Abstract
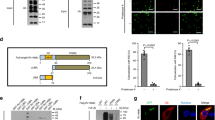
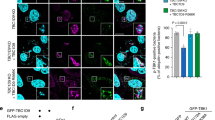
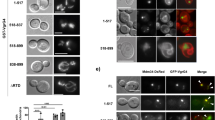
Autophagy degrades unnecessary organelles and misfolded protein aggregates1, as well as cytoplasm-invading bacteria2. Nevertheless, the bacteria Listeria monocytogenes efficiently escapes autophagy3,4. We show here that recruitment of the Arp2/3 complex and Ena/VASP, via the bacterial ActA protein, to the bacterial surface disguises the bacteria from autophagic recognition, an activity that is independent of the ability to mediate bacterial motility. L. monocytogenes expressing ActA mutants that lack the ability to recruit the host proteins initially underwent ubiquitylation, followed by recruitment of p62 (also known as SQSTM1) and LC3, before finally undergoing autophagy. The ability of ActA to mediate protection from ubiquitylation was further demonstrated by generating aggregate-prone GFP–ActA–Q79C and GFP–ActA–170* chimaeras, consisting of GFP (green fluorescent protein), the ActA protein and segments of polyQ5 or Golgi membrane protein GCP170 (ref. 6). GFP–ActA–Q79C and GFP–ActA–170* formed aggregates in the host cell cytoplasm, however, these ActA-containing aggregates were not targeted for association with ubiquitin and p62. Our findings indicate that ActA-mediated host protein recruitment is a unique bacterial disguise tactic to escape from autophagy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Williams, A. et al. Aggregate-prone proteins are cleared from the cytosol by autophagy: therapeutic implications. Curr. Top. Dev. Biol. 76, 89–101 (2006).
Deretic, V. & Levine, B. Autophagy, immunity, and microbial adaptations. Cell Host Microbe. 5, 527–549 (2009).
Py, B. F., Lipinski, M. M. & Yuan, J. Autophagy limits Listeria monocytogenes intracellular growth in the early phase of primary infection. Autophagy 3, 117–125 (2007).
Birmingham, C. L. et al. Listeria monocytogenes evades killing by autophagy during colonization of host cells. Autophagy 3, 442–451 (2007).
Ikeda, H. et al. Expanded polyglutamine in the Machado-Joseph disease protein induces cell death in vitro and in vivo. Nature Genet. 13, 196–202 (1996).
Fu, L. et al. Nuclear aggresomes form by fusion of PML-associated aggregates. Mol. Biol. Cell 16, 4905–4917 (2005).
Klionsky, D. J. & Emr, S. D. Autophagy as a regulated pathway of cellular degradation. Science 290, 1717–1721 (2000).
Nakagawa, I. et al. Autophagy defends cells against invading group A Streptococcus. Science 306, 1037–1040 (2004).
Gutierrez, M. G. et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119, 753–766 (2004).
Deretic, V. et al. Mycobacterium tuberculosis inhibition of phagolysosome biogenesis and autophagy as a host defence mechanism. Cell. Microbiol. 8, 719–727 (2006).
Otto, G. P. et al. Macroautophagy is dispensable for intracellular replication of Legionella pneumophila in Dictyostelium discoideum. Mol. Microbiol. 51, 63–72 (2004).
Gutierrez, M. G. et al. Autophagy induction favours the generation and maturation of the Coxiella-replicative vacuoles. Cell. Microbiol. 7, 981–993 (2005).
Dorn, B. R., Dunn, W. A., Jr. & Progulske-Fox, A. Porphyromonas gingivalis traffics to autophagosomes in human coronary artery endothelial cells. Infect. Immun. 69, 5698–5708 (2001).
Ogawa, M. et al. Escape of intracellular Shigella from autophagy. Science 307, 727–731 (2005).
Kocks, C. et al. L. monocytogenes-induced actin assembly requires the actA gene product, a surface protein. Cell 68, 521–531 (1992).
Stevens, J. M., Galyov, E. E. & Stevens, M. P. Actin-dependent movement of bacterial pathogens. Nature Rev. Microbiol. 4, 91–101 (2006).
Rich, K. A., Burkett, C. & Webster, P. Cytoplasmic bacteria can be targets for autophagy. Cell. Microbiol. 5, 455–468 (2003).
Birmingham, C. L. et al. Listeriolysin O allows Listeria monocytogenes replication in macrophage vacuoles. Nature 451, 350–354 (2008).
