Abstract
Trichoderma reesei is the main industrial source of cellulases and hemicellulases used to depolymerize biomass to simple sugars that are converted to chemical intermediates and biofuels, such as ethanol. We assembled 89 scaffolds (sets of ordered and oriented contigs) to generate 34 Mbp of nearly contiguous T. reesei genome sequence comprising 9,129 predicted gene models. Unexpectedly, considering the industrial utility and effectiveness of the carbohydrate-active enzymes of T. reesei, its genome encodes fewer cellulases and hemicellulases than any other sequenced fungus able to hydrolyze plant cell wall polysaccharides. Many T. reesei genes encoding carbohydrate-active enzymes are distributed nonrandomly in clusters that lie between regions of synteny with other Sordariomycetes. Numerous genes encoding biosynthetic pathways for secondary metabolites may promote survival of T. reesei in its competitive soil habitat, but genome analysis provided little mechanistic insight into its extraordinary capacity for protein secretion. Our analysis, coupled with the genome sequence data, provides a roadmap for constructing enhanced T. reesei strains for industrial applications such as biofuel production.
Similar content being viewed by others
Main
Trichoderma reesei (teleomorph Hypocrea jecorina) is a mesophilic soft-rot ascomycete fungus that is widely used in industry as a source of cellulases and hemicellulases for the hydrolysis of plant cell wall polysaccharides. For many years after its discovery during World War II1, T. reesei was believed to reproduce asexually. However, although it was subsequently shown to be the anamorph of the pantropical ascomycete Hypocrea jecorina2, the organism remains most widely recognized by its former name. It has enjoyed a long history of safe use for industrial enzyme production3 and as an important model system for studying lignocellulose degradation.
Lignocellulosic biomass from agricultural crop residues, grasses, wood and municipal solid waste represents an abundant renewable resource that is becoming increasingly important as a future source of biofuels. Although replacement of gasoline with cellulosic ethanol may substantially reduce greenhouse gases in the atmosphere and decrease global warming4, the high cost of hydrolyzing biomass polysaccharides to fermentable sugars remains a major obstacle that must be overcome before cellulosic ethanol can be effectively commercialized. As the costs of cellulases and hemicellulases contribute substantially to the price of bioethanol, much cheaper sources of these enzymes are needed5. Consequently, new studies aimed at understanding and improving cellulase efficiency and productivity are at the forefront of biomass research.
As T. reesei represents a paradigm for the production of enzymes that hydrolyze biomass polysaccharides, intensive research efforts and considerable government funding have been applied toward developing better industrial strains for producing bioethanol and a range of key biochemical building blocks, such as 1,4-dicarboxy acids (succinate, malate, fumarate), 3-hydroxypropionic acid, aspartic acid, glucaric acid, glutamic acid, itaconic acid, levulinic acid, glycerol, sorbitol, xylitol/arabinitol and hydroxybutyrolactone, that are currently derived from nonrenewable petroleum-based resources5. Although genetic engineering techniques, gene knockout protocols and DNA-mediated transformation systems3 have improved industrial enzyme–producing T. reesei strains, the need to better understand this fungus and expand its extraordinary biotechnological potential has given impetus to the quest to sequence its genome. To this end, we have now analyzed the T. reesei genome with particular emphasis on its potential contributions to fuel biotechnology and other industrial applications.
Results
Features of the T. reesei genome
The genome of T. reesei was shotgun sequenced6 to approximately ninefold coverage from three libraries (insert sizes of 3 kb, 8 kb and 40 kb) totaling 433,863 reads (Supplementary Table 1 online). These data, in addition to more than 6,000 BAC-end sequences7, yielded a high-quality draft assembly using the Department of Energy Joint Genome Institute (JGI) shotgun assembler Jazz8. 6,329 finishing reads were created with custom primers from the 3-kb and 8-kb libraries to close a majority of the gaps, and the Phred/Phrap/Consed software package was then used to assemble 89 scaffolds and 97 contigs totaling ∼34 Mb. This is only 2.9% larger than the size estimated from several karyotyping studies9,10,11 and agrees with the genome size estimated by physical means. All of the genetic markers that were used in the three studies were recovered, as were all protein and RNA sequences in the current release of GenBank (version 161.0). We are therefore confident that the T. reesei genome sequence reported here represents more than 99% of the genome.
We detected repetitive sequences with similarity to class I and II transposable elements (Supplementary Note 1 online), although all contained multiple stop codons. The apparent absence of active transposons may be explained by the presence of active defense mechanisms, such as repeat-induced point mutations. These transposable elements totaled less than 1% of the finished genome—among the lowest frequencies reported for a fungal genome. A repeated hexanucleotide sequence, TTAGGG, that is identical to the telomeric repeat of Neurospora crassa was found at the ends of seven scaffolds in the T. reesei genome assembly (Supplementary Note 1).
Gene modeling was performed using a combination of homology and ab initio methods, selecting a single gene model for each locus (see Methods). This yielded 9,129 gene models (Table 1). This total is relatively close to the number of gene models in N. crassa12, but is roughly 2,500 fewer than the number of predicted genes in Fusarium graminearum13 (anamorph, Gibberella zeae)—a surprising difference, given that F. graminearum and T. reesei share the most recent common ancestor among the genomes listed in Table 1 (ref. 14). The average gene length in T. reesei is 1,793 base pairs (bp), with 3.1 exons per gene (average exon length, 508 bp; average intron length, 120 bp). All data, manual curations and sequence files are available for viewing and download in the interactive JGI Genome Portal (http://www.jgi.doe.gov/Tre esei).
