Abstract
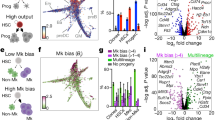
How the molecular programs of differentiated cells develop as cells transit from multipotency through lineage commitment remains unexplored. This reflects the inability to access cells undergoing commitment or located in the immediate vicinity of commitment boundaries. It remains unclear whether commitment constitutes a gradual process, or else represents a discrete transition. Analyses of in vitro self-renewing multipotent systems have revealed cellular heterogeneity with individual cells transiently exhibiting distinct biases for lineage commitment1,2,3,4,5,6. Such systems can be used to molecularly interrogate early stages of lineage affiliation and infer rules of lineage commitment. In haematopoiesis, population-based studies have indicated that lineage choice is governed by global transcriptional noise, with self-renewing multipotent cells reversibly activating transcriptome-wide lineage-affiliated programs7. We examine this hypothesis through functional and molecular analysis of individual blood cells captured from self-renewal cultures, during cytokine-driven differentiation and from primary stem and progenitor bone marrow compartments. We show dissociation between self-renewal potential and transcriptome-wide activation of lineage programs, and instead suggest that multipotent cells experience independent activation of individual regulators resulting in a low probability of transition to the committed state.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Enver, T., Pera, M., Peterson, C. & Andrews, P. W. Stem cell states, fates, and the rules of attraction. Cell Stem. Cell 4, 387–397 (2009).
Canham, M. A., Sharov, A. A., Ko, M. S. H. & Brickman, J. M. Functional heterogeneity of embryonic stem cells revealed through translational amplification of an early endodermal transcript. PLoS Biol. 8, e1000379 (2010).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Hayashi, K., Lopes, S. M., Tang, F. & Surani, M. A. Dynamic equilibrium and heterogeneity of mouse pluripotent stem cells with distinct functional and epigenetic states. Cell Stem. Cell 3, 391–401 (2008).
Hough, S. R., Laslett, A. L., Grimmond, S. B., Kolle, G. & Pera, M. F. A continuum of cell states spans pluripotency and lineage commitment in human embryonic stem cells. PLoS One 4, e7708 (2009).
Kalmar, T. et al. Regulated fluctuations in nanog expression mediate cell fate decisions in embryonic stem cells. PLoS Biol. 7, e1000149 (2009).
Chang, H. H., Hemberg, M., Barahona, M., Ingber, D. E. & Huang, S. Transcriptome-wide noise controls lineage choice in mammalian progenitor cells. Nature 453, 544–547 (2008).
Bussmann, L. H. et al. A robust and highly efficient immune cell reprogramming system. Cell Stem. Cell 5, 554–566 (2009).
Davis, R. L., Weintraub, H. & Lassar, A. B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51, 987–1000 (1987).
Feng, R. et al. PU.1 and C/EBP α/β convert fibroblasts into macrophage-like cells. Proc. Natl Acad. Sci. USA 105, 6057–6062 (2008).
Graf, T. & Enver, T. Forcing cells to change lineages. Nature 462, 587–594 (2009).
Heyworth, C., Pearson, S., May, G. & Enver, T. Transcription factor-mediated lineage switching reveals plasticity in primary committed progenitor cells. EMBO J. 21, 3770–3781 (2002).
Iwasaki, H. et al. GATA-1 converts lymphoid and myelomonocytic progenitors into the megakaryocyte/erythrocyte lineages. Immunity 19, 451–462 (2003).
Weintraub, H. et al. Activation of muscle-specific genes in pigment, nerve, fat, liver, and fibroblast cell lines by forced expression of MyoD. Proc. Natl Acad. Sci. USA 86, 5434–5438 (1989).
Tsai, S., Bartelmez, S., Sitnicka, E. & Collins, S. Lymphohematopoietic progenitors immortalized by a retroviral vector harboring a dominant-negative retinoic acid receptor can recapitulate lymphoid, myeloid, and erythroid development. Genes Dev. 8, 2831–2841 (1994).
Ye, Z. J., Kluger, Y., Lian, Z. & Weissman, S. M. Two types of precursor cells in a multipotential hematopoietic cell line. Proc. Natl Acad. Sci. USA 102, 18461–18466 (2005).
Kim, S. I. & Bresnick, E. H. Transcriptional control of erythropoiesis: emerging mechanisms and principles. Oncogene 26, 6777–6794 (2007).
Capron, C. et al. LYL-1 deficiency induces a stress erythropoiesis. Exp. Hematol. 39, 629–642 (2011).
Pronk, C. J. et al. Elucidation of the phenotypic, functional, and molecular topography of a myeloerythroid progenitor cell hierarchy. Cell Stem. Cell 1, 428–442 (2007).
