Abstract
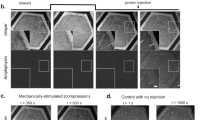
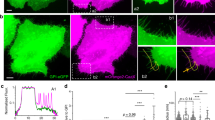
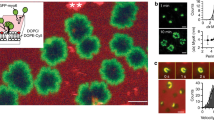
Many of the more than 20 mammalian proteins with N-BAR domains1,2 control cell architecture3 and endocytosis4,5 by associating with curved sections of the plasma membrane6. It is not well understood whether N-BAR proteins are recruited directly by processes that mechanically curve the plasma membrane or indirectly by plasma-membrane-associated adaptor proteins that recruit proteins with N-BAR domains that then induce membrane curvature. Here, we show that externally induced inward deformation of the plasma membrane by cone-shaped nanostructures (nanocones) and internally induced inward deformation by contracting actin cables both trigger recruitment of isolated N-BAR domains to the curved plasma membrane. Markedly, live-cell imaging in adherent cells showed selective recruitment of full-length N-BAR proteins and isolated N-BAR domains to plasma membrane sub-regions above nanocone stripes. Electron microscopy confirmed that N-BAR domains are recruited to local membrane sites curved by nanocones. We further showed that N-BAR domains are periodically recruited to curved plasma membrane sites during local lamellipodia retraction in the front of migrating cells. Recruitment required myosin-II-generated force applied to plasma-membrane-connected actin cables. Together, our results show that N-BAR domains can be directly recruited to the plasma membrane by external push or internal pull forces that locally curve the plasma membrane.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Habermann, B. The BAR-domain family of proteins: a case of bending and binding? EMBO Rep. 5, 250–255 (2004).
Suetsugu, S., Toyooka, K. & Senju, Y. Subcellular membrane curvature mediated by the BAR domain superfamily proteins. Semin. Cell Dev. Biol. 21, 340–349 (2010).
Rollason, R., Korolchuk, V., Hamilton, C., Jepson, M. & Banting, G. A CD317/tetherin-RICH2 complex plays a critical role in the organization of the subapical actin cytoskeleton in polarized epithelial cells. J. Cell Biol. 184, 721–736 (2009).
David, C., McPherson, P. S., Mundigl, O. & de Camilli, P. A role of amphiphysin in synaptic vesicle endocytosis suggested by its binding to dynamin in nerve terminals. Proc. Natl Acad. Sci. USA 93, 331–335 (1996).
Ringstad, N. et al. Endophilin/SH3p4 is required for the transition from early to late stages in clathrin-mediated synaptic vesicle endocytosis. Neuron 24, 143–154 (1999).
Bhatia, V. K. et al. Amphipathic motifs in BAR domains are essential for membrane curvature sensing. EMBO J. 28, 3303–3314 (2009).
Peter, B. J. et al. BAR domains as sensors of membrane curvature: the amphiphysin BAR structure. Science 303, 495–499 (2004).
Taylor, M. J., Perrais, D. & Merrifield, C. J. A high precision survey of the molecular dynamics of mammalian clathrin-mediated endocytosis. PLoS Biol. 9, e1000604 (2011).
Yamada, H. et al. Dynamic interaction of amphiphysin with N-WASP regulates actin assembly. J. Biol. Chem. 284, 34244–34256 (2009).
Yarar, D., Waterman-Storer, C. M. & Schmid, S. L. SNX9 couples actin assembly to phosphoinositide signals and is required for membrane remodeling during endocytosis. Dev. Cell 13, 43–56 (2007).
Rocca, D. L., Martin, S., Jenkins, E. L. & Hanley, J. G. Inhibition of Arp2/3-mediated actin polymerization by PICK1 regulates neuronal morphology and AMPA receptor endocytosis. Nat. Cell Biol. 10, 259–271 (2008).
Salazar, M. A. et al. Tuba, a novel protein containing bin/amphiphysin/Rvs and Dbl homology domains, links dynamin to regulation of the actin cytoskeleton. J. Biol. Chem. 278, 49031–49043 (2003).
Richnau, N. & Aspenstrom, P. Rich, a ρ GTPase-activating protein domain-containing protein involved in signaling by Cdc42 and RAC1. J. Biol. Chem. 276, 35060–35070 (2001).
Jeong, S., McDowell, M. T. & Cui, Y. Low-temperature self-catalytic growth of tin oxide nanocones over large areas. ACS Nano 5, 5800–5807 (2011).
Giannone, G. et al. Periodic lamellipodial contractions correlate with rearward actin waves. Cell 116, 431–443 (2004).
Gallop, J. L. et al. Mechanism of endophilin N-BAR domain-mediated membrane curvature. EMBO J. 25, 2898–2910 (2006).
Masuda, M. et al. Endophilin BAR domain drives membrane curvature by two newly identified structure-based mechanisms. EMBO J. 25, 2889–2897 (2006).
