Abstract
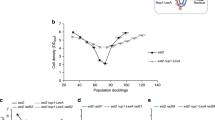
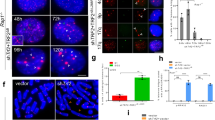
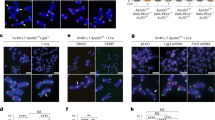
The ends of linear eukaryotic chromosomes are protected by telomeres, which serve to ensure proper chromosome replication and to prevent spurious recombination at chromosome ends. In this study, we show by single cell analysis that in the absence of telomerase, a single short telomere is sufficient to induce the recruitment of checkpoint and recombination proteins. Notably, a DNA damage response at eroded telomeres starts many generations before senescence and is characterized by the recruitment of Cdc13 (cell division cycle 13), replication protein A, DNA damage checkpoint proteins and the DNA repair protein Rad52 into a single focus. Moreover, we show that eroded telomeres, although remaining at the nuclear periphery, move to the nuclear pore complex. Our results link the DNA damage response at eroded telomeres to changes in subnuclear localization and suggest the existence of collapsed replication forks at eroded telomeres.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Gilson, E. & Geli, V. How telomeres are replicated. Nature Rev. Mol. Cell Biol. 8, 825–838 (2007).
Wellinger, R. J., Wolf, A. J. & Zakian, V. A. Saccharomyces telomeres acquire single-strand TG1–3 tails late in S phase. Cell 72, 51–60 (1993).
Larrivee, M., LeBel, C. & Wellinger, R. J. The generation of proper constitutive G-tails on yeast telomeres is dependent on the MRX complex. Genes Dev. 18, 1391–1396 (2004).
Vodenicharov, M. D. & Wellinger, R. J. DNA degradation at unprotected telomeres in yeast is regulated by the CDK1 (Cdc28/Clb) cell-cycle kinase. Mol. Cell 24, 127–137 (2006).
Bertuch, A. A. & Lundblad, V. The Ku heterodimer performs separable activities at double-strand breaks and chromosome termini. Mol. Cell. Biol. 23, 8202–8215 (2003).
Negrini, S., Ribaud, V., Bianchi, A. & Shore, D. DNA breaks are masked by multiple Rap1 binding in yeast: implications for telomere capping and telomerase regulation. Genes Dev. 21, 292–302 (2007).
Longhese, M. P. DNA damage response at functional and dysfunctional telomeres. Genes Dev. 22, 125–140 (2008).
Hug, N. & Lingner, J. Telomere length homeostasis. Chromosoma 115, 413–425 (2006).
Lundblad, V. & Szostak, J. W. A mutant with a defect in telomere elongation leads to senescence in yeast. Cell 57, 633–643 (1989).
Grandin, N., Bailly, A. & Charbonneau, M. Activation of Mrc1, a mediator of the replication checkpoint, by telomere erosion. Biol. Cell 97, 799–814 (2005).
d'Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198 (2003).
Takai, H., Smogorzewska, A. & de Lange, T. DNA damage foci at dysfunctional telomeres. Curr. Biol. 13, 1549–1556 (2003).
Bhattacharyya, M. K. & Lustig, A. J. Telomere dynamics in genome stability. Trends Biochem. Sci. 31, 114–122 (2006).
Le, S., Moore, J. K., Haber, J. E. & Greider, C. W. RAD50 and RAD51 define two pathways that collaborate to maintain telomeres in the absence of telomerase. Genetics 152, 143–152 (1999).
Krogh, B. & Symington, L. Recombination proteins in yeast. Annu. Rev. Genet. 38, 233–271 (2004).
Lisby, M., Barlow, J. H., Burgess, R. C. & Rothstein, R. Choreography of the DNA damage response; spatiotemporal relationships among checkpoint and repair proteins. Cell 118, 699–713 (2004).
Lisby, M., Mortensen, U. H. & Rothstein, R. Colocalization of multiple DNA double-strand breaks at a single Rad52 repair centre. Nature Cell Biol. 5, 572–577 (2003).
Marcand, S., Brevet, V. & Gilson, E. Progressive cis-inhibition of telomerase upon telomere elongation. EMBO J. 18, 3509–3519 (1999).
Gotta, M. et al. The clustering of telomeres and colocalization with Rap1, Sir3, and Sir4 proteins in wild-type Saccharomyces cerevisiae. J. Cell Biol. 134, 1349–1363 (1996).
Polotnianka, R. M., Li, J. & Lustig, A. J. The yeast Ku heterodimer is essential for protection of the telomere against nucleolytic and recombinational activities. Curr. Biol. 8, 831–834 (1998).