Pina, C., May, G., Soneji, S., Hong, D. & Enver, T. MLLT3 regulates early human erythroid and megakaryocytic cell fate. Cell Stem. Cell 2, 264–273 (2008).
Rodrigues, N. P. et al. Haploinsufficiency of GATA-2 perturbs adult hematopoietic stem-cell homeostasis. Blood 106, 477–484 (2005).
Tipping, A. J. et al. High GATA-2 expression inhibits human hematopoietic stem and progenitor cell function by effects on cell cycle. Blood 113, 2661–2672 (2009).
Porcher, C. et al. The T cell leukemia oncoprotein SCL/tal-1 is essential for development of all hematopoietic lineages. Cell 86, 47–57 (1996).
Wray, J., Kalkan, T. & Smith, A. G. The ground state of pluripotency. Biochem. Soc. Trans. 38, 1027–1032 (2010).
Coulombel, L. Identification of hematopoietic stem/progenitor cells: strength and drawbacks of functional assays. Oncogene 23, 7210–7222 (2004).
Adolfsson, J. et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential: a revised road map for adult blood lineage commitment. Cell 121, 295–306 (2005).
Huang, S., Eichler, G., Bar-Yam, Y. & Ingber, D. E. Cell fates as high-dimensional attractor states of a complex gene regulatory network. Phys. Rev. Lett. 94, 128701 (2005).
Orford, K. et al. Differential H3K4 methylation identifies developmentally poised hematopoietic genes. Dev. Cell 14, 798–809 (2008).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic. Acids Res. 29, e45 (2001).
Hu, M. et al. Multilineage gene expression precedes commitment in the hemopoietic system. Genes Dev. 11, 774–785 (1997).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet Mol. Biol. 3, 1–25 (2004).
Acknowledgements
The authors would like to thank R. Gupta and G. May for conceptual discussions; C. Waugh, K. Clark, A. Pizzey and T. Adejumo for cell sorting; S. McGowan for microarray data accessibility; and J. Wray for critical reading of the manuscript. J.T. is a student of the PhD Program in Computational Biology at Instituto Gulbenkian de Ciencia, Oeiras, Portugal, and was financially supported by Fundacao para a Ciencia e Tecnologia (SFRH/BD/33208/2007). C. Peterson is supported by the Swedish Foundation for Strategic Research (Senior Individual Grant). This work was financially supported by the Medical Research Council of the United Kingdom, Leukaemia and Lymphoma Research, EuroSyStem and STEMEXPAND.
Author information
Authors and Affiliations
Contributions
C. Pina initiated and led the study, conducted all single-cell RT–qPCR experiments, carried out clonal reconstitution assays and divisional tracking experiments, participated in population-based analyses of EML subcompartments, analysed and interpreted experimental data, participated in figure production and wrote the paper. C.F. processed mouse bone marrow samples, carried out western blotting, participated in characterization of EML subcompartments, including clonal reconstitution assays, analysed experimental data and contributed to its interpretation and produced the figures. A.J.T. carried out immunostaining, did cell sorting, participated in population-based analyses of EML subcompartments, analysed experimental data and contributed to its interpretation and contributed to figure production. J.B. carried out non-quantitative single-cell RT–PCR and processed microarray samples. S.S. analysed microarray data, contributed to analysis of single-cell RT–qPCR data and participated in figure production. J.T. and C. Peterson contributed to analysis of single-cell RT–qPCR data and contributed to data interpretation. T.E. supervised all aspects of the study, and wrote the paper. C. Pina, C.F. and A.J.T. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 391 kb)
Supplementary Table 1
Supplementary Information (XLS 31 kb)
Supplementary Table 2
Supplementary Information (XLSX 283 kb)
Supplementary Table 3
Supplementary Information (XLS 56 kb)
Supplementary Table 4
Supplementary Information (XLS 127 kb)
Rights and permissions
About this article
Cite this article
Pina, C., Fugazza, C., Tipping, A. et al. Inferring rules of lineage commitment in haematopoiesis. Nat Cell Biol 14, 287–294 (2012). https://doi.org/10.1038/ncb2442
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2442
This article is cited by
-
Critical transition and reversion of tumorigenesis
Experimental & Molecular Medicine (2023)
-
Intravital microscopy imaging of kidney injury and regeneration
Renal Replacement Therapy (2021)
-
Executable cancer models: successes and challenges
Nature Reviews Cancer (2020)
-
SCNS: a graphical tool for reconstructing executable regulatory networks from single-cell genomic data
BMC Systems Biology (2018)
-
Nanog induced intermediate state in regulating stem cell differentiation and reprogramming
BMC Systems Biology (2018)