Abstract
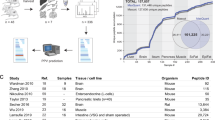
Mass spectrometry–based quantitative proteomics has become an important component of biological and clinical research. Although such analyses typically assume that a protein's peptide fragments are observed with equal likelihood, only a few so-called 'proteotypic' peptides are repeatedly and consistently identified for any given protein present in a mixture. Using >600,000 peptide identifications generated by four proteomic platforms, we empirically identified >16,000 proteotypic peptides for 4,030 distinct yeast proteins. Characteristic physicochemical properties of these peptides were used to develop a computational tool that can predict proteotypic peptides for any protein from any organism, for a given platform, with >85% cumulative accuracy. Possible applications of proteotypic peptides include validation of protein identifications, absolute quantification of proteins, annotation of coding sequences in genomes, and characterization of the physical principles governing key elements of mass spectrometric workflows (e.g., digestion, chromatography, ionization and fragmentation).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Gerber, S.A., Rush, J., Stemman, O., Kirschner, M.W. & Gygi, S.P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. USA 100, 6940–6945 (2003).
Ong, S.E., Foster, L.J. & Mann, M. Mass spectrometric–based approaches in quantitative proteomics. Methods 29, 124–130 (2003).
Wright, M.E. et al. Identification of androgen-coregulated protein networks from the microsomes of human prostate cancer cells. Genome Biol. 5, R4 (2003).
Durr, E. et al. Direct proteomic mapping of the lung microvascular endothelial cell surface in vivo and in cell culture. Nat. Biotechnol 22, 985–992 (2004).
Ranish, J.A. et al. The study of macromolecular complexes by quantitative proteomics. Nat.Genet. 33, 349–355 (2003).
Blagoev, B. et al. A proteomics strategy to elucidate functional protein-protein interactions applied to EGF signaling. Nat. Biotechnol. 21, 315–318 (2003).
Andersen, J.S. et al. Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, 570–574 (2003).
Marko-Varga, G. et al. Discovery of biomarker candidates within disease by protein profiling: principles and concepts. J.Proteome Res. 4, 1200–1212 (2005).
Old, W.M. et al. Comparison of label-free methods for quantifying human proteins by shotgun proteomics. Mol.Cell Proteomics 4, 1487–1502 (2005).
Flory, M.R., Griffin, T.J., Martin, D. & Aebersold, R. Advances in quantitative proteomics using stable isotope tags. Trends Biotechnol. 20, S23–29 (2002).
Kirkpatrick, D.S., Gerber, S.A. & Gygi, S.P. The absolute quantification strategy: a general procedure for the quantification of proteins and post-translational modifications. Methods 35, 265–273 (2005).
Kuster, B., Schirle, M., Mallick, P. & Aebersold, R. Innovation: Scoring proteomes with proteotypic peptide probes. Nat. Rev. Mol. Cell Biol. (2005).
Keller, A. et al. Experimental protein mixture for validating tandem mass spectral analysis. Omics 6, 207–212 (2002).
Desiere, F. et al. Integration with the human genome of peptide sequences obtained by high-throughput mass spectrometry. Genome Biol. 6, R9 (2005).
Craig, R., Cortens, J.P. & Beavis, R.C. Open source system for analyzing, validating, and storing protein identification data. J.Proteome Res. 3, 1234–1242 (2004).
Marzolf, B. et al. SBEAMS-Microarray: database software supporting genomic expression analyses for systems biology. BMC Bioinformatics 7, 286 (2006).
Jones, P. et al. PRIDE: a public repository of protein and peptide identifications for the proteomics community. Nucleic Acids Res. 34, D659–663 (2006).
Kawashima, S. & Kanehisa, M. AAindex: amino acid index database. Nucleic Acids Res. 28, 374 (2000).
De Strooper, B. et al. Deficiency of presenilin-1 inhibits the normal cleavage of amyloid precursor protein. Nature 391, 387–390 (1998).
Xing, Y. & Lee, C. Alternative splicing and RNA selection pressure--evolutionary consequences for eukaryotic genomes. Nat. Rev. Genet. 7, 499–509 (2006).
Eisenberg, D., Marcotte, E.M., Xenarios, I. & Yeates, T.O. Protein function in the post-genomic era. Nature 405, 823–826 (2000).
Rotzschke, O. et al. Exact prediction of a natural T cell epitope. Eur. J. Immunol. 21, 2891–2894 (1991).
Schwartz, D. & Gygi, S.P. An iterative statistical approach to the identification of protein phosphorylation motifs from large-scale data sets. Nat. Biotechnol. 23, 1391–1398 (2005).
Marques, J.T. et al. A structural basis for discriminating between self and nonself double-stranded RNAs in mammalian cells. Nat. Biotechnol. 24, 559–565 (2006).
Schirle, M. et al. Proceedings of the 52nd ASMS Conference on Mass Spectrometry and Allied Topics, Nashville, Tennessee, May 23–27, 2004 (American Society for Mass Spectrometry, Santa Fe, NM 2004).
