Abstract
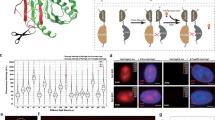
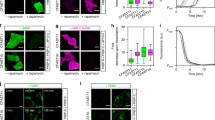
The specificity of biological regulatory mechanisms relies on selective interactions between different proteins in different cell types and in response to different extracellular signals. We describe a bimolecular fluorescence complementation (BiFC) approach for the simultaneous visualization of multiple protein interactions in the same cell. This approach is based on complementation between fragments of fluorescent proteins with different spectral characteristics. We have identified 12 bimolecular fluorescent complexes that correspond to 7 different spectral classes. Bimolecular complex formation between fragments of different fluorescent proteins did not differentially affect the dimerization efficiency of the bZIP domains of Fos and Jun or the subcellular sites of interactions between these domains. Multicolor BiFC enables visualization of interactions between different proteins in the same cell and comparison of the efficiencies of complex formation with alternative interaction partners.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hu, C.-D., Chinenov, Y. & Kerppola, T.K. Visualization of interactions among bZIP and Rel proteins in living cells using bimolecular fluorescence complementation. Mol. Cell 9, 789–798 (2002).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Miyawaki, A. et al. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388, 882–887 (1997).
Romoser, V.A., Hinkle, P.M. & Persechini, A. Detection in living cells of Ca2+-dependent changes in the fluorescence emission of an indicator composed of two green fluorescent protein variants linked by a calmodulin-binding sequence. A new class of fluorescent indicators. J. Biol. Chem. 272, 13270–13274 (1997).
Johnsson, N. & Varshavsky, A. Split ubiquitin as a sensor of protein interactions in vivo. Proc. Natl. Acad. Sci. USA 91, 10340–10344 (1994).
Rossi, F., Charlton, C.A. & Blau, H.M. Monitoring protein-protein interactions in intact eukaryotic cells by β-galactosidase complementation. Proc. Natl. Acad. Sci. USA 94, 8405–8410 (1997).
Pelletier, J.N., Campbell-Valois, F.X. & Michnick, S.W. Oligomerization domain-directed reassembly of active dihydrofolate reductase from rationally designed fragments. Proc. Natl. Acad. Sci. USA 95, 12141–12146 (1998).
Ghosh, I., Hamilton, A.D. & Regan, L. Antiparallel leucine zipper-directed protein reassembly: application to the green fluorescent protein. J. Am. Chem. Soc. 122, 5658–5659 (2000).
Galarneau, A., Primeau, M., Trudeau, L.E. & Michnick, S.W. Beta-Lactamase protein fragment complementation assays as in vivo and in vitro sensors of protein-protein interactions. Nat. Biotechnol. 20, 619–622 (2002).
Ormö, M. et al. Crystal structure of the Aequorea victoria green fluorescent protein. Science 273, 1392–1395 (1996).
Kerppola, T.K. & Curran, T. Selective DNA bending by a variety of bZIP proteins. Mol. Cell. Biol. 13, 5479–5489 (1993).
Tsai, E.Y. et al. A lipopolysaccharide-specific enhancer complex involving Ets, Elk-1, Sp1, and CREB binding protein and p300 is recruited to the tumor necrosis factor alpha promoter in vivo. Mol. Cell. Biol. 20, 6084–6094 (2000).
Falvo, J.V., Parekh, B.S., Lin, C.H., Fraenkel, E. & Maniatis, T. Assembly of a functional beta interferon enhanceosome is dependent on ATF-2-c-jun heterodimer orientation. Mol. Cell. Biol. 20, 4814–4825 (2000).
Daury, L. et al. Opposing functions of ATF2 and Fos-like transcription factors in c-Jun-mediated myogenin expression and terminal differentiation of avian myoblasts. Oncogene 20, 7998–8008 (2001).
O'Shea, E.K., Rutkowski, R., Stafford, W.D., Kim, P.S. & Stafford, W.F. Preferential heterodimer formation by isolated leucine zippers from fos and jun. Science 245, 646–648 (1989).
