Abstract
Members of the dynamin family of GTPases have unique structural properties that might reveal a general mechanochemical basis for membrane constriction. Receptor-mediated endocytosis, caveolae internalization and certain trafficking events in the Golgi all require dynamin for vesiculation1. The dynamin-related protein Drp1 (Dlp1) has been implicated in mitochondria fission2 and a plant dynamin-like protein phragmoplastin is involved in the vesicular events leading to cell wall formation3. A common theme among these proteins is their ability to self-assemble into spirals and their localization to areas of membrane fission. Here we present the first three-dimensional structure of dynamin at a resolution of ∼20 Å, determined from cryo-electron micrographs of tubular crystals in the constricted state. The map reveals a T-shaped dimer consisting of three prominent densities: leg, stalk and head. The structure suggests that the dense stalk and head regions rearrange when GTP is added, a rearrangement that generates a force on the underlying lipid bilayer and thereby leads to membrane constriction. These results indicate that dynamin is a force-generating 'contrictase'.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hinshaw, J. E. Annu. Rev. Cell. Dev. Biol. 16, 483–519 (2000).
Erickson, H. P. J. Cell Biol. 148, 1103–1105 (2000).
Gu, X. & Verma, D. P. Plant Cell 9, 157–169 (1997).
Hinshaw, J. E. & Schmid, S. L. Nature 374, 190–192 (1995).
Kosaka, T. & Ikeda, K. J. Neurobiol. 14, 207–225 (1983).
Takei, K., McPherson, P. S., Schmid, S. L. & De Camilli, P. Nature 374, 186–190 (1995).
Sweitzer, S. M. & Hinshaw, J. E. Cell 93, 1021–1029 (1998).
Warnock, D. E., Baba, T. & Schmid, S. L. Mol. Biol. Cell 8, 2553–2562 (1997).
Muhlberg, A. B., Warnock, D. E. & Schmid, S. L. EMBO J. 16, 6676–6683 (1997).
Bauerfeind, R., Takei, K. & De Camilli, P. J. Biol. Chem. 272, 30984–30992 (1997).
Ringstad, N. et al. Neuron 24, 143–154 (1999).
Sever, S., Damke, H. & Schmid, S. L. J. Cell Biol. 150, 1137–1148 (2000).
Hill, E., van der Kaay, J., Downes, C. P. & Smythe, E. J. Cell Biol. 152, 309–324 (2001).
Binns, D. D. et al. J. Protein Chem. 18, 277–290 (1999).
Toyoshima, C. & Unwin, N. J. Cell Biol. 111, 2623–2635 (1990).
Burger, K. N., Demel, R. A., Schmid, S. L. & de Kruijff, B. Biochemistry 39, 12485–12493 (2000).
Ferguson, K. M., Lemmon, M. A., Schlessinger, J. & Sigler, P. B. Cell 79, 199–209 (1994).
Timm, D. et al. Nature Struct. Biol. 1, 782–788 (1994).
Prakash, B., Praefcke, G. J., Renault, L., Wittinghofer, A. & Herrmann, C. Nature 403, 567–571 (2000).
Achiriloaie, M., Barylko, B. & Albanesi, J. P. Mol. Cell. Biol. 19, 1410–1415 (1999).
Vallis, Y., Wigge, P., Marks, B., Evans, P. R. & McMahon, H. T. Curr. Biol. 9, 257–260 (1999).
Lee, A., Frank, D. W., Marks, M. S. & Lemmon, M. A. Curr. Biol. 9, 261–264 (1999).
Smirnova, E., Shurland, D. L., Newman-Smith, E. D., Pishvaee, B. & van der Bliek, A. M. J. Biol. Chem. 274, 14942–14947 (1999).
Sever, S., Muhlberg, A. B. & Schmid, S. L. Nature 398, 481–486 (1999).
Okamoto, P. M., Tripet, B., Litowski, J., Hodges, R. S. & Vallee, R. B. J. Biol. Chem. 274, 10277–10286 (1999).
Gilbert, A., Paccaud, J. P. & Carpentier, J. L. J. Cell Sci. 110, 3105–3115 (1997).
Marks, B. et al. Nature 410, 231–235 (2001).
Schmidt, A. et al. Nature 401, 133–141 (1999).
Stowell, M. H., Marks, B., Wigge, P. & McMahon, H. T. Nature Cell Biol. 1, 27–32 (1999).
Carr, J. F. & Hinshaw, J. E. J. Biol. Chem. 272, 28030–28035 (1997).
Beroukhim, R. & Unwin, N. Ultramicroscopy 70, 57–81 (1997).
Carragher, B., Whittaker, M. & Milligan, R. A. J. Struct. Biol. 116, 107–112 (1996).
Collaborative Computational Project, number 4. Acta Cryst. D 50, 760–763 (1994).
Acknowledgements
We thank J. Hanover for helpful suggestions and critical reading of the manuscript; workers in S. Schmid's laboratory for providing ΔPRD high titre stock; R. Milligan and A. Lin for assistance with Phoelix and VolVis; B. Sheehan for general computer help; N. Unwin for the use of the MRC helix-processing package; B. Carragher and D. Weber for help with Suprim; A. Steven and M. Cerritelli for assistance with the STEM analysis; B. Bowers for the rotary shadowing work; and F. Dyda, D. Belnap and T. Hirai for help with O. The BNL STEM is an NIH Supported Resource Center, with additional support provided by the Department of Energy, Office of Biological and Environmental Research.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary movie
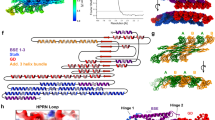
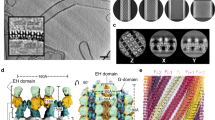
Movie 1 Quarter map of ΔPRD dynamin three-dimensional structure. The map shows two legs, one in front of the other, connected to the stalk region by a narrow linker. The stalk consists of a very dense globular domain that extends and bifurcates into two heads. The head has a distinct contour shape that accommodates the GTPase crystal structure (see Movie 2). (MOV 631 kb)
Supplementary movie
Movie 2 Docking the two crystal structures (PH17 and GTPase19) into the ΔDDPRD dynamin three-dimensional map.The movie demonstrates that GTPase (green) and PH (orange) crystal structures fit very well into the head and leg domains respectively. (MOV 565 kb)
Rights and permissions
About this article
Cite this article
Zhang, P., Hinshaw, J. Three-dimensional reconstruction of dynamin in the constricted state. Nat Cell Biol 3, 922–926 (2001). https://doi.org/10.1038/ncb1001-922
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1001-922
This article is cited by
-
Dynamin-dependent vesicle twist at the final stage of clathrin-mediated endocytosis
Nature Cell Biology (2021)
-
Mass photometry enables label-free tracking and mass measurement of single proteins on lipid bilayers
Nature Methods (2021)
-
Dynamin-2 R465W mutation induces long range perturbation in highly ordered oligomeric structures
Scientific Reports (2020)
-
Structure of a mitochondrial fission dynamin in the closed conformation
Nature Structural & Molecular Biology (2018)
-
Cryo-EM Studies of Drp1 Reveal Cardiolipin Interactions that Activate the Helical Oligomer
Scientific Reports (2017)