Abstract
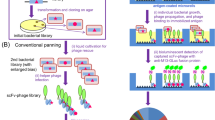
This report describes the selection of highly efficient antibody catalysts by combining chemical selection from a synthetic library with directed in vitro protein evolution. Evolution started from a naive antibody library displayed on phage made from fully synthetic, antibody-encoding genes (the Human Combinatorial Antibody Library; HuCAL-scFv). HuCAL-scFv was screened by direct selection for catalytic antibodies exhibiting phosphatase turnover. The substrate used was an aryl phosphate, which is spontaneously transformed into an electrophilic trapping reagent after cleavage. Chemical selection identified an efficient biocatalyst that then served as a template for error-prone PCR (epPCR) to generate randomized repertoires that were subjected to further selection cycles. The resulting superior catalysts displayed cumulative mutations throughout the protein sequence; the ten-fold improvement of their catalytic proficiencies (>1010 M−1) resulted from increased kcat values, thus demonstrating direct selection for turnover. The strategy described here makes the search for new catalysts independent of the immune system and the antibody framework.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schmidt-Dannert, C. Directed evolution of single proteins, metabolic pathways, and viruses. Biochemistry 40, 13125–13136 (2001).
Farinas, E.T., Bulter, T. & Arnold, F.H. Directed enzyme evolution. Curr. Opin. Biotechnol. 12, 545–551 (2001).
Griffiths, A.D. & Tawfik, D.S. Man-made enzymes—from design to in vitro compartmentalisation. Curr. Opin. Biotechnol. 11, 338–353 (2000).
Kazlauskas, R.J. Molecular modeling and biocatalysis: explanations, predictions, limitations, and opportunities. Curr. Opin. Chem. Biol. 4, 81–88 (2000).
Blackburn, G.M., Datta, A., Denham, H. & Wentworth, P. Jr. Catalytic antibodies. Adv. Phys. Org. Chem. 31, 249–392 (1998).
Hilvert, D. Critical analysis of antibody catalysis. Annu. Rev. Biochem. 69, 751–793 (2000).
Wentworth, P. Jr. & Janda, K.D. Catalytic antibodies: structure and function. Cell Biochem. Biophys. 35, 63–87 (2001).
Wirsching, P., Ashley, J.A., Lo, C.H., Janda, K.D. & Lerner, R.A. Reactive immunization. Science 270, 1775–1782 (1995).
Barbas, C.F., et al. Immune versus natural selection: antibody aldolases with enzymic rates but broader scope. Science 278, 2085–2092 (1997).
Gao, C. et al. Making chemistry selectable by linking it to infectivity. Proc. Natl. Acad. Sci. USA 94, 11777–11782 (1997).
Amstutz, P. et al. In vitro selection for catalytic activity with ribosome display. J. Am. Chem. Soc. 124, 9396–9403 (2002).
Janda, K.D. et al. Chemical selection for catalysis in combinatorial antibody libraries. Science 275, 945–948 (1997).
Soumillion, P. et al. Phage display of enzymes and in vitro selection for catalytic activity. Appl. Biochem. Biotechnol. 47, 175–190 (1994).
Tanaka, F., Lerner, R.A. & Barbas, C.F. Reconstructing aldolase antibodies to alter their substrate specificity and turnover. J. Am. Chem. Soc. 122, 4835–4836 (2000).
Danielsen, S. et al. In vitro selection of enzymatically active lipase variants from phage libraries using a mechanism-based inhibitor. Gene 272, 267–274 (2001).
Knappik, A. et al. Fully synthetic Human Combinatorial Antibody Libraries (HuCAL) based on modular consensus frameworks and CDRs randomized with trinucleotides. J. Mol. Biol. 296, 57–86 (2000).
Krebs, B. et al. High-throughput generation and engineering of recombinant human antibodies. J. Immunol. Methods 254, 67–84 (2001).
Betley, J.R. et al. Direct screening for phosphatase activity by turnover-based capture of protein catalysts. Angew. Chem., Int. Edn. Engl. 41, 775–777 (2002).
Myers, J.K., Cohen, J.D. & Widlanski, T.S. Substituent effects on the mechanism-based inactivation of prostatic acid phosphatase. J. Am. Chem. Soc. 117, 11049–11054 (1995).
Born, T.L., Myers, J.K., Widlanski, T.S. & Rusnak, F. 4-(Fluoromethyl)phenyl phosphate acts as a mechanism-based inhibitor of calcineurin. J. Biol. Chem. 270, 25651–25655 (1995).
Loubinoux, B., Miazimbakana, J. & Gerardin, P. Reactivity of new precursors of quinone methides. Tetrahedron Lett. 30, 1939–1942 (1989).
Vayron, P. et al. Toward antibody-catalyzed hydrolysis of organophosphorus poisons. Proc. Natl. Acad. Sci. USA 97, 7058–7063 (2000).
Scanlan, T.S., Prudent, J.R. & Schultz, P.G. Antibody-catalyzed hydrolysis of phosphate monoesters. J. Am. Chem. Soc. 113, 9397–9398 (1991).
Wentworth, P. Jr. et al. A bait and switch hapten strategy generates catalytic antibodies for phosphodiester hydrolysis. Proc. Natl. Acad. Sci. USA 95, 5971–5975 (1998).
Wentworth, P. & Janda, K.D. Catalytic antibodies. Curr. Opin. Chem. Biol. 2, 138–144 (1998).
Spivak, D.A., Hoffman, T.Z., Moore, A.H., Taylor, M.J. & Janda, K.D. A comparison of flexible and constrained haptens in eliciting antibody catalysts for paraoxon hydrolysis. Bioorg. Med. Chem. 7, 1145–1150 (1999).