Conserved synteny in T. reesei
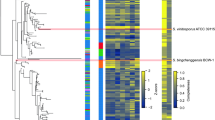
To gain insight into the effect of the environment on genome evolution, we constructed a comparative map by calculating syntenic regions shared by T. reesei, F. graminearum13 and N. crassa12 (Supplementary Table 2 online). As noted in other studies15, this map (Fig. 1) illustrates segments in which the gene order has changed since the divergence of these species, resulting in large gaps between syntenic blocks. In many cases, these gaps are conserved between T. reesei and the other Sordariomycetes, suggesting that they are prone to frequent insertions, duplications or chromosomal breaks. Regions without synteny to other genomes often contain genes that are important for the adaptation of the organism15,16,17. Another noteworthy feature of the comparative map (Fig. 1) is the number of chromosomal rearrangements that have occurred since the divergence of the three organisms, clearly illustrating the highly dynamic nature of the genome. The difference in the level of syntenic coverage relative to the two other Sordariomycetes (Fig. 1 and Supplementary Table 2a) is consistent with the view that F. graminearum and T. reesei share a more recent common ancestor18.
The eight scaffolds shown comprise ∼50% of the T. reesei genome. Small blocks internal to the T. reesei genome represent sections of the T. reesei genome that share synteny with the F. graminearum and N. crassa genomes. The calculation of syntenic blocks is described in Methods. Syntenic block coloring is by chromosome and linkage group for F. graminearum and N. crassa, respectively. Blue boxes represent regions of the T. reesei genome that have an increased density of genes encoding carbohydrate active enzymes (CAZymes), as described in the text. Overall syntenic comparisons and detailed descriptions of CAZyme blocks are presented in Supplementary Tables 2 and 7, respectively. Chr, chromosome; LG, linkage group.
To investigate the factors that determine gene synteny, we compared the general features of genes in syntenic blocks with those found in gaps (nonsyntenic regions) (Supplementary Table 2b). Although for many of these metrics there is little difference between the regions, there is a large difference in mean exon size between F. graminearum syntenic and nonsyntenic regions (89 nucleotides, P value 2.2 × 10−16). Analysis of InterPro19 domain content between the two groups of genes (Supplementary Table 2c) reveals a noticeable difference in the number of genes containing the domain IPR001680 (G-protein β WD-40 repeat). As genes with this domain seem to have unusually large exons, they probably contribute substantially to the shift in mean exon size. Another interesting finding from the InterPro comparison in Supplementary Table 2c is that the InterPro domain IPR000254 (cellulose-binding region, fungal) is found only in genes that lie in syntenic gaps and is not found in T. reesei genes that are in syntenic blocks shared with either F. graminearum or N. crassa.
Protein domains in T. reesei
We compared the protein domains encoded by the T. reesei genome to those of 13 fungal genomes using InterProScan19 to find regions in proteins with known functions. Compared to those of sequenced species within the Pezizomycotina, the proteome of T. reesei shows underrepresentation of many proteins with known functions (Supplementary Table 3 online). In particular, as outlined in the next section, T. reesei differs in its content of proteins related to plant biomass degradation. Consistent with its natural role as a necrophyte, T. reesei lacks several protein families related to infecting and degrading living plant tissue, such as pectate lyases and pectin esterases. Moreover, the failure to detect tannase and feruloyl esterase family members suggests that T. reesei is apparently handicapped in the degradation of hemicellulose.
Carbohydrate-active enzymes in T. reesei and other fungi
Carbohydrate-active enzymes (CAZymes) are categorized into different classes and families in the CAZy database (http://www.cazy.org)20. CAZymes that cleave, build and rearrange oligo- and polysaccharides play a central role in the biology of fungi such as T. reesei and are key to optimizing biomass degradation by these species. Given the relative importance of this protein family to the biotechnology community, we performed a detailed examination of the CAZome of T. reesei and compared it with the corresponding gene subsets from 13 fungi for which genome sequences are available (Table 2).
Although one might expect that T. reesei—an efficient plant polysaccharide degrader and important model of the degradation system—would contain expansions of genes whose products are involved in digesting cell wall compounds, it has surprisingly few genes encoding glycoside hydrolases (GHs). With a total of 200 GH-encoding genes, it has fewer GHs than the phytopathogens Magnaporthe grisea and F. graminearum (Table 1). This figure is also slightly below the average number of GHs found in this lineage (211), though the difference does not exceed the standard deviation (s.d. = 32). Compared to other fungal lineages, the Sordariomycetes represent the second most GH-rich lineage, preceded only by the Eurotiomycetes, which average 265 GHs and have a more homogeneous GH distribution (s.d. = 19).
With 103 glycosyltransferases, T. reesei is close to the average among Sordariomycetes (96) (Table 2). This enzyme class shows less variability in Sordariomycetes than do GHs (s.d. = 15), as is also noticeable in the other phyla in our dataset. This trend is maintained for both intralineage and interlineage variability, suggesting both that glycosyltransferases possess basal intracellular activities and that variations in composition may reflect species drift rather than environmental pressure.
The enzymes involved in plant polysaccharide depolymerization frequently carry a carbohydrate-binding module (CBM) appended to the catalytic domain. Unexpectedly, the T. reesei genome has the smallest number of CBM-containing proteins among the Sordariomycetes in our dataset (Table 2). However, it should be noted that the fact that the Sordariomycetes have the highest number of CBMs in this dataset can essentially be attributed to the significant enrichment of CBMs in the phytopathogens F. graminearum and M. grisea. Similarly, T. reesei has the lowest number (16) of carbohydrate esterases among the Sordariomycetes we analyzed. The difference from the average among Sordariomycetes (32) is approximately equal to the standard deviation (s.d. = 15).
