Abstract
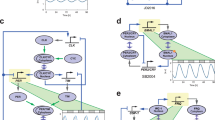
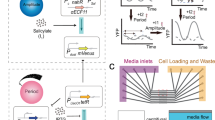
Mammalian circadian clocks consist of complex regulatory loops mediated through—at least—morning, daytime and night-time DNA elements. To prove the transcriptional logic of mammalian clocks, we developed an in cellulo mammalian cell-culture system that allowed us to design and implement artificial transcriptional circuits. Here we show that morning activation and night-time repression can yield the transcriptional output during the daytime, and similarly that daytime activation and morning repression can yield night-time transcriptional output. We also observed that the diverse transcriptional outputs of other phases can be generated through the expression of simple combinations of transcriptional activators and repressors. These results reveal design principles not only for understanding the continuous transcriptional outputs observed in vivo but also for the logical construction of artificial promoters working at novel phases. Logical synthesis of artificial circuits, with an identified structure and observed dynamics, provides an alternative strategy applicable to the investigation of complex biological systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hogenesch, J. B. et al. Characterization of a subset of the basic-helix–loop–helix-PAS superfamily that interacts with components of the dioxin signaling pathway. J. Biol. Chem. 272, 8581–8593 (1997).
Gekakis, N. Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569 (1998).
Yoo, S. H. et al. A noncanonical E-box enhancer drives mouse Period2 circadian oscillations in vivo. Proc. Natl Acad. Sci. USA 102, 2608–2613 (2005).
Ueda, H. R. et al. System-level identification of transcriptional circuits underlying mammalian circadian clocks. Nature Genet. 37, 187–192 (2005).
Falvey, E., Marcacci, L. & Schibler, U. DNA-binding specificity of PAR and C/EBP leucine zipper proteins: a single amino acid substitution in the C/EBP DNA-binding domain confers PAR-like specificity to C/EBP. Biol. Chem. 377, 797–809 (1996).
Harding, H. P. & Lazar, M. A. The orphan receptor Rev-ErbAα activates transcription via a novel response element. Mol. Cell. Biol. 13, 3113–3121 (1993).
Preitner, N. The orphan nuclear receptor REV-ERBα controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 110, 251–260 (2002).
Ueda, H. R. et al. A transcription factor response element for gene expression during circadian night. Nature 418, 534–539 (2002).
Reppert, S. M. & Weaver, D. R. Coordination of circadian timing in mammals. Nature 418, 935–941 (2002).
Bunger, M. K. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103, 1009–1017 (2000).
King, D. P. Positional cloning of the mouse circadian clock gene. Cell 89, 641–653 (1997).
Reick, M., Garcia, J. A., Dudley, C. & McKnight, S. L. NPAS2: an analog of clock operative in the mammalian forebrain. Science 293, 506–509 (2001).
Okano, T., Sasaki, M. & Fukada, Y. Cloning of mouse BMAL2 and its daily expression profile in the suprachiasmatic nucleus: a remarkable acceleration of Bmal2 sequence divergence after Bmal gene duplication. Neurosci. Lett. 300, 111–114 (2001).
Zheng, B. et al. The mPer2 gene encodes a functional component of the mammalian circadian clock. Nature 400, 169–173 (1999).
Shearman, L. P., Jin, X., Lee, C., Reppert, S. M. & Weaver, D. R. Targeted disruption of the mPer3 gene: subtle effects on circadian clock function. Mol. Cell. Biol. 20, 6269–6275 (2000).
Zheng, B. Nonredundant roles of the mPer1 and mPer2 genes in the mammalian circadian clock. Cell 105, 683–694 (2001).
Cermakian, N., Monaco, L., Pando, M. P., Dierich, A. & Sassone-Corsi, P. Altered behavioral rhythms and clock gene expression in mice with a targeted mutation in the Period1 gene. EMBO J. 20, 3967–3974 (2001).
Bae, K. Differential functions of mPer1, mPer2, and mPer3 in the SCN circadian clock. Neuron 30, 525–536 (2001).
van der Horst, G. T. Mammalian Cry1 and Cry2 are essential for maintenance of circadian rhythms. Nature 398, 627–630 (1999).
Vitaterna, M. H. et al. Differential regulation of mammalian period genes and circadian rhythmicity by cryptochromes 1 and 2. Proc. Natl Acad. Sci. USA 96, 12114–12119 (1999).
Honma, S. Dec1 and Dec2 are regulators of the mammalian molecular clock. Nature 419, 841–844 (2002).
Lowrey, P. L. Positional syntenic cloning and functional characterization of the mammalian circadian mutation tau. Science 288, 483–491 (2000).
Xu, Y. et al. Functional consequences of a CKIδ mutation causing familial advanced sleep phase syndrome. Nature 434, 640–644 (2005).
Lavery, D. J. et al. Circadian expression of the steroid 15 α-hydroxylase (Cyp2a4) and coumarin 7-hydroxylase (Cyp2a5) genes in mouse liver is regulated by the PAR leucine zipper transcription factor DBP. Mol. Cell. Biol. 19, 6488–6499 (1999).
Gachon, F. et al. The loss of circadian PAR bZip transcription factors results in epilepsy. Genes Dev. 18, 1397–1412 (2004).
Mitsui, S., Yamaguchi, S., Matsuo, T., Ishida, Y. & Okamura, H. Antagonistic role of E4BP4 and PAR proteins in the circadian oscillatory mechanism. Genes Dev. 15, 995–1006 (2001).
Sato, T. K. A functional genomics strategy reveals Rora as a component of the mammalian circadian clock. Neuron 43, 527–537 (2004).
André, E. et al. Disruption of retinoid-related orphan receptor β changes circadian behavior, causes retinal degeneration and leads to vacillans phenotype in mice. EMBO J. 17, 3867–3877 (1998).
