Abstract
To gain a better understanding of the critical role of mitochondria in cell function, we have compiled an extensive catalogue of the mitochondrial proteome using highly purified mitochondria from normal human heart tissue. Sucrose gradient centrifugation was employed to partially resolve protein complexes whose individual protein components were separated by one-dimensional PAGE. Total in-gel processing and subsequent detection by mass spectrometry and rigorous bioinformatic analysis yielded a total of 615 distinct protein identifications. All protein pI values, molecular weight ranges, and hydrophobicities were represented. The coverage of the known subunits of the oxidative phosphorylation machinery within the inner mitochondrial membrane was >90%. A significant proportion of identified proteins are involved in signaling, RNA, DNA, and protein synthesis, ion transport, and lipid metabolism. The biochemical roles of 19% of the identified proteins have not been defined. This database of proteins provides a comprehensive resource for the discovery of novel mitochondrial functions and pathways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chinnery, P.F., Howell, N., Andrews, R.M. & Turnbull, D.M. Clinical mitochondrial genetics. J. Med. Genet. 36, 425–36 (1999).
Terkeltaub, R., Johnson, K., Murphy, A. & Ghosh, S. The mitochondrion in osteoarthritis. Mitochondrion 1, 301–309 (2002).
Scharfe, C. et al. MITOP, the mitochondrial proteome database: 2000 update. Nucleic Acids Res. 28, 155–8 (2000).
Taylor, S.W., Fahy, E. & Ghosh, S.S. Global organellar proteomics. Trends Biotechnol. 21, 82–88 (2003).
Hanson, B.J., Schulenberg, B., Patton, W.F. & Capaldi, R.A. A novel subfractionation approach for mitochondrial proteins: a three-dimensional mitochondrial proteome map. Electrophoresis 22, 950–959 (2001).
Taylor, S.W. et al. An alternative strategy to determine the mitochondrial proteome using sucrose gradient fractionation and 1D PAGE on highly purified human heart mitochondria. J. Proteome Res. 1, 451–458 (2002).
Rappsilber, J., Ryder, U., Lamond, A.I. & Mann, M. Large-scale proteomic analysis of the human spliceosome. Genome Res. 12, 1231–1245 (2002).
Schwartz, R., Ting, C.S. & King, J. Whole proteome pI values correlate with subcellular localizations of proteins for organisms within the three domains of life. Genome Res. 11, 703–709 (2001).
Kyte, J. & Doolittle, R.F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982).
Sonnhammer, E.L., Eddy, S.R. & Durbin, R. Pfam: a comprehensive database of protein domain families based on seed alignments. Proteins 28, 405–420 (1997).
Fearnley, I.M. et al. GRIM-19, a cell death regulatory gene product, is a subunit of bovine mitochondrial NADH:ubiquinone oxidoreductase (complex I). J. Biol. Chem. 276, 38345–38348 (2001).
Carroll, J., Shannon, R.J., Fearnley, I.M., Walker, J.E. & Hirst, J. Definition of the nuclear encoded protein composition of bovine heart mitochondrial complex I: identification of two new subunits. J. Biol. Chem. 277, 50311–50317 (2002).
Yaffe, M.B. How do 14-3-3 proteins work?—Gatekeeper phosphorylation and the molecular anvil hypothesis. FEBS Lett. 513, 53–57 (2002).
Fountoulakis, M., Berndt, P., Langen, H. & Suter, L. The rat liver mitochondrial proteins. Electrophoresis 23, 311–328 (2002).
Lopez, M.F. et al. High-throughput profiling of the mitochondrial proteome using affinity fractionation and automation. Electrophoresis 21, 3427–3440 (2000).
Rabilloud, T. et al. Two-dimensional electrophoresis of human placental mitochondria and protein identification by mass spectrometry: toward a human mitochondrial proteome. Electrophoresis 19, 1006–1014 (1998).
