Abstract
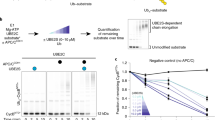
Protein ubiquitination regulates many cellular processes, including protein degradation, signal transduction, DNA repair and cell division. In the classical model, a uniform polyubiquitin chain that is linked through Lys 48 is required for recognition and degradation by the 26S proteasome. Here, we used a reconstituted system and quantitative mass spectrometry to demonstrate that cyclin B1 is modified by ubiquitin chains of complex topology, rather than by homogeneous Lys 48-linked chains. The anaphase-promoting complex was found to attach monoubiquitin to multiple lysine residues on cyclin B1, followed by poly-ubiquitin chain extensions linked through multiple lysine residues of ubiquitin (Lys 63, Lys 11 and Lys 48). These heterogeneous ubiquitin chains were sufficient for binding to ubiquitin receptors, as well as for degradation by the 26S proteasome, even when they were synthesized with mutant ubiquitin that lacked Lys 48. Together, our observations expand the context of what can be considered to be a sufficient degradation signal and provide unique insights into the mechanisms of substrate ubiquitination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Hershko, A. & Ciechanover, A. The ubiquitin system. Ann. Rev. Biochem. 67, 425–479 (1998).
Pickart, C. M. Back to the future with ubiquitin. Cell 116, 181–190 (2004).
Pickart, C. M. & Fushman, D. Polyubiquitin chains: polymeric protein signals. Curr. Opin. Chem. Biol. 8, 610–616 (2004).
Thrower, J. S., Hoffman, L., Rechsteiner, M. & Pickart, C. M. Recognition of the polyubiquitin proteolytic signal. EMBO J. 19, 94–102 (2000).
Finley, D. et al. Inhibition of proteolysis and cell cycle progression in a multiubiquitination-deficient yeast mutant. Mol. Cell. Biol. 14, 5501–5509 (1994).
Chau, V. et al. A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243, 1576–1583 (1989).
Deng, L. et al. Activation of the IκB kinase complex by TRAF6 requires a dimeric ubiquitin-conjugating enzyme complex and a unique polyubiquitin chain. Cell 103, 351–361 (2000).
Hofmann, R. M. & Pickart, C. M. Noncanonical MMS2-encoded ubiquitin-conjugating enzyme functions in assembly of novel polyubiquitin chains for DNA repair. Cell 96, 645–653 (1999).
Spence, J., Sadis, S., Haas, A. L. & Finley, D. A ubiquitin mutant with specific defects in DNA repair and multiubiquitination. Mol. Cell. Biol. 15, 1265–1273 (1995).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nature Biotechnol. 21, 921–926 (2003).
Peters, J. M. The anaphase-promoting complex: proteolysis in mitosis and beyond. Mol. Cell 9, 931–943 (2002).
Aristarkhov, A. et al. E2-C, a cyclin-selective ubiquitin carrier protein required for the destruction of mitotic cyclins. Proc. Natl Acad. Sci. USA 93, 4294–4299 (1996).
Yu, H., King, R. W., Peters, J. M. & Kirschner, M. W. Identification of a novel ubiquitin-conjugating enzyme involved in mitotic cyclin degradation. Curr. Biol. 6, 455–466 (1996).
Seino, H., Kishi, T., Nishitani, H. & Yamao, F. Two ubiquitin-conjugating enzymes, UbcP1/Ubc4 and UbcP4/Ubc11, have distinct functions for ubiquitination of mitotic cyclin. Mol. Cell. Biol. 23, 3497–3505 (2003).
Townsley, F. M. & Ruderman, J. V. Functional analysis of the Saccharomyces cerevisiae UBC11 gene. Yeast 14, 747–757 (1998).
Kirkpatrick, D. S., Denison, C. & Gygi, S. P. Weighing in on ubiquitin: the expanding role of mass spectrometry-based proteomics. Nature Cell Biol. 7, 750–757 (2005).
Gerber, S. A., Rush, J., Stemman, O., Kirschner, M. W. & Gygi, S. P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl Acad. Sci. USA 100, 6940–6945 (2003).
Raasi, S., Varadan, R., Fushman, D. & Pickart, C. M. Diverse polyubiquitin interaction properties of ubiquitin-associated domains. Nature Struct. Mol. Biol. 12, 708–714 (2005).
Elsasser, S., Chandler-Militello, D., Muller, B., Hanna, J. & Finley, D. Rad23 and Rpn10 serve as alternative ubiquitin receptors for the proteasome. J. Biol. Chem. 279, 26817–26822 (2004).
Elsasser, S. et al. Proteasome subunit Rpn1 binds ubiquitin-like protein domains. Nature Cell Biol. 4, 725–730 (2002).
