Abstract
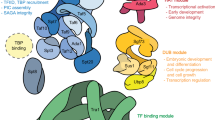
Targeting of a gene to the nuclear pore complexes (NPCs), known as gene gating, can affect its transcriptional state1,2,3,4,5,6,7,8,9. However, the mechanism underlying gene gating is poorly understood. Here, we have identified SAGA-associated Sgf73 (ref. 10), the yeast orthologue of human Ataxin-7 (ref. 11), as a regulator of histone H2B ubiquitin levels, a modification linked to both transcription initiation and elongation12,13. Sgf73 is a key component of a minimal histone-deubiquitinating complex. Activation of the H2B deubiquitinating protease, Ubp8, is cooperative and requires complex formation with the amino-terminal zinc-finger-containing domain of Sgf73 and Sgf11–Sus1. Through a separate domain, Sgf73 mediates recruitment of the TREX-2 mRNA export factors Sac3 and Thp1 to SAGA and their stable interaction with Sus1–Cdc31. This latter step is crucial to target TREX-2 to the NPC. Loss of Sgf73 from SAGA abrogates gene gating of GAL1 and causes a GAL1 mRNA export defect. Thus, Sgf73 provides a molecular scaffold to integrate the regulation of H2B ubiquitin levels, tethering of a gene to the NPC and export of mRNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Akhtar, A. & Gasser, S. M. The nuclear envelope and transcriptional control. Nature Rev. Genet. 8, 507–517 (2007).
Brickner, J. H. & Walter, P. Gene recruitment of the activated INO1 locus to the nuclear membrane. PLoS Biol. 2, e342 (2004).
Cabal, G. G. et al. SAGA interacting factors confine sub-diffusion of transcribed genes to the nuclear envelope. Nature 441, 770–773 (2006).
Casolari, J. M. et al. Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization. Cell 117, 427–439 (2004).
Dieppois, G., Iglesias, N. & Stutz, F. Cotranscriptional recruitment to the mRNA export receptor Mex67p contributes to nuclear pore anchoring of activated genes. Mol. Cell Biol. 26, 7858–7870 (2006).
Drubin, D. A., Garakani, A. M. & Silver, P. A. Motion as a phenotype: the use of live-cell imaging and machine visual screening to characterize transcription-dependent chromosome dynamics. BMC Cell Biol. 7, 19 (2006).
Taddei, A. et al. Nuclear pore association confers optimal expression levels for an inducible yeast gene. Nature 441, 774–778 (2006).
Abruzzi, K. C., Belostotsky, D. A., Chekanova, J. A., Dower, K. & Rosbash, M. 3´-end formation signals modulate the association of genes with the nuclear periphery as well as mRNP dot formation. EMBO J. 25, 4253–4262 (2006).
Dilworth, D. J. et al. The mobile nucleoporin Nup2p and chromatin-bound Prp20p function in endogenous NPC-mediated transcriptional control. J. Cell Biol. 171, 955–965 (2005).
McMahon, S. J., Pray-Grant, M. G., Schieltz, D., Yates, J. R., 3rd & Grant, P. A. Polyglutamine-expanded spinocerebellar ataxia-7 protein disrupts normal SAGA and SLIK histone acetyltransferase activity. Proc. Natl Acad. Sci. USA 102, 8478–8482 (2005).
Helmlinger, D. et al. Ataxin-7 is a subunit of GCN5 histone acetyltransferase-containing complexes. Hum. Mol. Genet. 13, 1257–1265 (2004).
Henry, K. W. et al. Transcriptional activation via sequential histone H2B ubiquitylation and deubiquitylation, mediated by SAGA-associated Ubp8. Genes Dev. 17, 2648–2663 (2003).
Pavri, R. et al. Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125, 703–717 (2006).
Daniel, J. A. & Grant, P. A. Multi-tasking on chromatin with the SAGA coactivator complexes. Mutat. Res. 618, 135–148 (2007).
Rodriguez-Navarro, S. et al. Sus1, a functional component of the SAGA histone acetylase complex and the nuclear pore-associated mRNA export machinery. Cell 116, 75–86 (2004).
Fischer, T. et al. Yeast centrin Cdc31 is linked to the nuclear mRNA export machinery. Nature Cell Biol. 6, 840–848 (2004).
Fischer, T. et al. The mRNA export machinery requires the novel Sac3p-Thp1p complex to dock at the nucleoplasmic entrance of the nuclear pores. EMBO J. 21, 5843–5852 (2002).
Ingvarsdottir, K. et al. H2B ubiquitin protease Ubp8 and Sgf11 constitute a discrete functional module within the Saccharomyces cerevisiae SAGA complex. Mol. Cell Biol. 25, 1162–1172 (2005).
Lee, K. K., Florens, L., Swanson, S. K., Washburn, M. P. & Workman, J. L. The deubiquitylation activity of Ubp8 is dependent upon Sgf11 and its association with the SAGA complex. Mol. Cell Biol. 25, 1173–1182 (2005).
Köhler, A. et al. The mRNA export factor Sus1 is involved in Spt/Ada/Gcn5 acetyltransferase-mediated H2B deubiquitinylation through its interaction with Ubp8 and Sgf11. Mol. Biol. Cell 17, 4228–4236 (2006).