Scheffler, N.K. et al. Two-dimensional electrophoresis and mass spectrometric identification of mitochondrial proteins from an SH-SY5Y neuroblastoma cell line. Mitochondrion 1, 161–179 (2001).
Palmieri, L. et al. Citrin and aralar1 are Ca2+-stimulated aspartate/glutamate transporters in mitochondria. Embo. J. 20, 5060–5069 (2001).
Simpson, P.B. & Russell, J.T. Role of sarcoplasmic/endoplasmic-reticulum Ca2+-ATPases in mediating Ca2+ waves and local Ca2+-release microdomains in cultured glia. Biochem. J. 325, 239–247 (1997).
Koc, E.C. et al. The large subunit of the mammalian mitochondrial ribosome. Analysis of the complement of ribosomal proteins present. J. Biol. Chem. 276, 43958–43969 (2001).
Suzuki, T. et al. Structural compensation for the deficit of rRNA with proteins in the mammalian mitochondrial ribosome. Systematic analysis of protein components of the large ribosomal subunit from mammalian mitochondria. J. Biol. Chem. 276, 21724–21736 (2001).
Cavdar Koc, E., Burkhart, W., Blackburn, K., Moseley, A. & Spremulli, L.L. The small subunit of the mammalian mitochondrial ribosome. Identification of the full complement of ribosomal proteins present. J. Biol. Chem. 276, 19363–19374 (2001).
Suzuki, T. et al. Proteomic analysis of the mammalian mitochondrial ribosome. Identification of protein components in the 28 S small subunit. J. Biol. Chem. 276, 33181–33195 (2001).
Grandier-Vazeille, X. et al. Yeast mitochondrial dehydrogenases are associated in a supramolecular complex. Biochemistry 40, 9758–9769 (2001).
Bardel, J. et al. A survey of the plant mitochondrial proteome in relation to development. Proteomics 2, 880–898 (2002).
Patterson, S.D. et al. Mass spectrometric identification of proteins released from mitochondria undergoing permeability transition. Cell Death. Differ. 7, 137–44 (2000).
Rabilloud, T. et al. The mitochondrial antioxidant defence system and its response to oxidative stress. Proteomics 1, 1105–1110 (2001).
Tanveer, A. et al. Involvement of cyclophilin D in the activation of a mitochondrial pore by Ca2+ and oxidant stress. Eur. J. Biochem. 238, 166–172 (1996).
Paschen, S.A. & Neupert, W. Protein import into mitochondria. IUBMB Life 52, 101–112 (2001).
Mori, M. & Terada, K. Mitochondrial protein import in animals. Biochim. Biophys. Acta 1403, 12–27 (1998).
Meisinger, C. et al. Protein import channel of the outer mitochondrial membrane: a highly stable Tom40-Tom22 core structure differentially interacts with preproteins, small tom proteins, and import receptors. Mol.Cell Biol. 21, 2337–2348 (2001).
Abdul, K.M. et al. Functional analysis of human metaxin in mitochondrial protein import in cultured cells and its relationship with the Tom complex. Biochem. Biophys. Res. Commun. 276, 1028–1034 (2000).
Hartmann, C.M., Gehring, H. & Christen, P. The mature form of imported mitochondrial proteins undergoes conformational changes upon binding to isolated mitochondria. Eur. J. Biochem. 218, 905–10 (1993).
Parry, D.M. & Pedersen, P.L. Intracellular localization of rat kidney hexokinase. evidence for an association with low density mitochondria. J. Biol. Chem. 259, 8917–8923 (1984).
Pastorino, J.G., Shulga, N. & Hoek, J.B. Mitochondrial binding of hexokinase II inhibits Bax-induced cytochrome c release and apoptosis. J. Biol. Chem. 277, 7610–7618 (2002).
Heggeness, M.H., Simon, M. & Singer, S.J. Association of mitochondria with microtubules in cultured cells. Proc. Natl. Acad. Sci. USA 75, 3863–3866 (1978).