The Sordariomycetes, including T. reesei, show a relative paucity of polysaccharide lyase genes—a category that typically contains 3–4 genes—with the exception of F. graminearum, which has an expansion of 20 genes. Such a high number of polysaccharide lyases is found only in the Eurotiomycetes, which have an average of 18 polysaccharide lyases. No polysaccharide lyases are found in unicellular Ascomycetes. In conclusion, the T. reesei genome encodes a number of CAZymes that is slightly below the average found among Sordariomycetes. Detailed statistical analyses are presented in Supplementary Note 2 and Supplementary Table 4 online.
Unexpectedly, a thorough inspection of the T. reesei genome revealed only seven genes encoding well-known cellulases (endoglucanases and cellobiohydrolases), giving T. reesei the fewest cellulases of all the fungi listed in Table 3 that are able to degrade plant cell walls. This trend is further amplified if one adds the family of GH61 proteins (Table 3). Hemicellulose comprises a diverse group of complex polysaccharides, and their complete degradation requires an arsenal of enzymes. With only 16 hemicellulase genes, T. reesei has the smallest set of hemicellulases among all fungi analyzed (Table 4). Similarly, T. reesei has the smallest set of enzymes for the breakdown of pectin among the plant cell wall–degrading fungi (Supplementary Table 5 online).
Protein secretion
T. reesei is an extraordinarily efficient producer of extracellular enzymes, with certain industrial strains producing 100 g of extracellular protein per liter.21. This apparently remarkable efficacy of the protein secretion machinery of T. reesei makes analysis of its genes encoding secretory pathway components of particular interest. Not surprisingly, homologs of proteins that function in the secretory pathway of Saccharomyces cerevisiae were found in the T. reesei genome. Although generally these are present as single-copy genes in T. reesei and show greater similarity to the yeast orthologs than to their mammalian counterparts, there are a few notable exceptions to this trend. T. reesei seems to have three proteins whose closest homolog in yeast is protein disulfide isomerase, Pdi1p. This could be connected to the fact that the major secreted cellulases of this fungus have many disulfide bonds22. The ER-associated protein degradation (ERAD) pathway of T. reesei seems to be more redundant than the secretory pathway in general, as two orthologs of the yeast DER1 and UFD1 genes are found. We found clear homologs of most of the other known ERAD components in T. reesei, despite an apparent lack of orthologs or little sequence similarity to yeast ERAD components in the Aspergillus niger genome23.
Orthologs of most S. cerevisiae proteins that are known to be involved in protein trafficking can be found as single copies in the T. reesei genome. Whereas yeast lacks counterparts of the mammalian GTPase proteins Rab2, Rab4 and Rab5 and Arf6 and Arf10—signaling proteins involved in membrane fusion or vesicle budding in diverse cellular locations—T. reesei and N. crassa seem to have orthologs of these proteins (Supplementary Table 6 online). The t-SNARE protein Sso1p of yeast, a receptor for the secretory vesicles on the plasma membrane, has two homologs in T. reesei, and a recent study indicates that the two Sso1 homologs have divergent functions24. Taken together, these findings suggest that the membrane trafficking system in T. reesei is more diverse than that in S. cerevisiae.
CAZyme gene clusters in T. reesei
Many of the T. reesei genes encoding CAZymes are nonrandomly distributed within the genome. In a previous study, nine known genes whose products are involved in cellulose and hemicellulose degradation were shown to be colocated in several areas of the genome7. We have extended this work to the location of all CAZymes in the genome and found that in total, 130 of the 316 (41%) CAZyme genes are found in 25 discrete regions ranging from 14 kb to 275 kb in length (roughly 2.4 Mb, or 7% of the genome) (Fig. 2 and Supplementary Table 7 online). These regions contain a density of CAZyme genes averaging fivefold greater than the expected density for randomly distributed genes. Based on the hypergeometric distribution (see Methods), we have calculated the P values of the clusters, which range from 0.015 to 1 × 10−4. Each region contains between two (as adjacent pairs) and ten CAZyme genes.
In all sections of the genome shown in this figure, the CAZyme genes are in gaps between syntenic blocks. (a) Region of scaffold 1 containing a gene encoding a candidate chitinase and acetyl xylan esterase (axe1), as well as cip1 (encoding a previously identified CBM-containing protein42) and cel61a (encoding endoglucanase IV). The polyketide synthase (PKS) genes in this region are Tr73621 and Tr73618. Tr73621 appears in a clade with the lovF gene (encoding lovostatin-producing protein) of Aspergillus terreus, Tr73618, as described in the text. (b) Region of scaffold 29 showing that cel7a (CBHI), swollenin and xylanase 4 are co-induced. The upstream orange arrows denote genes that show co-induction on cellulose or sophorose as well, but are unknown conserved hypotheticals. (c) This region contains bxl1 and cel3a (encoding β-xylosidase and β-glucosidase 1, respectively), which are co-induced. Region also contains two unknown genes (purple arrows) encoding predicted secretion signals that are also coexpressed along with bxl1 and cel3a. (d) A candidate xylanase gene and cip2, encoding the CBM-containing protein and identified in published work42, are both adjacent to an unknown gene that is also induced on cellulose/sophorose.
To gain insight into how such clusters arise, we analyzed the number of orthologs within the clusters. Ninety-five of the 130 (73%) CAZyme genes that are in clusters are in gaps of synteny. Of those 95 CAZyme genes, 69 (72%) have orthologs in F. graminearum. There are a mere 16 CAZyme orthologs that are in synteny with F. graminearum, indicating that gene movement is the major factor in the organization of the clusters, whereas gene duplications have a minor role. With respect to the nonorthologous CAZyme genes (the potential duplicates), all have homologs in almost all the fungal genomes sequenced to date. In addition, few CAZyme genes in the same cluster are from the same CAZyme family, with a few notable exceptions (see Supplementary Table 7); only ten genes in four different clusters are from the same subfamily, including a pair of GH3s and a triplet of GH3s. This suggests that the few paralogs that are colocated in the clusters indicate that gene relocation rather than duplication is responsible for the formation of the CAZyme clusters.
