Abstract
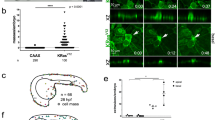
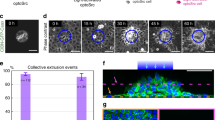
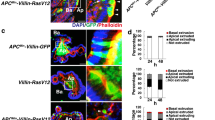
In most cancers, transformation begins in a single cell in an epithelial cell sheet1,2,3. However, it is not known what happens at the interface between non-transformed (normal) and transformed cells once the initial transformation has occurred. Using Madin-Darby canine kidney (MDCK) epithelial cells that express constitutively active, oncogenic Ras (RasV12) in a tetracycline-inducible system, we investigated the cellular processes arising at the interface between normal and transformed cells. We show that two independent phenomena occur in a non-cell-autonomous manner: when surrounded by normal cells, RasV12 cells are either apically extruded from the monolayer, or form dynamic basal protrusions and invade the basal matrix. Neither apical extrusion nor basal protrusion formation is observed when RasV12 cells are surrounded by other RasV12 cells. We show that Cdc42 and ROCK (also known as Rho kinase) have vital roles in these processes. We also demonstrate that E-cadherin knockdown in normal cells surrounding RasV12 cells reduces the frequency of apical extrusion, while promoting basal protrusion formation and invasion. These results indicate that RasV12-transformed cells are able to recognize differences between normal and transformed cells, and consequently leave epithelial sheets either apically or basally, in a cell-context-dependent manner.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Fialkow, P. J. Clonal origin of human tumors. Biochim. Biophys. Acta. 458, 283–321 (1976).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Schoenenberger, C. A., Zuk, A., Kendall, D. & Matlin, K. S. Multilayering and loss of apical polarity in MDCK cells transformed with viral K-ras. J. Cell Biol. 112, 873–889 (1991).
Rosenblatt, J., Raff, M. C. & Cramer, L. P. An epithelial cell destined for apoptosis signals its neighbors to extrude it by an actin- and myosin-dependent mechanism. Curr. Biol. 11, 1847–1857 (2001).
Bos, J. L. Ras-like GTPases. Biochim. Biophys. Acta. 1333, M19–31 (1997).
Baena-Lopez, L. A., Pastor-Pareja, J. C. & Resino, J. Wg and Egfr signalling antagonise the development of the peripodial epithelium in Drosophila wing discs. Development 130, 6497–6506 (2003).
Rodriguez-Viciana, P. et al. Role of phosphoinositide 3-OH kinase in cell transformation and control of the actin cytoskeleton by Ras. Cell 89, 457–467 (1997).
Cantrell, D. A. Phosphoinositide 3-kinase signalling pathways. J. Cell Sci. 114, 1439–1445 (2001).
Karnoub, A. E. & Weinberg, R. A. Ras oncogenes: split personalities. Nature Rev. Mol. Cell Biol. 9, 517–531 (2008).
Kimura, K. et al. Regulation of myosin phosphatase by Rho and Rho-associated kinase (Rho-kinase). Science 273, 245–248 (1996).
Jaffe, A. B. & Hall, A. Rho GTPases: biochemistry and biology. Annu. Rev. Cell Dev. Biol. 21, 247–269 (2005).
Vega, F. M. & Ridley, A. J. Rho GTPases in cancer cell biology. FEBS Lett. 582, 2093–2101 (2008).
Sahai, E., Olson, M. F. & Marshall, C. J. Cross-talk between Ras and Rho signalling pathways in transformation favours proliferation and increased motility. EMBO J. 20, 755–766 (2001).
de la Cova, C., Abril, M., Bellosta, P., Gallant, P. & Johnston, L. A. Drosophila Myc regulates organ size by inducing cell competition. Cell 117, 107–116 (2004).
Moreno, E. & Basler, K. dMyc transforms cells into super-competitors. Cell 117, 117–129 (2004).
Vidal, M., Larson, D. E. & Cagan, R. L. Csk-deficient boundary cells are eliminated from normal Drosophila epithelia by exclusion, migration, and apoptosis. Dev. Cell 10, 33–44 (2006).
Brumby, A. M. & Richardson, H. E. scribble mutants cooperate with oncogenic Ras or Notch to cause neoplastic overgrowth in Drosophila. EMBO J. 22, 5769–5779 (2003).