Carré, M. et al. Tubulin is an inherent component of mitochondrial membranes that interacts with the voltage-dependent anion channel. J. Biol. Chem. 277, 33664–33669 (2002).
Dzeja, P.P., Bortolon, R., Perez-Terzic, C., Holmuhamedov, E.L. & Terzic, A. Energetic communication between mitochondria and nucleus directed by catalyzed phosphotransfer. Proc. Natl. Acad. Sci. USA 99, 10156–10161 (2002).
Pflieger, D. et al. Systematic identification of mitochondrial proteins by LC-MS/MS. Anal. Chem. 74, 2400–2406 (2002).
Murray, J., Gilkerson, R. & Capaldi, R. Quantitative proteomics: the copy number of pyruvate dehydrogenase is more than 102-fold lower than that of complex III in human mitochondria. FEBS Lett. 529, 173 (2002).
Model, K. et al. Protein translocase of the outer mitochondrial membrane: role of import receptors in the structural organization of the TOM complex. J. Mol. Biol. 316, 657–666 (2002).
Kumar, A. et al. Subcellular localization of the yeast proteome. Genes Dev. 16, 707–719 (2002).
Pfeffer, S.R. Rab GTPases: specifying and deciphering organelle identity and function. Trends Cell Biol. 11, 487–491 (2001).
Endele, S., Fuhry, M., Pak, S.J., Zabel, B.U. & Winterpacht, A. LETM1, a novel gene encoding a putative EF-hand Ca2+-binding protein, flanks the Wolf-Hirschhorn syndrome (WHS) critical region and is deleted in most WHS patients. Genomics 60, 218–225 (1999).
Caggese, C. et al. Identification of nuclear genes encoding mitochondrial proteins: isolation of a collection of D. melanogaster cDNAs homologous to sequences in the Human Gene Index database. Mol. Gen. Genet. 261, 64–70 (1999).
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002).
Storrie, B. & Madden, E.A. Isolation of subcellular organelles. Methods Enzymol. 182, 203–225 (1990).
Aggeler, R.J. et al. A functionally-active human F1F0 ATPase can be purified by immunocapture from heart tissue and fibroblast cell lines: subunit structure and activity studies. J. Biol. Chem. 277, 33906–33912 (2002).
Clauser, K.R., Baker, P. & Burlingame, A.L. Role of accurate mass measurement (+/−10 ppm) in protein identification strategies employing MS or MS/MS and database searching. Anal. Chem. 71, 2871–2882 (1999).
Ducret, A., Van Oostveen, I., Eng, J.K., Yates, J.R., 3rd & Aebersold, R. High throughput protein characterization by automated reverse-phase chromatography/electrospray tandem mass spectrometry. Protein Sci. 7, 706–719 (1998).
Field, H.I., Fenyo, D. & Beavis, R.C. RADARS, a bioinformatics solution that automates proteome mass spectral analysis, optimises protein identification, and archives data in a relational database. Proteomics 2, 36–47 (2002).
Acknowledgements
The authors thank Neil Howell and Christen Anderson for helpful comments, and Paul Haynes and Ross Hoffman for technical advice.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
MitoKor is a for-profit biotechnology company that has focused on using the mitochondrion as a platform for drug discrovery. Some proteins disclosed in the article are potential molecular drug targets for which patent applications have been, or will be, filed.
Supplementary information
Supplementary Figure 1.
Metrizamide gradient purified human heart mitochondria are highly depleted of plasma membrane, cytosol and organellar markers. (PDF 997 kb)
The purity of metrizamide purified mitochondria (lanes 3) compared to the initial human heart homogenate (lanes 1) and differential centrifugation purified mitochondria (lanes 2) was assessed. Equivalent loads of 25 μg (Panels A. B. and C.) or 90 μg (Panel D.) of each sample were separated by 4-12% NuPAGE. Western analysis was carried out with antibodies reactive toward proteins from various cellular locations.