Verma, R. et al. Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome. Science 298, 611–615 (2002).
Leggett, D. S. et al. Multiple associated proteins regulate proteasome structure and function. Mol. Cell 10, 495–507 (2002).
Mastrandrea, L. D., You, J., Niles, E. G. & Pickart, C. M. E2/E3-mediated assembly of lysine 29-linked polyubiquitin chains. J. Biol. Chem. 274, 27299–27306 (1999).
Johnson, E. S., Ma, P. C., Ota, I. M. & Varshavsky, A. A proteolytic pathway that recognizes ubiquitin as a degradation signal. J. Biol. Chem. 270, 17442–17456 (1995).
Petroski, M. D. & Deshaies, R. J. Mechanism of lysine 48-linked ubiquitin-chain synthesis by the cullin-RING ubiquitin-ligase complex SCF–Cdc34. Cell 123, 1107–1120 (2005).
Rape, M. & Kirschner, M. W. Autonomous regulation of the anaphase-promoting complex couples mitosis to S-phase entry. Nature 432, 588–595 (2004).
Ortolan, T. G. et al. The DNA repair protein rad23 is a negative regulator of multi-ubiquitin chain assembly. Nature Cell Biol. 2, 601–608 (2000).
Richly, H. et al. A series of ubiquitin binding factors connects CDC48/p97 to substrate multiubiquitylation and proteasomal targeting. Cell 120, 73–84 (2005).
Carroll, C. W. & Morgan, D. O. The Doc1 subunit is a processivity factor for the anaphase-promoting complex. Nature Cell Biol. 4, 880–887 (2002).
Rape, M., Reddy, S. K. & Kirschner, M. W. The processivity of multiubiquitination by the APC determines the order of substrate degradation. Cell 124, 89–103 (2006).
Guterman, A. & Glickman, M. H. Complementary roles for Rpn11 and Ubp6 in deubiquitination and proteolysis by the proteasome. J. Biol. Chem. 279, 1729–1738 (2004).
Hershko, A. & Heller, H. Occurrence of a polyubiquitin structure in ubiquitin-protein conjugates. Biochem. Biophys. Res. Commun. 128, 1079–1086 (1985).
Baboshina, O. V. & Haas, A. L. Novel multiubiquitin chain linkages catalyzed by the conjugating enzymes E2EPF and RAD6 are recognized by 26 S proteasome subunit 5. J. Biol. Chem. 271, 2823–2831 (1996).
Hofmann, R. M. & Pickart, C. M. In vitro assembly and recognition of Lys-63 polyubiquitin chains. J. Biol. Chem. 276, 27936–27943 (2001).
Flick, K. et al. Proteolysis-independent regulation of the transcription factor Met4 by a single Lys 48-linked ubiquitin chain. Nature Cell Biol. 6, 634–641 (2004).
Flick, K., Raasi, S., Zhang, H., Yen, J. L. & Kaiser, P. A ubiquitin-interacting motif protects polyubiquitinated Met4 from degradation by the 26S proteasome. Nature Cell Biol. 8, 509–515 (2006)
Petroski, M. D. & Deshaies, R. J. Context of multiubiquitin chain attachment influences the rate of Sic1 degradation. Mol. Cell 11, 1435–1444 (2003).
Acknowledgements
The authors wish to thank S. Gerber for technical advice and helpful discussions, and D.O. Morgan for the gift of cyclin B1 and cdc2 baculovirus. Additionally, the authors thank members of the Gygi, Finley and King labs for helpful discussions. R.W.K. is supported by a National Institutes of Health (NIH) grant (GM66492) and the McKenzie Family Foundation, and is a Damon Runyon Scholar. Work in the lab of D.F. is funded by a NIH grant (GM065592). Work in the lab of S.P.G. is supported by NIH grants (HG3456 and GM67945).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1, S2, S3, Supplementary Table S1 and Supplementary Text (PDF 398 kb)
Rights and permissions
About this article
Cite this article
Kirkpatrick, D., Hathaway, N., Hanna, J. et al. Quantitative analysis of in vitro ubiquitinated cyclin B1 reveals complex chain topology. Nat Cell Biol 8, 700–710 (2006). https://doi.org/10.1038/ncb1436
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1436
This article is cited by
-
Current methodologies in protein ubiquitination characterization: from ubiquitinated protein to ubiquitin chain architecture
Cell & Bioscience (2022)
-
Deubiquitinating enzymes and the proteasome regulate preferential sets of ubiquitin substrates
Nature Communications (2022)
-
Emerging functions of branched ubiquitin chains
Cell Discovery (2021)
-
Antibody toolkit reveals N-terminally ubiquitinated substrates of UBE2W
Nature Communications (2021)
-
The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates
Nature Communications (2020)