The profile of CAZyme genes found in the clusters suggests a specific biological role. Approximately 70% of the CAZyme genes in the clusters encode GHs. The finding that 24% of the glycosyltransferase genes and 46% of the GH genes in the genome are found in these regions indicates that the majority of the CAZyme genes in these clusters encode proteins involved in polysaccharide degradation (Supplementary Table 7). This is supported by the finding that many of the genes with products previously shown to be involved in plant cell wall degradation fall into the CAZyme-rich regions (Supplementary Table 8 online). Three of the four expansin-like genes in T. reesei, including the previously described swollenin gene25, are located in these clusters (Fig. 2). It is intriguing that the few glycosyltransferases whose genes are found in CAZyme clusters are largely mannosyltransferases, chitin synthases (four of nine in T. reesei), α-glycosyltransferases and β-glycosyltransferases—all enzymes that may be involved in synthesizing fungal cell walls26.
A portion of the data from two transcriptomics projects identifying T. reesei genes induced by sophorose27 and cellulose28 were mapped to the genome. Although not all of the clustered GH genes were coexpressed in the above studies, we found four examples in which adjacent or nearly adjacent genes were coexpressed (Fig. 2), giving further evidence for the biological importance of the CAZyme clustering. Notably, in these regions there is no syntenic signal with any of the other fungal genomes as shown in Figure 1, suggesting that these genes are reordered in T. reesei and that this organization is evolutionarily advantageous for the fungus.
Several of the regions of high CAZyme gene density also contain genes encoding proteins involved in secondary metabolism (Supplementary Table 7). Specifically, five of the 25 CAZyme clusters contain either a polyketide synthase (PKS) gene or a nonribosomal peptide synthase (NRPS) gene. In particular, we found two nonreducing PKS genes (scaffold_1:410,000–530,000 and scaffold_6:10,000–148,000, Fig. 2) that in our phylogenetic analysis (maximum likelihood performed with PHYML and RAXYML, data not shown) appear in a clade with previously undescribed PKS genes. In addition, the PKS gene in the region of scaffold_6:10,000–148,000 is fused with an NRPS gene that resides in a clade with NRPS genes encoding proteins involved in lovastatin and citrinin production (maximum likelihood performed with PHYML and RAXML, data not shown). Another intriguing finding is that T. reesei has retained most NRPS paralogs as compared to other Sordariomycetes analyzed thus far. Supplementary Table 9 online lists the NRPS and PKS genes found in the T. reesei genome.
Discussion
As compared with the fungi listed in Tables 2 3 4, T. reesei has the smallest repertoire of genes for cellulases, hemicellulases and pectinases—the three categories of enzymes involved in depolymerizing plant cell wall polysaccharides (Tables 3 and 4 and Supplementary Table 5, respectively). This is unexpected, as the cellulolytic enzyme machinery of T. reesei is efficient and represents the paradigm for the enzymatic breakdown of cellulose and hemicellulose. An inability to rationalize the diversity observed in the composition of cellulolytic enzymes among fungal proteomes underscores our poor understanding of plant cell wall degradation and suggests that there may be room for improving T. reesei strains by augmenting their inventory of CAZymes with genes from other sources. On the other hand, this limited enzyme set is sufficient to enable T. reesei to compete in nature with other fungi that degrade cellulose and hemicellulose. The degree to which its success is enhanced by an array of secondary metabolites is unknown. However, it is tempting to speculate that the clustering of GH genes (in some cases near genes encoding proteins involved in secondary metabolite production) has enabled T. reesei to control the expression of these genes more efficiently.
The T. reesei genome reveals that several enzyme families involved in polysaccharide degradation are reduced or absent. Of all the possible CAZyme genes involved in pectin degradation, only members of the GH28 family are found, and there is no expansion of this family that could compensate for the lack of other pectinolytic enzymes. This deficiency of pectinolytic enzymes is confounding when one compares T. reesei with other Sordariomycetes (Supplementary Table 2), but it is consistent with the poor growth of T. reesei on D-galacturonic acid and L-rhamnose29, two constituents of the pectin backbone. L-Arabinose and D-galactose, which make up the majority of the side chains in 'hairy regions' of pectin, are readily metabolized. Possibly the pectin backbone is depolymerized by other fungi and bacteria in the soil, where T. reesei exists primarily as a secondary colonizer. The absence of invertase (EC 3.2.1.26; family GH32) is also consistent with the fungus' role as a secondary colonizer, if sucrose is consumed rapidly by primary colonizers.
Previous studies indicate that, in both bacterial and eukaryotic genomes, the locations of genes are not necessarily random30. In fungi, there are examples of gene clusters encoding proteins that are involved in the production of secondary metabolites, including NRPS/PKS pathways, or in the oxidation of substrates, for example the cytochrome P450 genes in Phanerochaete chrysosporium31. In several Clostridium species, there is an intriguing parallel to the T. reesei CAZyme clusters in that the genes of the cellulosome complex encoding GH enzymes needed for cellulose and hemicellulose degradation are also clustered32. However, the distances between GH genes are much shorter than in T. reesei, aside from the cases shown in Figure 2. Thus, in Clostridium cellulosome gene clusters, as well as in the T. reesei CAZyme clusters, functional coupling of genes whose products are involved in the hydrolysis of cellulose and hemicellulose creates pressure to maintain their positions relative to one another. This is in agreement with the chemical complexity of plant cell wall polysaccharides, which requires a diverse mixture of enzymes for complete depolymerization. Given these observations, it is reasonable to conclude that the clustering of the CAZyme genes is favored by selection for the enhanced degradative efficiency and coordinated regulation that a colocalization strategy may offer.
