Abstract
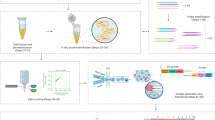
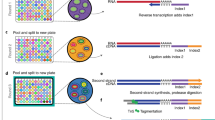
We describe a transcriptional analysis platform consisting of a universal micro-array system (UMAS) combined with an enzymatic manipulation step that is capable of generating expression profiles from any organism without requiring a priori species-specific knowledge of transcript sequences. The transcriptome is converted to cDNA and processed with restriction endonucleases to generate low-complexity pools (∼80–120) of equal length DNA fragments. The resulting material is amplified and detected with the UMAS system, comprising all possible 4,096 (46) DNA hexamers. Ligation to the arrays yields thousands of 14-mer sequence tags. The compendium of signals from all pools in the array-of-universal arrays comprises a full-transcriptome expression profile. The technology was validated by analysis of the galactose response of Saccharomyces cerevisiae, and the resulting profiles showed excellent agreement with the literature and real-time PCR assays. The technology was also used to demonstrate expression profiling from a hybrid organism in a proof-of-concept experiment where a T-cell receptor gene was expressed in yeast.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Janssen, P. et al. Beyond 100 Genomes. Genome Biol. 4, 402 (2003).
Liang, P. & Pardee, A. Differential display of eukaryotic messenger RNA by means of the polymerase chain reaction. Science 257, 967–971 (1992).
Saha, S. et al. Using the transcriptome to annotate the genome. Nat. Biotechnol. 19, 508–512 (2002).
Velculescu, V.E., Zhang, L., Vogelstein, B. & Kinzler, K.W. Serial analysis of gene expression. Science 270, 484–487 (1995).
Brenner, S. et al. Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat. Biotechnol. 18, 630–634 (2000).
Brenner, S. et al. In vitro cloning of complex mixtures of DNA on microbeads: physical separation of differentially expressed cDNAs. Proc. Natl. Acad. Sci. USA 97, 1665–1670 (2000).
Sutcliffe, J.G. et al. TOGA: An automated parsing technology for analyzing expression on nearly all genes. Proc. Natl. Acad. Sci. USA 97, 1976–1981 (2002).
Shimkets, R.A. et al. Gene expression analysis by transcript profiling coupled to a gene database query. Nat. Biotechnol. 17, 798–803 (1999).
Schena, M., Shalon, D., Davis, R.W. & Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Lockhart, D.J. et al. Expression monitoring by hybridization to high-density oligonucleotides arrays. Nat. Biotechnol. 14, 1675–1680 (1996).
Ramakrishnan, R. et al. An assessment of Motorola codelink microarray performance for expression profiling applications. Nucleic Acids Res. 30, e30, (2002).
Hughes, T.R. et al. Expression profiling using microarrays fabricated by an ink-jet oligonucleotides synthesizer. Nat. Biotechnol. 19, 342–347 (2001).
Hughes, T.R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
van Dam, R.M. & Quake, S.R. Gene expression analysis with universal n-mer array. Genome Res. 12, 145–152 (2002).
Housby, J.N. & Southern, E.M. Fidelity of DNA ligation: a novel experimental approach based on the polymerization of libraries of oligonucleotides. Nucleic Acids Res. 26, 4259–4266 (1998).
Prichard, C.E. & Southern, E.M. Effects of base mismatches on joining of short oligodeoxynulceotides by DNA ligases. Nucleic Acids Res. 25, 3403–3407 (1997).
Cherepanov, A., Yildirim, E. & de Vries, S. Joining of short DNA oligonucleotides with base pair mismatches by T4 DNA ligase. J. Biochem. 129, 61–68 (2001).
James, K.D., Boles, A.R., Henckel, D. & Ellington, A.D. The fidelity of template-directed oligonucleotides ligation and its relevance to DNA computation. Nucleic Acids Res. 26, 5203–5211 (1998).
James, K.D. & Ellington, A.D. Surprising fidelity of template-directed chemical ligation of oligonucleotides. Chem. Biol. 4, 595–605 (1997).
Rossi, R., Montecucco, A., Ciarrocchi, G. & Biamonti, G. Functional characterization of the T4 DNA ligase: a new insight into the mechanism of action. Nucleic Acids Res. 25, 2106–2113 (1997).
Aoi, Y., Yoshinobu, T., Tanizawa, K., Kinoshita, K. & Iwasaki, H. Ligation errors in DNA computing. Biosystems 52, 181–187 (1999).
Faulhammer, D., Lipton, R.J. & Landweber, L.F. Fidelity of enzymatic ligation for DNA computing. J. Comput. Bio. 7, 839–848 (2000).