Welch, M. D., Iwamatsu, A. & Mitchison, T. J. Actin polymerization is induced by Arp2/3 protein complex at the surface of Listeria monocytogenes. Nature 385, 265–269 (1997).
Chakraborty, T. et al. A focal adhesion factor directly linking intracellularly motile Listeria monocytogenes and Listeria ivanovii to the actin-based cytoskeleton of mammalian cells. EMBO J. 14, 1314–1321 (1995).
Footer, M. J., Lyo, J. K. & Theriot, J. A. Close packing of Listeria monocytogenes ActA, a natively unfolded protein, enhances F-actin assembly without dimerization. J. Biol. Chem. 283, 23852–23862 (2008).
Portnoy, D. A., Chakraborty, T., Goebel, W. & Cossart, P. Molecular determinants of Listeria monocytogenes pathogenesis. Infect. Immun. 60, 1263–1267 (1992).
Pistor, S. et al. Mutations of arginine residues within the 146-KKRRK-150 motif of the ActA protein of Listeria monocytogenes abolish intracellular motility by interfering with the recruitment of the Arp2/3 complex. J. Cell Sci. 113 (Pt 18), 3277–3287 (2000).
Kuma, A. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036 (2004).
Bjorkoy, G. et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J. Cell Biol. 171, 603–614 (2005).
Komatsu, M. et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131, 1149–1163 (2007).
Gal, J., Strom, A. L., Kilty, R., Zhang, F. & Zhu, H. p62 accumulates and enhances aggregate formation in model systems of familial amyotrophic lateral sclerosis. J. Biol. Chem. 282, 11068–11077 (2007).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitylated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Long, J. et al. Ubiquitin recognition by the UBA domain of p62 involves a novel conformational switch. J. Biol. Chem. 283, 5427–5440 (2008).
Van Troys, M. et al. The actin propulsive machinery: the proteome of Listeria monocytogenes tails. Biochem. Biophys. Res. Commun. 375, 194–199 (2008).
Yano, T. et al. Autophagic control of listeria through intracellular innate immune recognition in drosophila. Nature Immunol. 9, 908–916 (2008).
Acknowledgements
We thank all members of the Sasakawa laboratory for discussions and technical advice and A. Amend and N. Schlarenko for help with the generation of several ActA mutants. This work was supported by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan to M.O. and C.S, the Special Coordination Funds for Promoting Science from the Japan Science and Technology Agency to C.S, and the ERA-NET Pathogenomics Network funded by the German Ministry of Education and Research and the EU to T.H. and T.C.
Author information
Authors and Affiliations
Contributions
C.S. and T.C. conceived the study. Y.Y. and M.O. performed experiments. Y.Y., M.O., C.S. and T.C. analysed data. M.F., K.M., M.H. and I.N. provided technical advice. M.Y. performed electon microscopic analysis. T.Y. and T.I. generated p62-null mouse embryonic fibroblasts. A.K. generated the Q79C plasmid. E.S. generated the GFP-170* plasmid. T.H. and T.C. generated bacterial strains and antibodies. C.S. and T.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 3672 kb)
Supplementary Information
Supplementary Movie 1 (MOV 4018 kb)
Supplementary Information
Supplementary Movie 2 (MOV 8135 kb)
Supplementary Information
Supplementary Movie 3 (MOV 3769 kb)
Supplementary Information
Supplementary Movie 4 (MOV 8404 kb)
Supplementary Information
Supplementary Movie 5 (MOV 3466 kb)
Supplementary Information
Supplementary Movie 6 (MOV 10188 kb)
Supplementary Information
Supplementary Movie 7 (MOV 3634 kb)
Supplementary Information
Supplementary Movie 8 (MOV 7396 kb)
Rights and permissions
About this article
Cite this article
Yoshikawa, Y., Ogawa, M., Hain, T. et al. Listeria monocytogenes ActA-mediated escape from autophagic recognition. Nat Cell Biol 11, 1233–1240 (2009). https://doi.org/10.1038/ncb1967
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1967
This article is cited by
-
A glycine-rich PE_PGRS protein governs mycobacterial actin-based motility
Nature Communications (2022)
-
The crosstalk between bacteria and host autophagy: host defense or bacteria offense
Journal of Microbiology (2022)
-
Tailored co-localization analysis of intracellular microbes and punctum-distributed phagosome–lysosome pathway proteins using ImageJ plugin EzColocalization
Scientific Reports (2021)
-
Lysosome repositioning as an autophagy escape mechanism by Mycobacterium tuberculosis Beijing strain
Scientific Reports (2021)
-
Regulation of eosinophil functions by autophagy
Seminars in Immunopathology (2021)