Abstract
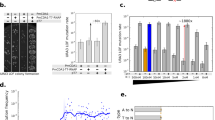
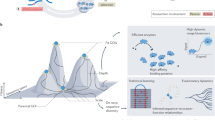
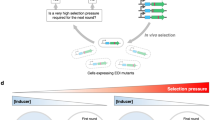
Combining tunable transcription with an enzyme-degradation tag affords an effective means to reduce intracellular enzyme concentrations from high to very low levels. Such fine-tuned control allows selection pressure to be systematically increased in directed-evolution experiments. This facilitates identification of mutants with wild-type activity, as shown here for an engineered chorismate mutase. Numerous selection formats and cell-based screening methodologies may benefit from the large dynamic range afforded by this easily implemented strategy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Taylor, S.V., Kast, P. & Hilvert, D. Angew. Chem. Int. Ed. Engl. 40, 3310–3335 (2001).
MacBeath, G., Kast, P. & Hilvert, D. Protein Sci. 7, 325–335 (1998).
Kast, P., Asif-Ullah, M., Jiang, N. & Hilvert, D. Proc. Natl. Acad. Sci. USA 93, 5043–5048 (1996).
Vamvaca, K., Butz, M., Walter, K.U., Taylor, S.V. & Hilvert, D. Protein Sci. 14, 2103–2114 (2005).
Yano, T., Oue, S. & Kagamiyama, H. Proc. Natl. Acad. Sci. USA 95, 5511–5515 (1998).
Khlebnikov, A., Risa, O., Skaug, T., Carrier, T.A. & Keasling, J.D. J. Bacteriol. 182, 7029–7034 (2000).
Hansen, L.H., Ferrari, B., Sorensen, A.H., Veal, D. & Sorensen, S.J. Appl. Environ. Microbiol. 67, 239–244 (2001).
Lutz, R. & Bujard, H. Nucleic Acids Res. 25, 1203–1210 (1997).
Karzai, A.W., Roche, E.D. & Sauer, R.T. Nat. Struct. Biol. 7, 449–455 (2000).
Burton, S.G., Cowan, D.A. & Woodley, J.M. Nat. Biotechnol. 20, 37–45 (2002).
Turner, N.J. in Enzyme Assays (ed. J.L. Reymond) 139–161, (Wiley-VCH, Weinheim, 2006).
Jestin, J.L. & Kaminski, P.A. J. Biotechnol. 113, 85–103 (2004).
Hillen, W. & Berens, C. Annu. Rev. Microbiol. 48, 345–369 (1994).
Gamper, M., Hilvert, D. & Kast, P. Biochemistry 39, 14087–14094 (2000).
Acknowledgements
This work was generously supported by the Schweizerischer Nationalfonds and the ETH Zurich.
Author information
Authors and Affiliations
Contributions
M.N., P.K. and D.H. designed research; M.N., M.B. and C.H. performed the experiments; M.N., M.B., P.K. and D.H. analyzed data; M.N., P.K. and D.H. wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–4; Supplementary Table 1; Supplementary Methods (PDF 226 kb)
Rights and permissions
About this article
Cite this article
Neuenschwander, M., Butz, M., Heintz, C. et al. A simple selection strategy for evolving highly efficient enzymes. Nat Biotechnol 25, 1145–1147 (2007). https://doi.org/10.1038/nbt1341
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1341
This article is cited by
-
Stochastic chain termination in bacterial pilus assembly
Nature Communications (2023)
-
A growth selection system for the directed evolution of amine-forming or converting enzymes
Nature Communications (2022)
-
An in vivo selection system with tightly regulated gene expression enables directed evolution of highly efficient enzymes
Scientific Reports (2021)
-
Archimedes’ principle for characterisation of recombinant whole cell biocatalysts
Scientific Reports (2018)
-
Sequence-based prediction of permissive stretches for internal protein tagging and knockdown
BMC Biology (2017)