Breslauer, K.J., Frank, R., Blocker, H. & Marky, L.A. Predicting DNA duplex stability from the base sequence. Proc. Natl. Acad. Sci. USA 83, 3746–3750 (1986).
Frierer, S.M. et al. Improved free-energy parameters for prediction of RNA duplex stability. Proc. Natl. Acad. Sci. USA 83, 9373–9377 (1986).
SantaLucia, J. Jr. A unified view of polymer, dumbbell, and oligonucleotides DNA nearest-neighbor thermodynamics. Proc. Natl. Acad. Sci. USA 95, 1460–1465 (1998).
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000).
Roth, F.P., Hughes, J.D., Estep, P.W. & Church, G.M. Finding DNA regulatory motifs within unaligned noncoding sequences clustered by whole-genome mRNA quantitation. Nat. Biotechnol. 16, 939–945 (1998).
Ideker, T. et al. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292, 929–934 (2001).
Oshiro, G. et al. Parallel identification of new genes in Saccharomyces cerevisiae. Genome Res. 12, 1210–1220 (2002).
Kumar, A. et al. An integrated approach for finding overlooked genes in yeast. Nat. Biotechnol. 20, 58–63 (2002).
Velculescu, V.E. et al. Characterization of the yeast transcriptome. Cell 88, 243–251 (1997).
Martin, V.J.J., Pitera, D.J., Withers, S.T., Newman, J.D. & Keasling, J.D. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat. Biotechnol. 21, 796–802 (2003).
Szczebara, F.M. et al. Total biosynthesis of hydrocortisone from a simple carbon source in yeast. Nat. Biotechnol. 21, 143–149 (2003).
Boder, E.T. & Wittrup, K.D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
Holler, P.D. et al. In vitro evolution of a T-cell receptor with high affinity for peptide/MHC. Proc. Natl. Acad. Sci. USA 97, 5387–5392 (2000).
Sasik, R., Calvo, E. & Corbeil, J. Statistical analysis of high-density oligonucleotides arrays: a multiplicative noise model. Bioinformatics 18, 1633–1640 (2002).
Rinn, J.L. et al. The transcriptional activity of human chromosome 22. Gen. & Dev. 17, 529–540.
Baug, L.R., Hill, A.A., Brown, E.L. & Hunter, C. Quantitative analysis of mRNA amplification by in vitro transcription. Nucleic Acids Res. 29, e29 (2001).
Demers, L.M. et al. Direct patterning of modified oligonucleotides on metals and insulators by dip-pen nanolithography. Science 296, 1836–1838 (2002).
Park, S.J., Taton, T.A. & Mirkin, C.A. Array-based electrical detection of DNA with nanoparticle probes. Science 295, 1503–1506 (2002).
Edgar, R., Domrachev, M. & Lash, A.E. Gene Expression Omnibus: NCBI gene and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Zhao, J., Hyman, L., Moore, C. & Smith, T.F. Genomic detection of new yeast pre-mRNA 3-end processing. Nucleic Acids Res. 27, 888–894 (1999).
Acknowledgements
The authors are grateful to Gavin Sherlock for expert technical advice; Nam-Hai Chua, Jon Morrow and Paul Kaplan for their scientific input and encouragement; Jose Lage, Grant Carlson and Gisela Carlson for comments on the manuscript; Christina Mityas, Mike Murtha, Yih-Woei Fridell and Shane Weber for their efforts in the early stages of this work; Tom Owen for technical assistance; and Meri Ross for manuscript preparation. We also thank all of our colleagues at Agilix for helpful discussions, continuing scientific input and excellent support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The Agilix employees who are listed as coauthors, and Junhyong Kim and Paul Lizardi (who are scientific advisors to the company), either have or are eligible for stock options through the company's stock option program.
Rights and permissions
About this article
Cite this article
Roth, M., Feng, L., McConnell, K. et al. Expression profiling using a hexamer-based universal microarray. Nat Biotechnol 22, 418–426 (2004). https://doi.org/10.1038/nbt948
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt948
This article is cited by
-
Repression of RAD51 gene expression by E2F4/p130 complexes in hypoxia
Oncogene (2007)
-
PathogenMIPer: a tool for the design of molecular inversion probes to detect multiple pathogens
BMC Bioinformatics (2006)
-
A sequence-oriented comparison of gene expression measurements across different hybridization-based technologies
Nature Biotechnology (2006)
-
Sociogenomics: social life in molecular terms
Nature Reviews Genetics (2005)
-
Unshackling expression profiling arrays
Nature Biotechnology (2004)