The concentration of CAZyme genes (primarily GH genes) in syntenic gaps with F. graminearum and N. crassa further supports the notion that selective pressure can maintain the clustering of genes encoding proteins involved in biomass degradation. In comparison, previous studies15,16,17 indicate that syntenic gaps in other genomes are enriched in genes that are important for species-specific attributes. Although it is possible that duplications may play a role in the loss of synteny, the CAZyme clusters in T. reesei show little evidence of expansion in comparison with the other fungi analyzed. Indeed, there are few clusters that contain appreciable numbers of genes from the same subfamily (Supplementary Table 7), indicating that recent duplication has not played an important role in their creation. It is therefore likely that the majority of the breaks in synteny at which CAZyme genes are clustered arise from movement of CAZyme genes into these regions, followed by pressure to fix the genomic rearrangements in the population.
The reduction in duplicated genes in T. reesei could be attributed to the effects of repeat-induced point mutation, similar to the limitation seen in N. crassa (Supplementary Note 1). As mentioned previously, a repeat-induced point mutation–like pattern of mutation is observed in the transposons of T. reesei, albeit at a lower density of mutations than in N. crassa. This could explain why the genome sizes of T. reesei and N. crassa are similar and why both genomes contain few intact repeats. It may also account for the lack of gene family expansion in GHs, forcing the organism to favor gene translocations to facilitate adaptation to the environment.
The biased placement of several secondary metabolism genes near CAZyme clusters presents the intriguing possibility that they may enable T. reesei to fend off competition for nutrients. In addition, the number of conserved PKS and NRPS genes in T. reesei suggests that the organism's survival requires an arsenal of antimicrobial secondary metabolites, particularly in light of the limited repertoire of CAZymes. The only GH family that contains any appreciable enrichment is that of the chitinase genes in family GH18 (Table 3), nearly half of which can be found in clusters. Other members of the genus Trichoderma (such as T. harzianum and T. atroviride) are capable of mycoparasitism, and both chitinases and secondary metabolites could be important in attacking other fungi33.
Although the organization of GH genes may contribute to the ability of T. reesei to efficiently degrade plant material, the lack of key enzyme activities clearly suggests opportunities for industry to generate improved enzyme cocktails that may be used for the conversion of plant biomass to fermentable sugars. As complete hydrolysis of cellulosic and hemicellulosic substrates requires multiple enzymes acting synergistically, development of superior enzyme blends is likely to occur through genetic engineering of suitable industrial strains. The capacity for secreting copious amounts of extracellular enzymes, the availability of genetic tools and the straightforward, inexpensive fermentation of T. reesei make it an ideal candidate for producing enzymes useful for the conversion of biomass feedstocks such as corn stover, cereal straw and switch grass34 to fuel ethanol and manufacturing chemicals that are currently derived from nonrenewable resources. Production of these enzymes at economically viable levels will require an increased understanding of the dynamics of cell growth and enzyme production. Mathematical and kinetic models are being developed to optimize these processes35, and the availability of a complete genome sequence will provide a blueprint to improve the models and to empower strain improvement strategies for creating superior enzyme mixtures from a single highly productive strain.
Methods
Automated annotation.
In addition to using previously published methods36, to predict genes in the T. reesei genome we used an ab initio gene predictor, Fgenesh37, specifically trained for this genome, and two homology-based gene predictors, Fgenesh+ (http://www.softberry.com) and Genewise38. All three methods predict only coding sequence regions in genes, which we then corrected and, where possible, extended into putatively full-length genes using 42,916 T. reesei expressed sequence tags (ESTs). Finally, using a heuristic approach implemented in the JGI pipeline, we combined all predicted gene models to produce a nonredundant set of genes, in which a single best gene model per locus was selected on the basis of sequence similarity to known proteins and support from available ESTs. This representative set included 9,129 genes and was subject to manual curation and genome analysis as described here.
The majority (82%) of predicted genes contain multiple exons, with an average of 3.1 exons per gene. Average gene density, similar between most of the larger scaffolds, is 3.7 kb per gene. Average gene, transcript and protein lengths are 1.8 kb, 1.6 kb and 492 amino acids, respectively (Table 1). In total, 7,887 (86%) gene models were predicted to be complete. There are 42,916 ESTs that support 46.1% of the predicted genes. Approximately 94% of the predicted proteins show sequence similarity to other proteins, primarily from fungi. A total of 2,164 manually curated genes from version 1.2 of the T. reesei Genome Portal were mapped forward to version 2.0.
We annotated and classified genes according to Gene Ontology (GO)39, eukaryotic orthologous groups (KOGs)40 and KEGG metabolic pathways41. We assigned GO terms to 4,977 (54.5%) of the predicted T. reesei proteins, including 3,547, 1,913 and 4,651 proteins with molecular function, cellular component and biological process, respectively. We also assigned 5,420 (59.4%) proteins to KOG clusters. We assigned 751 distinct EC numbers to 2,264 (25%) proteins mapped to KEGG pathways.
Manual curation.
We manually curated gene function assignments for 2,164 gene models using an interactive website (http://genome.jgi.doe.gov/Trire2). To assign confidence to these functional calls as well as to standardize the nomenclature methods, we used a qualifier system based on the homolog for which a functional assignment was made, which is used throughout the paper. This nomenclature was based on the following naming convention. A three-letter code was assigned to a gene only when the gene had experimental evidence describing the function of the gene product in T. reesei. If an experiment had not been performed in T. reesei, the tag tre<gene_id> was used—for example, tre167435. The definition line, or “def_line,” was assigned on the basis of sequence similarity to proteins in other organisms and InterPro domains. If the best sequence similarity was above 80% identity and 80% coverage (calculated as alignment length divided by predicted protein length) with respect to a protein for which there was published experimental evidence of its function, no def_line qualifier was used. If the sequence similarity was above 70% identity and 70% coverage, yet lower than 80% identity and 80% coverage, with respect to a protein with published evidence of function, the def_line qualifier “Candidate” was used (for example, candidate α,α-trehalase). If the sequence similarity was above 50% identity and 50% coverage with respect to a protein with published evidence of function, the def_line qualifier “Related to” was used, as in “Related to α-fucosidase.” Below this last threshold, all def_lines are tagged with the qualifier “Hypothetical.” Unknown and hypothetical proteins with hits to only other unknown hypothetical proteins are assigned the def_line “Conserved Hypothetical.”
