Abstract
A useful approach for exploring gene function involves generating mutant mice from genetically modified embryonic stem (ES) cells. Recent advances in genetic engineering of ES cells have shifted the bottleneck in this process to the generation of mice. Conventional injections of ES cells into blastocyst hosts produce F0 generation chimeras that are only partially derived from ES cells, requiring additional breeding to obtain mutant mice that can be phenotyped. The tetraploid complementation approach directly yields mice that are almost entirely derived from ES cells, but it is inefficient, works only with certain hybrid ES cell lines and suffers from nonspecific lethality and abnormalities, complicating phenotypic analyses. Here we show that laser-assisted injection of either inbred or hybrid ES cells into eight cell–stage embryos efficiently yields F0 generation mice that are fully ES cell–derived and healthy, exhibit 100% germline transmission and allow immediate phenotypic analysis, greatly accelerating gene function assignment.
Similar content being viewed by others
Main
Perhaps the most efficient way to elucidate gene function is by engineering gene mutations in ES cells and from these cells deriving mice that contain these genetic changes1,2,3,4. Mutant phenotypes in these genetically altered mice often provide important insights into gene function. Technological advances in producing genetic alterations in ES cells have resulted in large collections of modified ES cells5,6,7,8 that will soon include mutations for every gene in the genome9,10. These advances have shifted the bottleneck in gene function assignment to the time-consuming steps involved in generating mutant mice that can be phenotyped. Currently, mutant mice are usually generated by introducing genetically altered ES cells into blastocyst-stage host embryos, resulting in F0 generation chimeric mice that are only partially derived from the modified ES cells. If part of the germ line is derived from the modified ES cells, these chimeras can be bred to obtain F1 generation mice that are uniformly heterozygous for the mutation of interest. Subsequent interbreeding of these heterozygous mice can result in F2 generation mice that are homozygous for the intended mutation. Because few mutant phenotypes can be detected in chimeric or heterozygous mutant mice, phenotyping requires derivation of homozygous mutant F2 mice. In addition, chimeras that are estimated to be >90% ES cell–derived based on coat color can be inefficient germline transmitters, because coat color chimerism does not fully reflect ES cell contribution to internal organs (including germ cells). In conclusion, chimeras render the production of homozygous mutant mice suitable for phenotyping inefficient and time-consuming.
Use of tetraploid embryos as hosts for the ES cells can accelerate phenotyping because the resulting F0 generation mice are almost completely ES cell–derived and thus immediately available for phenotypic analyses11,12,13. Unfortunately, the tetraploid approach is an inefficient process for the generation of live mice; it requires certain low-passage hybrid strain ES cell lines and fails to yield viable mice when ES cells from inbred strains are used. In addition, the resulting F0 mice, which can possess up to 2% host contamination, exhibit poor viability and have other abnormalities (such as changes in growth rates and body weight), unrelated to the introduced genetic mutation, that complicate phenotypic analyses11,12,13. Although the tetraploid approach is a major advance over the blastocyst method in terms of speed to phenotype mutant mice, its drawbacks limit its usefulness for large-scale functional genomic efforts.
Here we demonstrate that laser-assisted injection of ES cells into eight cell–stage embryos efficiently generates viable and healthy F0 generation mice that contain no more than 0.1% host contamination. Most importantly, these F0 generation mice, derived from either heterozygous or homozygous mutant ES cells, can be used directly in phenotypic analyses. We demonstrate that the mutant phenotypes in these F0 mice are indistinguishable from those observed in mice derived by conventional breeding. This approach can also use both inbred and hybrid ES cells with either inbred or outbred host embryos. Furthermore, this method works not only with standard male XY ES cells, but also with their XO congenic clones, thus rapidly and efficiently yielding not only male but also female F0 founder mice. As both male and female ES cell–derived F0 generation mice demonstrate 100% germline transmission, these mice can be efficiently intercrossed to generate homozygous mutant mice.