Stec, B., Hehir, M.J., Brennan, C., Nolte, M. & Kantrowitz, E.R. Kinetic and X-ray structural studies of three mutant E. coli alkaline phosphatases: insights into the catalytic mechanism without the nucleophile Ser102. J. Mol. Biol. 277, 647–662 (1998).
Coleman, J.E. Structure and mechanism of alkaline phosphatase. Annu. Rev. Biophys. Biomol. Struct. 21, 441–483 (1992).
Thatcher, G.R.J. & Kluger, R. Mechanism and catalysis of nucleophilic substitution in phosphate esters. Adv. Phys. Org. Chem. 25, 99–265 (1989).
Peracchi, A. Enzyme catalysis: removing chemically 'essential' residues by site-directed mutagenesis. Trends Biochem. Sci. 26, 497–503 (2001).
Davies, D.R., Padlan, E.A. & Sheriff, S. Antibody-antigen complexes. Annu. Rev. Biochem. 59, 439–473 (1990).
Zaccolo, M., Williams, D.M., Brown, D.M. & Gherardi, E. An approach to random mutagenesis of DNA using mixtures of triphosphate derivatives of nucleoside analogues. J. Mol. Biol. 255, 589–603 (1996).
Arnold, F.H. When blind is better: protein design by evolution. Nat. Biotechnol. 16, 617–618 (1998).
Chen, Y.L., Tang, T.Y. & Cheng, K.J. Directed evolution to produce an alkalophilic variant from a Neocallimastix patriciarum xylanase. Can. J. Microbiol. 47, 1088–1094 (2001).
Christians, F.C. & Loeb, L.A. Novel human DNA alkyltransferases obtained by random substitution and genetic selection in bacteria. Proc. Natl. Acad. Sci. USA 93, 6124–6128 (1995).
Martinez, M.A., Pezo, V., Marliere, P. & Wain-Hobson, S. Exploring the functional robustness of an enzyme by in vitro evolution. EMBO J. 15, 1203–1210 (1995).
Oue, S., Okamoto, A., Yano, T. & Kagamiyama, H. Redesigning the substrate specificity of an enzyme by cumulative effects of the mutations of non-active site residues. J. Biol. Chem. 274, 2344–2349 (1999).
Daugherty, P.S., Chen, G., Iverson, B.L. & Georgiou, G. Quantitative analysis of the effect of the mutation frequency on the affinity maturation of single chain Fv antibodies. Proc. Natl. Acad. Sci. USA 97, 2029–2034 (2000).
Takahashi, N., Kakinuma, H., Liu, L., Nishi, Y. & Fujii, I. In vitro abzyme evolution to optimize antibody recognition for catalysis. Nat. Biotechnol. 19, 563–567 (2001).
Ladner, R.C. & Ley, A.C. Novel frameworks as a source of high-affinity ligands. Curr. Opin. Biotechnol. 12, 406–410 (2001).
Jürgens, C. et al. Directed evolution of a (βα)8-barrel enzyme to catalyze related reactions in two different metabolic pathways. Proc. Natl. Acad. Sci. USA 97, 9925–9930 (2000).
Hanes, J., Schaffitzel, C., Knappik, A. & Plückthun, A. Picomolar affinity antibodies from a fully synthetic naive library selected and evolved by ribosome display. Nat. Biotechnol. 18, 1287–1292 (2000).
Wolfenden, R., Ridgway, C. & Young, G. Spontaneous hydrolysis of ionized phosphate monoesters and diesters and the proficiencies of phosphohydrolases as catalysts. J. Am. Chem. Soc. 120, 833–834 (1998).
O'Brien, P.J. & Herschlag, D. Alkaline phosphatase revisited: hydrolysis of alkyl phosphates. Biochemistry 41, 3207–3225 (2002).
Honegger, A. & Plückthun, A. Yet another numbering scheme for immunoglobulin variable domains: an automatic modeling and analysis tool. J. Mol. Biol. 309, 657–670 (2001).
Kabat, E.A. & Wu, T.T. Identical V region amino acid sequences and segments of sequences in antibodies of different specificities. Relative contributions of VH and VL genes, minigenes, and complementarity-determining regions to binding of antibody-combining sites. J. Immunol. 147, 1709–1719 (1991).
Acknowledgements
S.C.-T. and D.L. were supported by Studentships under the European Commission Training and Mobility of Researchers program (grant ERBFMRXCT 980193). J.R. was supported by a Biotechnology and Biological Sciences Research Council Studentship. We are grateful to Aziz Mekhalfia and Jason Betley for assistance with chemical synthesis. We are indebted to Bernhard Schimmele, Lutz Jermutus, Stephen Marino and Patrik Forrer for help, advice and discussion, and to MorphoSys AG for the constructive collaboration on HuCAL-scFv.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
A.P. owns shares of Morphosys AG.
Rights and permissions
About this article
Cite this article
Cesaro-Tadic, S., Lagos, D., Honegger, A. et al. Turnover-based in vitro selection and evolution of biocatalysts from a fully synthetic antibody library. Nat Biotechnol 21, 679–685 (2003). https://doi.org/10.1038/nbt828
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt828
This article is cited by
-
Evolution of protease activation and specificity via alpha-2-macroglobulin-mediated covalent capture
Nature Communications (2023)
-
Compartmentalized partnered replication for the directed evolution of genetic parts and circuits
Nature Protocols (2017)
-
mRNA display for the selection and evolution of enzymes from in vitro-translated protein libraries
Nature Protocols (2011)
-
Kemp elimination catalysts by computational enzyme design
Nature (2008)
-
Selection and evolution of enzymes from a partially randomized non-catalytic scaffold
Nature (2007)