij, E. et al. Dysregulation of microRNAs after myocardial infarction reveals a role of miR-29 in cardiac fibrosis. Proc. Natl Acad. Sci. USA 105, 13027–13032 (2008).
Roderburg, C. et al. Micro-RNA profiling reveals a role for miR-29 in human and murine liver fibrosis. Hepatology 53, 209–218 (2010).
Wang, B. et al. Suppression of microRNA-29 expression by TGF- β1 promotes collagen expression and renal fibrosis. J. Am. Soc. Nephrol. 2, 252–265 (2011).
Fabbri, M. et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl Acad. Sci. USA 104, 15805–15810 (2007).
Nguyen, T. et al. Downregulation of microRNA-29c is associated with hypermethylation of tumor-related genes and disease outcome in cutaneous melanoma. Epigenetics 6, 388–394 (2010).
Xiong, Y. et al. Effects of microRNA-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology 51, 836–845 (2009).
Sethi, A., Mao, W., Wordinger, R. J. & Clark, A. F. Transforming growth factor- β induces extracellular matrix protein cross-linking lysyl oxidase (LOX) genes in human trabecular meshwork cells. Invest. Ophthalmol. Vis. Sci. 52, 5240–5250 (2011).
Dumont, N. & Arteaga, C. L. Transforming growth factor- β and breast cancer: tumor promoting effects of transforming growth factor- β. Breast Cancer Res. 2, 125–132 (2000).
Sanchez-Elsner, T. et al. Synergistic cooperation between hypoxia and transforming growth factor- β pathways on human vascular endothelial growth factor gene expression. J. Biol. Chem. 276, 38527–38535 (2001).
Hurst, D. R., Edmonds, M. D. & Welch, D. R. Metastamir: the field of metastasis-regulatory microRNA is spreading. Cancer Res. 69, 7495–7498 (2009).
Lujambio, A. & Lowe, S. W. The microcosmos of cancer. Nature 482, 347–355 (2012).
Ma, L. et al. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat. Biotechnol. 28, 341–347 (2010).
Kota, J. et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137, 1005–1017 (2009).
Elenbaas, B. et al. Human breast cancer cells generated by oncogenic transformation of primary mammary epithelial cells. Genes Dev. 15, 50–65 (2001).
Hwang, H. W., Wentzel, E. A. & Mendell, J. T. A hexanucleotide element directs microRNA nuclear import. Science 315, 97–100 (2007).
Tong, Q. et al. Function of GATA transcription factors in preadipocyte-adipocyte transition. Science 290, 134–138 (2000).
Wrana, J. L. et al. TGF β signals through a heteromeric protein kinase receptor complex. Cell 71, 1003–1014 (1992).
Zawel, L. et al. Human Smad3 and Smad4 are sequence-specific transcription activators. Mol. Cell 1, 611–617 (1998).
Watnick, R. S., Cheng, Y. N., Rangarajan, A., Ince, T. A. & Weinberg, R. A. Ras modulates Myc activity to repress thrombospondin-1 expression and increase tumor angiogenesis. Cancer Cell 3, 219–231 (2003).
Gough, P. J., Gomez, I. G., Wille, P. T. & Raines, E. W. Macrophage expression of active MMP-9 induces acute plaque disruption in apoE-deficient mice. J. Clin. Invest. 116, 59–69 (2006).
Stingl, J. et al. Purification and unique properties of mammary epithelial stem cells. Nature 439, 993–997 (2006).
Acknowledgements
We thank members of the Werb laboratory for discussions, P. Shahi, J. Dai and J. Tai for experimental assistance and E. Atamaniuc, Y. Yu, and H. Capili for technical assistance. We thank T. Rambaldo and M. Kissner for flow cytometer assistance, the UCSF Biological Imaging Development Center for microscopy assistance, J. Debnath, G. Bergers, and D. Sheppard for discussions, and A. Goga, V. Weaver, J. Mott, P. Gonzalez, K. Xie and E. Raines for reagents. We also thank C. Choi for discussion and support. This research was supported by funds from the National Cancer Institute (R01 CA129523 to Z.W.), a Developmental Research grant from the Bay Area Breast Cancer SPORE (P50 CA058207 to Z.W.), a Department of Defense Predoctoral Fellowship (W81XWH-10-1-0168 to J.C.) and the UCSF Medical Scientist Training Program (J.C.). We dedicate this work to the memory of L. Verber.
Author information
Authors and Affiliations
Contributions
J.H.L. and A.B. contributed equally to this work. J.C. designed and performed experiments, with assistance from J.H.L., A.B., J-w.K. and S.P. Z.W. designed experiments and supervised research. J.C. and Z.W. wrote the manuscript, and all authors discussed the results and provided comments and feedback.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1770 kb)
Supplementary Tables 1–5
Supplementary Information (XLS 69 kb)
Time-lapse imaging of MDA231-Control breast cancer cells cultured in 3D Matrigel.
MDA231-Control cells were embedded into growth factor reduced Matrigel after re-aggregation overnight on low adhesion plates. 3D cultures were grown in serum-free media with 2.5 nM FGF2. Cells were imaged using a Zeiss Axiovert inverted brightfield microscope every 20 min for 48 h at 37 °C and 5% CO2. Images were assembled and played at 8 frames s−1. (MOV 824 kb)
Time-lapse imaging of MDA231-GATA3 breast cancer cells cultured in 3D Matrigel.
MDA231-GATA3 cells were embedded into growth factor reduced Matrigel after re-aggregation overnight on low adhesion plates. 3D cultures were grown in serum-free media with 2.5 nM FGF2. Cells were imaged using a Zeiss Axiovert inverted brightfield microscope every 20 min for 48 h at 37 °C and 5% CO2. Images were assembled and played at 8 frames s−1. (MOV 1018 kb)
Rights and permissions
About this article
Cite this article
Chou, J., Lin, J., Brenot, A. et al. GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression. Nat Cell Biol 15, 201–213 (2013). https://doi.org/10.1038/ncb2672
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2672
This article is cited by
-
Rutaecarpine induces the differentiation of triple-negative breast cancer cells through inhibiting fumarate hydratase
Journal of Translational Medicine (2023)
-
Loss of function of GATA3 regulates FRA1 and c-FOS to activate EMT and promote mammary tumorigenesis and metastasis
Cell Death & Disease (2023)
-
N6-methyladenosine RNA methylation in liver diseases: from mechanism to treatment
Journal of Gastroenterology (2023)
-
An N-glycoproteomic site-mapping analysis reveals glycoprotein alterations in esophageal squamous cell carcinoma
Journal of Translational Medicine (2022)
-
MiR-29b may suppresses peritoneal metastases through inhibition of the mesothelial–mesenchymal transition (MMT) of human peritoneal mesothelial cells
Scientific Reports (2022)