Ullmann, A., Jacob, F. & Monod, J. On the subunit structure of wild-type versus complemented β-galactosidase of Escherichia coli. J. Mol. Biol. 32, 1–13 (1968).
Kippen, A.D. & Fersht, A.R. Analysis of the mechanism of assembly of cleaved barnase from two peptide fragments and its relevance to the folding pathway of uncleaved barnase. Biochemistry 34, 1464–1468 (1995).
Koebnik, R. In vivo membrane assembly of split variants of the E. coli outer membrane protein OmpA. EMBO J. 15, 3529–3537 (1996).
Acknowledgements
We thank members of the Kerppola laboratory for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors have applied for a patent (application no. 60/375,949) on the molecular fluorescence complementation assay described in this paper.
Supplementary information
41587_2003_BFnbt816_MOESM1_ESM.jpg
Supplementary Fig. 1. Comparison of expression of bFos, bJun and bATF2 fused to fragments of different fluorescent proteins. (A) Plasmids encoding the fragments indicated fused to bJun and bFos respectively were co-transfected into cells. The YN155 used in this experiment (but not in others presented in this manuscript) was fused to Jun225-334, resulting in a larger fusion protein (lane 1). (B) Plasmids encoding the proteins indicated above each lane were co-transfected into cells. Cell lysates were analyzed by Western blotting using antibodies directed against the FLAG and HA epitopes. (JPG 82 kb)
41587_2003_BFnbt816_MOESM2_ESM.gif
Supplementary Fig. 2. Selectivity of bimolecular fluorescence complementation between fragments of fluorescent proteins fused to bFos and bJun. The excitation/emission maxima of all complexes that exhibited detectable fluorescence complementation are shown. None of the fragments exhibited detectable fluorescence alone. Cells with no detectable complementation (indicated by -) produced less than 1% of the fluorescence emissions of cells expressing YN155 and YC155 or CN155 and CC155 at every wavelength tested. (GIF 25 kb)
41587_2003_BFnbt816_MOESM3_ESM.gif
Supplementary Table 1. Effects of a deletion in the leucine zipper of Fos on complementation between fragments of different fluorescent proteins fused to bFos and bJun. The relative fluorescence intensities of cells transfected with the expression vectors indicated were evaluated by fluorescence microscopy. For a calibration of the qualitative scale, please compare with the quantitative data in Fig. 3. (GIF 38 kb)
41587_2003_BFnbt816_MOESM4_ESM.gif
Supplementary Table 2. Selectivity of dimerization among the bZIP domains of bFos, bJun and bATF2. The fluorescence intensities of cells transfected with plasmids expressing the proteins indicated were measured using 436/470 nm (C) or 500/530 nm (Y) filters. The average fluorescence intensities and the average of the ratios for all cells examined are shown. (GIF 44 kb)
Rights and permissions
About this article
Cite this article
Hu, CD., Kerppola, T. Simultaneous visualization of multiple protein interactions in living cells using multicolor fluorescence complementation analysis. Nat Biotechnol 21, 539–545 (2003). https://doi.org/10.1038/nbt816
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt816
This article is cited by
-
An optimum study on the laser scanning confocal microscopy techniques for BiFC assay using plant protoplast
Botanical Studies (2024)
-
Combined proteomics and CRISPR‒Cas9 screens in PDX identify ADAM10 as essential for leukemia in vivo
Molecular Cancer (2023)
-
Dopamine-induced neural activity detection onto a cell-cultured plasmonic nanograting platform
Applied Physics A (2023)
-
Novel Bimolecular Fluorescence Complementation (BiFC) Assay for Visualization of the Protein–Protein Interactions and Cellular Protein Complex Localizations
Molecular Biotechnology (2023)
-
Monitoring the interactions between alpha-synuclein and Tau in vitro and in vivo using bimolecular fluorescence complementation
Scientific Reports (2022)