Ferguson, S. M. et al. Coordinated actions of actin and BAR proteins upstream of dynamin at endocytic clathrin-coated pits. Dev. Cell 17, 811–822 (2009).
Inoue, T. & Meyer, T. Synthetic activation of endogenous PI3K and Racidentifies an AND-gate switch for cell polarization and migration. PLoS One 3, e3068 (2008).
Wu, Y. I. et al. A genetically encoded photoactivatable Rac controls the motility of living cells. Nature 461, 104–108 (2009).
Burnette, D. T. et al. A role for actin arcs in the leading-edge advance of migrating cells. Nat. Cell Biol. 13, 371–381 (2011).
Machacek, M. et al. Coordination of ρ GTPase activities during cell protrusion. Nature 461, 99–103 (2009).
Giannone, G. et al. Lamellipodial actin mechanically links myosin activity with adhesion-site formation. Cell 128, 561–575 (2007).
Abercrombie, M., Heaysman, J. E. & Pegrum, S. M. The locomotion of fibroblasts in culture. IV. Electron microscopy of the leading lamella. Exp. Cell Res. 67, 359–367 (1971).
Raucher, D. & Sheetz, M. P. Cell spreading and lamellipodial extension rate is regulated by membrane tension. J. Cell Biol. 148, 127–136 (2000).
Shimada, A. et al. Curved EFC/F-BAR-domain dimers are joined end to end into a filament for membrane invagination in endocytosis. Cell 129, 761–772 (2007).
Henne, W. M. et al. Structure and analysis of FCHo2 F-BAR domain: a dimerizing and membrane recruitment module that effects membrane curvature. Structure 15, 839–852 (2007).
Millard, T. H. et al. Structural basis of filopodia formation induced by the IRSp53/MIM homology domain of human IRSp53. EMBO J. 24, 240–250 (2005).
Johnson, H. W. & Schell, M. J. Neuronal IP3 3-kinase is an F-actin-bundling protein: role in dendritic targeting and regulation of spine morphology. Mol. Biol. Cell 20, 5166–5180 (2009).
Inoue, T., Heo, W. D., Grimley, J. S., Wandless, T. J. & Meyer, T. An inducible translocation strategy to rapidly activate and inhibit small GTPase signaling pathways. Nat. Methods 2, 415–418 (2005).
Wei, Q. & Adelstein, R. S. Conditional expression of a truncated fragment of nonmuscle myosin II-A alters cell shape but not cytokinesis in HeLa cells. Mol. Biol. Cell 11, 3617–3627 (2000).
Tsai, F. C. & Meyer, T. Ca(2+) pulses control local cycles of lamellipodia retraction and adhesion along the front of migrating cells. Curr. Biol. 22, 837–842 (2012).
Acknowledgements
The authors thank the members of the Meyer laboratory for comments and discussion. M.G. was supported by the Swiss National Science Foundation (No. PBBSP3-123159), and a Novartis Jubilaeumsstiftung and Stanford Deans Postdoctoral Fellowship. S.J. was supported by a Korea Foundation for Advanced Studies graduate fellowship. Y.C. acknowledges the partial support from a DOE-EFRC at Stanford: Center on Nanostructuring for Efficient Energy Conversion (No. DE-SC0001060). T.M. acknowledges financial support from the National Institutes of Health, MH064801, MH095087 and GM063702.
Author information
Authors and Affiliations
Contributions
M.G. performed all experiments and analysed the data. F-C.T. developed the temporal cross-correlation analysis. S.J. and Y.C. designed the nanocones. L-M.J. helped with the SEM. Y.I.W. and K.M.H. developed the photoactivatable RAC construct. M.G. and T.M. designed the experiments, interpreted the results and wrote the manuscript. All authors discussed the results and the manuscript. T.M. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1345 kb)
Supplementary Movie 1
Supplementary Information (AVI 28875 kb)
Supplementary Movie 2
Supplementary Information (AVI 845 kb)
Supplementary Movie 3
Supplementary Information (AVI 120 kb)
Rights and permissions
About this article
Cite this article
Galic, M., Jeong, S., Tsai, FC. et al. External push and internal pull forces recruit curvature-sensing N-BAR domain proteins to the plasma membrane. Nat Cell Biol 14, 874–881 (2012). https://doi.org/10.1038/ncb2533
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2533
This article is cited by
-
Biointerface design for vertical nanoprobes
Nature Reviews Materials (2022)
-
Enantiomer-dependent immunological response to chiral nanoparticles
Nature (2022)
-
Curvature dependence of BAR protein membrane association and dissociation kinetics
Scientific Reports (2022)
-
TGFβ-induced changes in membrane curvature influence Ras oncoprotein membrane localization
Scientific Reports (2022)
-
Insights into Membrane Curvature Sensing and Membrane Remodeling by Intrinsically Disordered Proteins and Protein Regions
The Journal of Membrane Biology (2022)