Tabb, D.L., Huang, Y., Wysocki, V.H. & Yates, J.R. 3rd Influence of basic residue content on fragment ion peak intensities in low-energy collision-induced dissociation spectra of peptides. Anal. Chem. 76, 1243–1248 (2004).
Breci, L.A., Tabb, D.L., Yates, J.R., 3rd & Wysocki, V.H. Cleavage N-terminal to proline: analysis of a database of peptide tandem mass spectra. Anal. Chem. 75, 1963–1971 (2003).
Peng, J., Elias, J.E., Thoreen, C.C., Licklider, L.J. & Gygi, S.P. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC-MS/MS) for large-scale protein analysis: the yeast proteome. J. Proteome Res. 2, 43–50 (2003).
Gavin, A.C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Acknowledgements
The authors are grateful to Julien Gagneur for fruitful discussions and the Cellzome biochemistry, mass spectrometry and informatics teams for generating and managing data. The work was supported in part with federal funds from the National Heart, Lung, and Blood Institute, National Institutes of Health, under contract N01-HV-28179.
Author information
Authors and Affiliations
Contributions
P.M., data (yeast MUDPIT-ESI), data analysis, idea and concept, wrote most of manuscript. M.S., data (yeast PAGE-MALDI, human PAGE-ESI), data mining, wrote part of manuscript. S.C., data (yeast MUDPIT-ESI), idea and concept. M.F., data (yeast MUDPIT-ICAT). H.L, data (yeast MUDPIT-ICAT), D.M., data (yeast MUDPIT-ESI). J.R., data (yeast MUDPIT-ESI, MUDPIT-ICAT). B.R., data (yeast MUDPIT-ICAT). R.S., computation underlying Figure 1b. T.W., data (yeast PAGE-ESI). B.K., idea and concept, wrote part of manuscript. R.A., idea and concept, wrote part of manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Number of Peptide Observations/Protein in Model Mixture.
Supplementary Fig. 2
Histogram of number of confidently predicted proteotypic peptides/protein.
Supplementary Fig. 3
Intersection of Proteins and Peptides among Experiment Types.
Supplementary Fig. 4
Cysteine 2D Histogram.
Supplementary Fig. 5
Mass Distribution of Observed Peptides.
Supplementary Fig. 6
Length (Size) Distribution of Observed Peptides.
Supplementary Table 1
Description of 4 large scale datasets used in study.
Supplementary Table 2
List of observed proteins used in study.
Supplementary Table 3
List of experimentally derived proteotypic peptides used in study.
Supplementary Table 4
Interrogated Physico-Chemical Properties and their discrimination potential for PAGE_ESI .
Supplementary Table 5
Interrogated Physico-Chemical Properties and their discrimination potential for PAGE_MALDI.
Supplementary Table 6
Interrogated Physico-Chemical Properties and their discrimination potential for MUDPIT_ESI.
Supplementary Table 7
Interrogated Physico-Chemical Properties and their discrimination potential for MUDPIT_ICAT.
Supplementary Table 8
Comparison of empirically observed peptide presence with those made by prediction for human γ–secretase.
Supplementary Table 9
Predicted Proteotypic Peptides for Yeast.
Supplementary Table 10
Predicted Proteotypic Peptides for Human.
Supplementary Table 11
Extended Predicted Proteotypic Peptides for Yeast.
Supplementary Table 12
Extended Predicted Proteotypic Peptides for Human.
Supplementary Table 13
Intersection of Proteins by Experimental Approach.
Supplementary Table 14
Intersection of Proteotypic Peptides by Experimental Approach.
Supplementary Table 15
Intersection of Proteins with Predicted Proteotypic Peptides by Experimental Approach.
Supplementary Table 16
Intersection of High Confidence Predicted Proteotypic Peptides by Experimental Approach.
Rights and permissions
About this article
Cite this article
Mallick, P., Schirle, M., Chen, S. et al. Computational prediction of proteotypic peptides for quantitative proteomics. Nat Biotechnol 25, 125–131 (2007). https://doi.org/10.1038/nbt1275
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1275
This article is cited by
-
Understanding How Silicon Fertilization Impacts Chemical Ecology and Multitrophic Interactions Among Plants, Insects and Beneficial Arthropods
Silicon (2023)
-
Proteomic discovery of non-invasive biomarkers of localized prostate cancer using mass spectrometry
Nature Reviews Urology (2021)
-
Elucidation of protein biomarkers for verification of selected biological warfare agents using tandem mass spectrometry
Scientific Reports (2020)
-
Purification and Production of Novel Angiotensin I-Converting Enzyme (ACE) Inhibitory Bioactive Peptides Derived from Fermented Goat Milk
International Journal of Peptide Research and Therapeutics (2020)
-
Quantitative assay of targeted proteome in tomato trichome glandular cells using a large-scale selected reaction monitoring strategy
Plant Methods (2019)