Calculation of syntenic blocks.
The areas of relationship known as syntenic (meaning 'same ribbon') regions or syntenic blocks are anchored with orthologs (calculated as mutual best hits or bidirectional best hits) between the two genomes in question, and are built by controlling for the minimum number of genes, minimum density and maximum gap (containing genes not from the same genome area) as compared with randomized data, as described in published work42. A version of the algorithm was programmed in PERL and runs in less than 1 min on an AMD Opteron dual-CPU machine with 6 gigabytes of RAM. This savings in time is largely due to the requirement that the orthologs be precalculated from BLAST results (minimum expectation value 10−5, 40% coverage required). The algorithm code is available from the authors upon request.
Although this technique may produce artificial breaks, it highlights regions that are dynamic and have recently picked up a large number of insertions or duplications. From the analysis shown in Supplementary Table 2, N. crassa shares 5,624 mutual best hit (MBH) genes with T. reesei (62% of T. reesei genes), and F. graminearum shares 6,580 MBH genes with T. reesei (72% of T. reesei genes) that have maintained their general location since diverging from their most recent common ancestor.
Protein domains.
The proteomes used in this study included Aspergillus fumigatus, Aspergillus nidulans, F. graminearum (not yet published), T. reesei, M. grisea, N. crassa, Ashbya gossypii, Candida albicans, Candida glabrata, Debaryomyces hansenii, Kluyveromyces lactis, Yarrowia lipolytica and S. cerevisiae. The number of genes found to have a certain InterPro entry was counted. To obtain robust results that would not be clouded by differences in sequencing coverage, assembly or version of the genomes used, we searched for over-represented InterPro entries by selecting those that had at least twice as many corresponding genes in T. reesei than in any other euascomycete, and vice versa for under-represented entries. Differences in InterPro entry counts can be due to the actual presence or absence of a domain or to mutations in the domain's sequence that renders it unrecognizable to InterProScan. To classify the results accordingly and verify them, we carried out BLAST searches and studied alignments of homologous genes.
Detection of carbohydrate-active enzymes in fungal proteomes.
The search for carbohydrate-active modules (GHs, glycosyltransferases, polysaccharide lyases and carbohydrate esterases) and their associated carbohydrate-binding modules (CBMs) in T. reesei was performed exactly as for the daily updates of the Carbohydrate-Active enZYme (CAZy) database (http://afmb.cnrs-mrs.fr/CAZY/). Briefly, the sequences of the proteins in CAZy were cut into their constitutive modules (catalytic modules, CBMs and other noncatalytic modules or domains of unknown function). The resulting fragments were assembled and formatted as a sequence library for BLAST searches. Accordingly, each protein model from T. reesei (and other fungal proteomes) was searched via BLAST against the library of approximately 100,000 individual modules using a database size parameter identical to that of the NCBI nonredundant database. All models that gave an expectation value lower than 0.1 were automatically kept and manually analyzed. Manual analysis involved examination of the alignment of the model with the various members of each family (whether of catalytic or noncatalytic modules), with a search of the conserved signatures and motifs characteristic of each family. The presence of the catalytic machinery was verified for borderline cases whenever known in the family. The models that showed the usual features that would lead to their inclusion in the CAZy database were kept for annotation and classified in the appropriate class and family.
Functional annotation of protein models corresponding to carbohydrate-active enzymes.
The analysis of the sequence-based families of GHs and glycosyltransferases shows that those families rarely coincide with a single substrate (or product) specificity43. As a consequence, many of these families group together enzymes that have different EC numbers. Our annotation strategy aims at producing (as much as possible) annotations that will 'age' well—for example, that are designed to survive experimental validation while avoiding overinterpretation. For instance, in a family that contains β-mannosidases, β-galactosidases and β-glucuronidases, all enzymes hydrolyze equatorially oriented glycosidic bonds. A strong similarity to β-galactosidases allows annotation as “candidate β-galactosidase,” but if similarity is not sufficient for a safe prediction of substrate specificity, the best possible annotation is “candidate β-glycosidase.” Each protein model kept from the modular annotation step was thus annotated using that scheme. The proteins were analyzed via BLAST again against the manually curated CAZy database, and we assigned a functional annotation according to the relevance of the BLAST matches. Only when the enzyme of the species itself has been experimentally characterized was the protein given an EC number. All uncharacterized protein models were thus at best “candidates” or “related to” or “distantly related to” their characterized match. Because the threshold of similarity that correlates with a change of substrate specificity is extremely variable from one family to another, the criteria were tightened or loosened appropriately for several protein families.
Fungal CAZome comparisons.
We used several statistical analyses to identify the significant features in the comparison of the sets of carbohydrate-active enzymes encoded by 13 fungal genomes, taking into account both taxonomic and CAZyme families variability. Basically, the approach consisted in applying a χ2 independence test and other statistical analyses to identify the most unexpected points for a given CAZyme family per species according to the general distribution.
For each class of CAZymes, the statistical test first required placing the data in a table of k columns (representing the different families) and l rows (representing the different species). The Aij value will represent the number of CAZymes from family i and species j. We next calculate the values of:

then,

The last value allows the rejection of the χ2 independence hypothesis, and the Aij that contributes the most to the total sum represents the points (families) that are the most significantly different for a given species.