Shukla, A., Stanojevic, N., Duan, Z., Sen, P. & Bhaumik, S. R. Ubp8p, a histone deubiquitinase whose association with SAGA is mediated by Sgf11p, differentially regulates lysine 4 methylation of histone H3 in vivo. Mol. Cell Biol. 26, 3339–3352 (2006).
Tanny, J. C., Erdjument-Bromage, H., Tempst, P. & Allis, C. D. Ubiquitylation of histone H2B controls RNA polymerase II transcription elongation independently of histone H3 methylation. Genes Dev. 21, 835–847 (2007).
Kao, C. F. et al. Rad6 plays a role in transcriptional activation through ubiquitylation of histone H2B. Genes Dev. 18, 184–195 (2004).
Xiao, T. et al. Histone H2B ubiquitylation is associated with elongating RNA polymerase II. Mol. Cell Biol. 25, 637–651 (2005).
Wyce, A. et al. H2B ubiquitylation acts as a barrier to Ctk1 nucleosomal recruitment prior to removal by Ubp8 within a SAGA-related complex. Mol. Cell 27, 275–288 (2007).
Daniel, J. A. et al. Deubiquitination of histone H2B by a yeast acetyltransferase complex regulates transcription. J. Biol. Chem. 279, 1867–1871 (2004).
Govind, C. K., Zhang, F., Qiu, H., Hofmeyer, K. & Hinnebusch, A. G. Gcn5 promotes acetylation, eviction and methylation of nucleosomes in transcribed coding regions. Mol. Cell 25, 31–42 (2007).
Bryant, G. O. & Ptashne, M. Independent recruitment in vivo by gal4 of two complexes required for transcription. Mol. Cell 11, 1301–1309 (2003).
Larschan, E. & Winston, F. The S. cerevisiae SAGA complex functions in vivo as a coactivator for transcriptional activation by Gal4. Genes Dev. 15, 1946–1956 (2001).
Palhan, V. B. et al. Polyglutamine-expanded ataxin-7 inhibits STAGA histone acetyltransferase activity to produce retinal degeneration. Proc. Natl Acad. Sci. USA 102, 8472–8477 (2005).
Shukla, A., Bajwa, P. & Bhaumik, S. R. SAGA-associated Sgf73p facilitates formation of the preinitiation complex assembly at the promoters either in a HAT-dependent or independent manner in vivo. Nucleic Acids Res. 34, 6225–6232 (2006).
Dang, L. C., Melandri, F. D. & Stein, R. L. Kinetic and mechanistic studies on the hydrolysis of ubiquitin C-terminal 7-amido-4-methylcoumarin by deubiquitinating enzymes. Biochemistry 37, 1868–1879 (1998).
Helmlinger, D., Tora, L. & Devys, D. Transcriptional alterations and chromatin remodeling in polyglutamine diseases. Trends Genet. 22, 562–570 (2006).
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nature Biotech. 17, 1030–1032 (1999).
Kao, C. F. & Osley, M. A. In vivo assays to study histone ubiquitylation. Methods 31, 59–66 (2003).
Acknowledgements
We thank Sabine Merker, Petra Ihrig and J. Lechner for performing mass spectrometry; Heiner Sähr for technical assitance and Silke Hauf (Max-Planck-Institut, Tübingen), Suzanne Elsasser, Dan Finley (Harvard Medical School, Boston) and Dieter Kressler for comments on the mansucript. We thank Mary Ann Osley for yeast strains and Catherine Dargemont for the GAL1 RNA probe. E.H. is a recipient of grants from the Deutsche Forschungsgemeinschaft (Leibniz Programme, SFB 638/B3) and Fonds der Chemischen Industrie. G.G.C. is a recipient of a fellowship from the Association pour la Researche sur le Cancer (ARC).
Author information
Authors and Affiliations
Contributions
A.K. and E.H. designed the project; A.K. and M.S. performed the experiments; G.G.C. conducted the gene gating assay (Fig. 5a) and was supervised by U.N.; A.K. and E.H. wrote the manuscript and all authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1, S2, S3, S4 and Supplementary Table 1 (PDF 1062 kb)
Rights and permissions
About this article
Cite this article
Köhler, A., Schneider, M., Cabal, G. et al. Yeast Ataxin-7 links histone deubiquitination with gene gating and mRNA export. Nat Cell Biol 10, 707–715 (2008). https://doi.org/10.1038/ncb1733
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1733
This article is cited by
-
SAGA–CORE subunit Spt7 is required for correct Ubp8 localization, chromatin association and deubiquitinase activity
Epigenetics & Chromatin (2020)
-
Dynamic modules of the coactivator SAGA in eukaryotic transcription
Experimental & Molecular Medicine (2020)
-
A yeast phenomic model for the influence of Warburg metabolism on genetic buffering of doxorubicin
Cancer & Metabolism (2019)
-
Chaperone-mediated ordered assembly of the SAGA and NuA4 transcription co-activator complexes in yeast
Nature Communications (2019)
-
SAGA DUBm-mediated surveillance regulates prompt export of stress-inducible transcripts for proteostasis
Nature Communications (2019)