Panel A shows Coomassie staining of protein in each fraction. Panel B. depicts the enrichment of components of the electron transport chain. Complex V was detected with Molecular Probes antibody cat.# A-21350 to ATPase alpha. Complex III was detected with Molecular Probes antibody cat.# A-21362 to Core I. Complex I was detected with Molecular Probes antibody cat.# A -21344 to NDUFA9. Complex IV was detected with Molecular Probes antibody cat.# A-21348 specific for Cox4. Further analysis of mitochondria specific markers is shown in Panel C. HSP 60, a matrix protein, was visualized with an antibody from Stressgen cat.# SPA-806. VDAC, a marker of the outer mitochondrial membrane, was detected with CalBioChem antibody cat.# 529538. Rab-11, after LC/MS/MS identification described in this paper, was confirmed to localize specifically to mitochondria using BD Transduction Labs antibody cat.#610656. These Western analyses demonstrate a good enrichment and integrity of mitochondria following metrizamide purification.
Reduced reactivity was detected in the DC mitochondria (lanes 2) and metrizamide mitochondria (lanes 3) by varous protein markers for nonmitochondrial proteomes, shown in Panel D. Dynamin II (cytosol/plasma membane), detected with BD Transduction Labs antibody cat.#D27120, Grp 95 and Grp 76 which contain the KDEL epitope (endoplasmic reticulum), detected with Stressgen antibody cat.#SPA-827, LAMP-1 (lysosome), detected with Santa Cruz Biotechnology antibody cat.#sc -17768, and alpha actin (cytosol), detected with Sigma antibody cat.#A2172 are depicted. Filters shown in Panel D were scanned on a Fluor-S Max Imaging System (BioRad) and quantitated using QuantityOne software. By this analysis, dynamin II was depleted by 95%, LAMP-1 by 98%, a-actin by 99%, Grp95 by 97%, and Grp76 by 95% in the metrizamide purified mitochondria relative to the initial human heart homogenate.
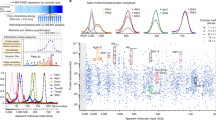
Supplementary Figure 2.
Plots of gel slice vs average molecular weight value of SEQUEST hits for sucrose density gradient fractions and pellet. (PDF 23 kb)
Supplementary Figure 3.
Distribution of the five complexes of the oxidative phosphorylation machinery within the sucrose gradient plotted as the number of SEQUEST matches (according to the threshold criteria in the Supplementary Information) to tryptic peptides of each of their subunits. Compare with the plot of Western intensities in Taylor et al. J. Proteome Research 1, 451–458 (2002). The distribution of selected proteins discussed in the text throughout the gradient is shown. Abbreviations: PHB's, prohibitins; SSBP, single stranded mitochondrial DNA binding protein; mtRBP's, mitochondrial ribosomal DNA binding proteins; cMDH, cytosolic malate dehydrogenase; GK, glycerol kinase; ICDH, isocitrate dehydrogenase; MXN, metaxin 2; mtMDH, mitochondrial MDH; CS, citrate synthase; AC, aconitase; FH, fumarate hydratase; LDH, lactate dehydrogenase; ADH, aldehyde dehydrogenase. (PDF 13 kb)
Rights and permissions
About this article
Cite this article
Taylor, S., Fahy, E., Zhang, B. et al. Characterization of the human heart mitochondrial proteome. Nat Biotechnol 21, 281–286 (2003). https://doi.org/10.1038/nbt793
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt793
This article is cited by
-
Mitochondrial proteome research: the road ahead
Nature Reviews Molecular Cell Biology (2024)
-
Mitochondrial heterogeneity in diseases
Signal Transduction and Targeted Therapy (2023)
-
FAM210A is essential for cold-induced mitochondrial remodeling in brown adipocytes
Nature Communications (2023)
-
Tracing the lactate shuttle to the mitochondrial reticulum
Experimental & Molecular Medicine (2022)
-
Respiratory substrate preferences in mitochondria isolated from different tissues of three fish species
Fish Physiology and Biochemistry (2022)