Stoker, M. G., Shearer, M. & O'Neill, C. Growth inhibition of polyoma-transformed cells by contact with static normal fibroblasts. J. Cell Sci. 1, 297–310 (1966).
Bignami, M., Rosa, S., La Rocca, S. A., Falcone, G. & Tato, F. Differential influence of adjacent normal cells on the proliferation of mammalian cells transformed by the viral oncogenes myc, ras and src. Oncogene 2, 509–514 (1988).
Alexander, D. B. et al. Normal cells control the growth of neighboring transformed cells independent of gap junctional communication and SRC activity. Cancer Res. 64, 1347–1358 (2004).
Wells, C. M., Walmsley, M., Ooi, S., Tybulewicz, V. & Ridley, A. J. Rac1-deficient macrophages exhibit defects in cell spreading and membrane ruffling but not migration. J. Cell Sci. 117, 1259–1268 (2004).
Hogan, C. et al. Rap1 regulates the formation of E-cadherin-based cell-cell contacts. Mol. Cell. Biol. 24, 6690–6700 (2004).
Acknowledgements
We thank G. K. Ojakian for the anti-gp135 antibody, A. Hall, A. Lloyd, R. Y. Tsien, S. Lowe and E. Sahai for constructs, and A. Vaughan for technical assistance with microscopes. We also thank Y. Morishita for discussion on physical forces at cell–cell adhesions. S.D-C. was supported by a FEBS Long Term Fellowship. A.E.P. acknowledges the Interdisciplinary Research Collaboration (IRC) in Nanotechnology (Cambridge, EPSRC UK) and the Dr Mortimer and Theresa Sackler Trust for financial support. This work is supported by MRC funding of the Cell Biology Unit.
Author information
Authors and Affiliations
Contributions
C.H. designed the experiments and generated most of the data; S.D-C. established stable MDCK cell lines and performed statistical analyses (Fig. 1d); M.N. analysed clonal expression of RasV12, RasN17 and RasWT in Drosophila wing imaginal discs (Fig. 1e and Supplementary Information, Fig. S3); M.K. performed western blot analyses (Supplementary Information, Fig. S1b), immunofluorescence studies (Supplementary Information, Fig. S4a) and established stable MDCK cell lines; C.Z. performed western blot and time-lapse analyses (Supplementary Information, Figs S2a, d and S5e), and established stable MDCK cell lines; A.E.P. performed AFM experiments and analyses (Supplementary Information, Fig. S9); E.P., L.A.B-L. and J-P.V. analysed clonal expression of RasV12, RasN17 and RasWT in Drosophila wing imaginal discs (Fig. 1e and Supplementary Information, Fig. S3); Y.I. provided technical expertise on use of collagen; H.H. provided technical expertise on myosin-II; F.P. assisted with Drosophila experiments; Y.F. conceived and designed the study and acted as principal investigator.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 3948 kb)
Supplementary Information
Supplementary Movie 1 (MOV 3033 kb)
Supplementary Information
Supplementary Movie 2 (MOV 2748 kb)
Supplementary Information
Supplementary Movie 3 (MOV 1275 kb)
Supplementary Information
Supplementary Movie 4 (MOV 130 kb)
Rights and permissions
About this article
Cite this article
Hogan, C., Dupré-Crochet, S., Norman, M. et al. Characterization of the interface between normal and transformed epithelial cells. Nat Cell Biol 11, 460–467 (2009). https://doi.org/10.1038/ncb1853
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1853
This article is cited by
-
Topological data analysis of spatial patterning in heterogeneous cell populations: clustering and sorting with varying cell-cell adhesion
npj Systems Biology and Applications (2023)
-
The ECM and tissue architecture are major determinants of early invasion mediated by E-cadherin dysfunction
Communications Biology (2023)
-
A mathematical model with aberrant growth correction in tissue homeostasis and tumor cell growth
Journal of Mathematical Biology (2023)
-
Pleiotropic effects of cell competition between normal and transformed cells in mammalian cancers
Journal of Cancer Research and Clinical Oncology (2023)
-
Self-assembly of tessellated tissue sheets by expansion and collision
Nature Communications (2022)