Gene cluster identification.
The gene families in question were collected by visual inspection using the JGI Genome Portal for the T. reesei genome. A cluster is defined as a region containing a statistically higher proportion of a particular gene family and must begin and end with a gene from the family in question. We then calculated the probability that a proportion in the cluster of the particular gene family is higher than the current one using the hypergeometric distribution (expressed as a P value). In gathering such clusters, it is possible to take a smaller section and get a higher P value; however, our goal was to take the longest reasonable cluster that had a P value <0.05 (outside the 95% confidence interval). In the CAZyme clusters presented here, the mean P value is 3.9 × 10−3, and only 4 of the 25 clusters has a P value that is >0.01, outside the 99% interval, but still <0.05.
Accession number.
The T. reesei nucleotide sequence and annotation data have been deposited in GenBank under accession number AAIL00000000.
Note: Supplementary information is available on the Nature Biotechnology website.
Accession codes
Change history
09 October 2008
In the version of this article initially published, an author's name was misspelled as Barbote. The correct spelling is Barabote. The error has been corrected in the HTML and PDF versions of the article.
References
Mandels, M. & Reese, E.T. Induction of cellulase in Trichoderma viride as influenced by carbon sources and metals. J. Bacteriol. 73, 269–278 (1957).
Kuhls, K. et al. Molecular evidence that the asexual industrial fungus Trichoderma reesei is a clonal derivative of the ascomycete Hypocrea jecorina. Proc. Natl. Acad. Sci. USA 93, 7755–7760 (1996).
Nevalainen., H., Suominen, P. & Taimisto, K. On the safety of Trichoderma reesei. J. Biotechnol. 37, 193–200 (1994).
Farrell, A. et al. Ethanol can contribute to energy and environmental goals. Science 311, 506–508 (2006).
Patel-Predd, P. Overcoming the hurdles to producing ethanol from cellulose. Environ. Sci. Technol. 40, 4052–4053 (2006).
Detter, J. et al. Isothermal strand-displacement amplification applications for high-throughput genomics. Genomics 80, 691–698 (2002).
Diener, S. et al. Insight into Trichoderma reesei's genome content, organization and evolution revealed through BAC library characterization. Fungal Genet. Biol. 41, 1077–1087 (2004).
Shapiro, H. Outline of the assembly process: Jazz, the JGI in-house assembler (LBNL Paper LBNL-58236). (Lawrence Berkeley National Laboratory, Berkeley, California, USA, 2005). http://repositories.cdlib.org/lbnl/LBNL-58236/.
Herrera-Estrella, A. et al. Electrophoretic karyotype and gene assignment to resolved chromosomes of Trichoderma spp. Mol. Microbiol. 7, 515–521 (1993).
Carter, G. et al. Chromosomal and genetic-analysis of the electrophoretic karyotype of Trichoderma reesei—mapping of the cellulase and xylanase genes. Mol. Microbiol. 6, 2167–2174 (1992).
Mantyla, A. et al. Electrophoretic karyotyping of wild-type and mutant Trichoderma longibrachiatum (reesei) strains. Curr. Genet. 21, 471–477 (1992).
Galagan, J. et al. The genome sequence of the filamentous fungus Neurospora crassa. Nature 422, 859–868 (2003).
Cuomo, C. et al. The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 317, 1400–1402 (2007).
Taylor, J. & Berbee, M. Dating divergences in the Fungal Tree of Life: review and new analyses. Mycologia 98, 838–849 (2006).
Galagan, J. et al. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005).
Machida, M. et al. Genome sequencing and analysis of Aspergillus oryzae. Nature 438, 1157–1161 (2005).
Nierman, W. et al. Genomic sequence of the pathogenic and allergenic filamentous fungus Aspergillus fumigatus. Nature 438, 1151–1156 (2005).
Zhang, N. et al. An overview of the systematics of the Sordariomycetes based on a four-gene phylogeny. Mycologia 98, 1076–1087 (2006).
Zdobnov, E. & Apweiler, R. InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847–848 (2001).
Coutinho, P. & Henrissat, B. Carbohydrate-active enzymes: an integrated database approach. in Recent Advances in Carbohydrate Bioengineering (eds. Gilbert, H.J., Davies, G., Henrissat, B. & Svensson, B.) 3–14 (Royal Society of Chemistry, Cambridge, 1999).
Cherry, J. & Fidantsef, A. Directed evolution of industrial enzymes: an update. Curr. Opin. Biotechnol. 14, 438–443 (2003).
Divne, C. et al. The 3-dimensional crystal structure of the catalytic core of cellobiohydrolase-I from Trichoderma reesei. Science 265, 524–528 (1994).
Pel, H. et al. Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513.88. Nat. Biotechnol. 25, 221–231 (2007).
Valkonen, M. et al. Spatially segregated SNARE protein interactions in living fungal cells. J. Biol. Chem. 282, 22775–22785 (2007).
Saloheimo, M. et al. Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. Eur. J. Biochem. 269, 4202–4211 (2002).
Cabib, E. et al. The yeast cell wall and septum as paradigms of cell growth and morphogenesis. J. Biol. Chem. 276, 19679–19682 (2001).
Foreman, P. et al. Transcriptional regulation of biomass-degrading enzymes in the filamentous fungus Trichoderma reesei. J. Biol. Chem. 278, 31988–31997 (2003).
Bashkirova, E., Rey, M.W. & Berka, R.M. Combination of suppression subtraction hybridization and microarray technologies to enumerate biomass-induced genes in the cellulolytic fungus Trichoderma reesei. in Applied Mycology and Biotechnology Vol. 5, Genes and Genomics (eds. Arora, D.K. & Berka, R.M.) 275–299 (Elsevier B.V., Amsterdam, The Netherlands, 2005).