Adams Martin, A., Dionne, I., Wellinger, R. J. & Holm, C. The function of DNA polymerase alpha at telomeric G tails is important for telomere homeostasis. Mol. Cell. Biol. 20, 786–796 (2000).
Doye, V., Wepf, R. & Hurt, E. C. A novel nuclear pore protein Nup133p with distinct roles in poly(A)+ RNA transport and nuclear pore distribution. EMBO J. 13, 6062–6075 (1994).
Nugent, C. I., Hughes, T. R., Lue, N. F. & Lundblad, V. Cdc13p: a single-strand telomeric DNA-binding protein with a dual role in yeast telomere maintenance. Science 274, 249–252 (1996).
Lin, J. J. & Zakian, V. A. The Saccharomyces CDC13 protein is a single-strand TG1–3 telomeric DNA-binding protein in vitro that affects telomere behavior in vivo. Proc. Natl. Acad. Sci. USA 93, 13760–13765 (1996).
Hackett, J. A. & Greider, C. W. End resection initiates genomic instability in the absence of telomerase. Mol. Cell. Biol. 23, 8450–8461 (2003).
Chang, M., Arneric, M. & Lingner, J. Telomerase repeat addition processivity is increased at critically short telomeres in a Tel1-dependent manner in Saccharomyces cerevisiae. Genes Dev. 21, 2485–2494 (2007).
Nagai, S. et al. Functional targeting of DNA damage to a nuclear pore-associated SUMO-dependent ubiquitin ligase. Science 322, 597–602 (2008).
Sherman, F., Fink, G. R. & Hicks, J. B. Methods in Yeast Genetics. (Cold Spring Harbor Laboratory, 1986).
Belli, G., Gari, E., Aldea, M. & Herrero, E. Functional analysis of yeast essential genes using a promoter-substitution cassette and the tetracycline-regulatable dual expression system. Yeast 14, 1127–1138 (1998).
Lisby, M., Rothstein, R. & Mortensen, U. H. Rad52 forms DNA repair and recombination centers during S phase. Proc. Natl. Acad. Sci. USA 98, 8276–8282 (2001).
Forstemann, K., Hoss, M. & Lingner, J. Telomerase-dependent repeat divergence at the 3′ ends of yeast telomeres. Nucl. Acids Res. 28, 2690–2694 (2000).
Acknowledgements
We thank members of the Géli, Gilson and Lisby laboratories for helpful discussions concerning this work, A. Nicolas, X. Zhao, R. Rothstein, B. Palancade, V. Lundblad and R. Wellinger for sharing reagents and for fruitful discussions, S. Larose and R. Wellinger, who engineered the pTet-off-TLC1 construct, and S. Brill for the anti-RPA antibody. This work was supported by The Danish Agency for Science, Technology and Innovation (M.L.), the Villum Kann Rasmussen Foundation (M.L.), the Deutscher Akademischer Austausch Dienst (S.M.G.) and the Lundbeck Foundation (N.E.B.). The Agence Nationale de la Recherche (ANR programme blanc) and the Ligue Nationale Contre le Cancer (LNCC, équipes labelisées) supported V.G. and E.G. laboratories; B.K. is a recipient of a fellowship from the LNNC and P.A. is supported by the Lebanese National council for Scientific Research (CNRSL) and the Association pour la Recherche sur le Cancer (ARC).
Author information
Authors and Affiliations
Contributions
B.K., P.L. and M.N.S. performed the ChIP and telomeric blot analyses. N.E.B., I.G. and M.L. performed the microscopy. B.K., N.E.B., S.M.G., M.T.T. and P.A. constructed strains and assisted with data analysis. M.L. and P.L. performed telomere length PCR. V.G., M.L., E.G. and N.E.B. conceived and designed research, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1243 kb)
Rights and permissions
About this article
Cite this article
Khadaroo, B., Teixeira, M., Luciano, P. et al. The DNA damage response at eroded telomeres and tethering to the nuclear pore complex. Nat Cell Biol 11, 980–987 (2009). https://doi.org/10.1038/ncb1910
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1910
This article is cited by
-
A R-loop sensing pathway mediates the relocation of transcribed genes to nuclear pore complexes
Nature Communications (2023)
-
Telomere shortening causes distinct cell division regimes during replicative senescence in Saccharomyces cerevisiae
Cell & Bioscience (2021)
-
DNA repair by Rad52 liquid droplets
Nature Communications (2020)
-
The nuclear pore primes recombination-dependent DNA synthesis at arrested forks by promoting SUMO removal
Nature Communications (2020)
-
The nuclear pore complex prevents sister chromatid recombination during replicative senescence
Nature Communications (2020)