Sato, T. K. et al. Feedback repression is required for mammalian circadian clock function. Nature Genet. 38, 312–319 (2006).
Akashi, M. & Takumi, T. The orphan nuclear receptor RORα regulates circadian transcription of the mammalian core-clock Bmal1. Nature Struct. Mol. Biol. 12, 441–448 (2005).
Sadowski, I., Ma, J., Triezenberg, S. & Ptashne, M. GAL4–VP16 is an unusually potent transcriptional activator. Nature 335, 563–564 (1988).
Gallego, M. & Virshup, D. M. Post-translational modifications regulate the ticking of the circadian clock. Nature Rev. Mol. Cell Biol. 8, 139–148 (2007).
Kojima, S., Hirose, M., Tokunaga, K., Sakaki, Y. & Tei, H. Structural and functional analysis of 3′ untranslated region of mouse Period1 mRNA. Biochem. Biophys. Res. Commun. 301, 1–7 (2003).
Lee, C., Etchegaray, J., Cagampang, F. R., Loudon, A. S. & Reppert, S. M. Posttranslational mechanisms regulate the mammalian circadian clock. Cell 107, 855–867 (2001).
Eide, E. J., Vielhaber, E. L., Hinz, W. A. & Virshup, D. M. The circadian regulatory proteins BMAL1 and cryptochromes are substrates of casein kinase Iepsilon. J. Biol. Chem. 277, 17248–17254 (2002).
Sanada, K., Okano, T. & Fukada, Y. Mitogen-activated protein kinase phosphorylates and negatively regulates basic helix–loop–helix-PAS transcription factor BMAL1. J. Biol. Chem. 277, 267–271 (2002).
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043 (2001).
Yin, L., Wang, J., Klein, P. S. & Lazar, M. A. Nuclear receptor Rev-erbα is a critical lithium-sensitive component of the circadian clock. Science 311, 1002–1005 (2006).
Akashi, M., Tsuchiya, Y., Yoshino, T. & Nishida, E. Control of intracellular dynamics of mammalian period proteins by casein kinase I ε (CKIε) and CKIδ in cultured cells. Mol. Cell. Biol. 22, 1693–1703 (2002).
Yagita, K. et al. Nucleocytoplasmic shuttling and mCRY-dependent inhibition of ubiquitylation of the mPER2 clock protein. EMBO J. 21, 1301–1314 (2002).
Cardone, L. et al. Circadian clock control by SUMOylation of BMAL1. Science 309, 1390–1394 (2005).
Vielhaber, E. L., Duricka, D., Ullman, K. S. & Virshup, D. M. Nuclear export of mammalian PERIOD proteins. J. Biol. Chem. 276, 45921–45927 (2001).
Ripperger, J. A. & Schibler, U. Rhythmic CLOCK-BMAL1 binding to multiple E-box motifs drives circadian Dbp transcription and chromatin transitions. Nature Genet. 38, 369–374 (2006).
Akhtar, R. A. et al. Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr. Biol. 12, 540–550 (2002).
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002).
Storch, K. F. et al. Extensive and divergent circadian gene expression in liver and heart. Nature 417, 78–83 (2002).
Salghetti, S. E., Caudy, A. A., Chenoweth, J. G. & Tansey, W. P. Regulation of transcriptional activation domain function by ubiquitin. Science 293, 1651–1653 (2001).
Lee, K., Loros, J. J. & Dunlap, J. C. Interconnected feedback loops in the Neurospora circadian system. Science 289, 107–110 (2000).
Dunlap, J. C. et al. A circadian clock in Neurospora: how genes and proteins cooperate to produce a sustained, entrainable, and compensated biological oscillator with a period of about a day. Cold Spring Harb. Symp. Quant. Biol. 72, 57–68 (2007).
Acknowledgements
This research was supported by an intramural Grant-in-Aid from CDB (H.R.U.), the Uehara Memorial Foundation (H.R.U.) and KAKENHI (Grant-in-Aid for Scientific Research) on Priority Areas 'Systems Genomics' from the Ministry of Education, Culture, Sports, Science and Technology of Japan (H.R.U.). We thank Hideki Ukai for critical discussion, Rikuhiro G. Yamada and Tetsuya J. Kobayashi for programming the real-time bioluminescence data analysis, Yuichi Kumaki for the Promoter Database, and Erik A. Kuld, Douglas Sipp and Michael Royle for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
H.R.U. developed the concept of the proof-by-synthesis of the transcriptional logic underlying mammalian circadian clocks. M.U. constructed all materials used in this work and performed the real-time luminescence assays. T.K. developed the data analysis methods for the in cellulo and in silico data, and conducted these analyses. All the authors discussed the results and commented on the manuscript text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 4002 kb)
Rights and permissions
About this article
Cite this article
Ukai-Tadenuma, M., Kasukawa, T. & Ueda, H. Proof-by-synthesis of the transcriptional logic of mammalian circadian clocks. Nat Cell Biol 10, 1154–1163 (2008). https://doi.org/10.1038/ncb1775
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1775
This article is cited by
-
Clock genes are expressed in cementum and regulate the proliferation and mineralization of cementoblasts
In Vitro Cellular & Developmental Biology - Animal (2023)
-
Circadian rhythms and cancers: the intrinsic links and therapeutic potentials
Journal of Hematology & Oncology (2022)
-
Explaining features of fine-grained phenomena using abstract analyses of phenomena and mechanisms: two examples from chronobiology
Synthese (2021)
-
Metabolic rivalry: circadian homeostasis and tumorigenesis
Nature Reviews Cancer (2020)
-
The adrenal gland circadian clock exhibits a distinct phase advance in spontaneously hypertensive rats
Hypertension Research (2019)