Druzhinina, I. et al. Global carbon utilization profiles of wild-type, mutant, and transformant strains of Hypocrea jecorina. Appl. Environ. Microbiol. 72, 2126–2133 (2006).
Hurst, L., Pal, C. & Lercher, M. The evolutionary dynamics of eukaryotic gene order. Nat. Rev. Genet. 5, 299–310 (2004).
Martinez, D. et al. Genome sequence of the lignocellulose degrading fungus Phanerochaete chrysosporium strain RP78. Nat. Biotechnol. 22, 695–700 (2004).
Doi, R. & Kosugi, A. Cellulosomes: plant-cell-wall-degrading enzyme complexes. Nat. Rev. Microbiol. 2, 541–551 (2004).
Seidl, V. et al. A complete survey of Trichoderma chitinases reveals three distinct subgroups of family 18 chitinases. FEBS J. 272, 5923–5939 (2005).
Rosgaard, L. et al. Efficiency of new fungal cellulase systems in boosting enzymatic degradation of barley straw lignocellulose. Biotechnol. Prog. 22, 493–498 (2006).
Tholudur, A., Ramirez, W. & McMillan, J. Mathematical modeling and optimization of cellulase protein production using Trichoderma reesei RL-P37. Biotechnol. Bioeng. 66, 1–16 (1999).
Arora, D.K., Berka, R. & Singh, G.B. Bioinformatics edn. 6 (Elsevier, Amsterdam, 2006).
Salamov, A. & Solovyev, V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000).
Birney, E. & Durbin, R. Using GeneWise in the Drosophila annotation experiment. Genome Res. 10, 547–548 (2000).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Koonin, E. et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 5, R7 (2004).
Kanehisa, M. et al. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, D277–D280 (2004).
Hoberman, R., Sankoff, D. & Durand, D. The statistical analysis of spatially clustered genes under the maximum gap criterion. J. Comput. Biol. 12, 1083–1102 (2005).
Stam, M. et al. Evolutionary and mechanistic relationships between glycosidases acting on alpha- and beta-bonds. Carbohydr. Res. 340, 2728–2734 (2005).
Dean, R. et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434, 980–986 (2005).
Jones, T. et al. The diploid genome sequence of Candida albicans. Proc. Natl. Acad. Sci. USA 101, 7329–7334 (2004).
Goffeau, A. et al. Life with 6000 genes. Science 274, 546–567 (1996).
Dujon, B. et al. Genome evolution in yeasts. Nature 430, 35–44 (2004).
Wood, V. et al. The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880 (2002).
Loftus, B. et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science 307, 1321–1324 (2005).
Acknowledgements
We would like to thank Maggie Werner-Washburne for a critical review of this work, Robert Sensibaugh for his consultation on soil chemistry issues and Glenn A. Stark and Osorio Meirelles for their consultation on statistics. This work was performed under the auspices of the US Department of Energy's Office of Science, Biological and Environmental Research Program and was supported by the University of California, by Lawrence Livermore National Laboratory under Contract No. W-7405-Eng-48, by Lawrence Berkeley National Laboratory under contract No. DE-AC03-76SF00098, by Los Alamos National Laboratory under contract No. W-7405-ENG-36 and by US National Institutes of Health grant GM060201. The work was also funded in part by the European Commission (STREP FungWall grant, contract LSHB-CT-2004-511952).
Author information
Authors and Affiliations
Contributions
D.M., CAZyme gene organization and synteny calculation; R.M.B., annotation of translation machinery; B.H., E.G.J.D. and P.M.C., annotation and comparative analysis of CAZyme genes; M.A., protein domains; M.S. and M.W., annotation of the secretory pathway; S.E.B., B.R. and C.L.S., annotation of secondary metabolism; D.C., annotation of repetitive elements and repeat-induced point mutation; O.C., C.S.H. and T.S.B., genome finishing; I.V.G. and A.S.S., gene modeling; J.C., I.H. and N.P., genome assembly; P.H., annotation of DNA replication, repair and recombination; C.P.K. and M.S., annotation of signaling pathway; L.F.L., annotation of oxidases; A.L.d.L., annotation of transcription factors; J.K.M., central metabolism; S.M., annotation of sexual development; B.N., amino acid metabolism; A.T., N.T., G.X., M.J. and M.M., gene model manual curation; R.B., annotation of transporters; A.W.P., annotation of microtubules and molecular motors; J.Y., annotation of proteases; M.A.N. and N.D.C., genomic analysis; P.R., S.M.L., C.D., D.B., C.R.K, D.S.R. and E.M.R., sequencing.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Notes 1 and 2 and Supplementary Tables 1–9 (PDF 708 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike license (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar license.
About this article
Cite this article
Martinez, D., Berka, R., Henrissat, B. et al. Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nat Biotechnol 26, 553–560 (2008). https://doi.org/10.1038/nbt1403
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1403
This article is cited by
-
Improved consolidated bioprocessing for itaconic acid production by simultaneous optimization of cellulase and metabolic pathway of Neurospora crassa
Biotechnology for Biofuels and Bioproducts (2024)
-
Molecular techniques for the improvement of microbial biocontrol agents against plant pathogens
Egyptian Journal of Biological Pest Control (2023)
-
Effect of itaconic acid production on Neurospora crassa in consolidated bioprocessing of cellulose
Microbial Cell Factories (2023)
-
Role of cellulose response transporter-like protein CRT2 in cellulase induction in Trichoderma reesei
Biotechnology for Biofuels and Bioproducts (2023)
-
Current genetic strategies to investigate gene functions in Trichoderma reesei
Microbial Cell Factories (2023)