Results
Eight-cell method markedly enhances coat-color contribution
Conventionally, ES cells are injected into the blastocoel within a blastocyst, at which point the cells must integrate within the already formed epiblast (also known as the inner cell mass, ICM), which gives rise to the embryo proper. This injection procedure results in mice that are chimeric with varying degrees of ES cell contribution (Fig. 1a). In an effort to increase the contribution of ES cells, we injected ES cells into earlier embryos at the eight-cell stage before formation of the ICM, because early studies suggested that injection into earlier embryos might result in higher degrees of ES cell contribution14,15. We reasoned that introduction before ICM formation might provide a competitive advantage to the injected ES cells—which already are committed to the epiblast lineage—over the endogenous blastomeres in terms of giving rise to the ICM and the embryo proper. Because previous attempts to introduce ES cells into eight cell–stage embryos involved laborious manipulations to penetrate the zona pellucida14,15, perhaps explaining the absence of any follow-up studies using such injections, we attempted to facilitate such eight cell–embryo injections by perforating the zona pellucida with a laser pulse from the XYClone laser system, and subsequently injecting ES cells through this perforation into the perivitelline space (Fig. 1b).
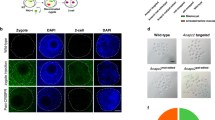
(a) In a conventional blastocyst injection (illustrated in cartoon at top, left) ES cells are injected into the blastocoel by piercing the trophectoderm at a cell-cell junction. The merged fluorescent/bright-field photomicrograph (middle, left) shows a real example in which ES cells that express green fluorescent protein (eGFP) from the Gt(ROSA)26Sor locus promoter were injected into a blastocyst. The injected ES cells mingle with the preexisting cells of the inner cell mass (ICM) (see Supplementary Video 1 online). The injected ES cells compete with the host ICM of the blastocyst to yield F0 Chimeras. (b) In an eight cell–stage embryo injection, ES cells are injected through a perforation in the zona pelucida (ZP) created by a laser pulse (illustrated in cartoon at top, right). The photomicrograph on the left beneath the cartoon shows a bright field image of an eight cell–stage embryo on a holding pipette immediately following the laser pulse just before injection (red arrow points to the perforation in the ZP). The adjacent merged fluorescent/bright-field image and the image immediately beneath show eGFP-expressing ES cells, as in (a), immediately after injection into the perivitelline space and in a resulting blastocyst stage embryo (white arrowheads indicate the boundary of the blastoceol, see Supplementary Video 2 online). The ES cells appear to have an advantage over the existing blastomeres in the formation of the ICM lineage, which results in fully ES cell–derived F0 mice. (c,d) Chimeras (c) and fully ES cell–derived mice (d) produced by introduction of inbred C57BL/6 ES cells into either blastocysts (c) or eight cell–stage embryos (d) from the C57BL/6Tyrc-2J albino strain (see also Table 1 and Supplementary Fig. 1 online).
We compared conventional blastocyst injections16 with laser-assisted injection of ES cells into eight-cell embryos using a wide assortment of ES cells and host embryo combinations. In every case, the eight cell–embryo injections resulted in dramatically higher ES cell contribution than the blastocyst injections, as judged by the extent of ES cell–derived coat color (Table 1 and Fig. 1c,d, in which increased black coat color reflects increased ES cell contribution). Remarkably, several of the combinations using the eight-cell method resulted in a preponderance of male mice with 100% ES cell–derived coat color (e.g., Fig. 1d), as well as a smaller number of male and female mice with 100% host-derived coat color, and no chimeras. These results suggested that the eight-cell method could lead to a much greater contribution to the developing embryo via an 'all or none' phenomenon—either the ES cells contributed substantially to the ICM, thus resulting in complete coat color contribution and gender conversion to males, or the mice were entirely host derived.
To begin to understand the process by which the eight-cell method may improve ES cell contribution compared to the traditional blastocyst method, we followed the fate of ES cells expressing green fluorescent protein (eGFP) that were injected by both approaches. Fluorescent images of blastocysts resulting from the eight-cell method suggested that the ICM was derived entirely from the green fluorescent ES cells, whereas the ICM of blastocysts following traditional injections was—as expected—clearly chimeric (compare blastocyst images in Fig. 1a,b, and Supplementary Fig. 1d,e; Supplementary Videos 1 and 2 online depict serial confocal sections through these blastocysts, further demonstrating the marked differences in ICM contributions using the two methods). Supplementary Figure 1 shows all the steps of the eight-cell method from laser perforation of the zona pellucida through ES cell contribution to the ICM. Combinations of ES cells and host embryos that worked exceedingly well (resulting in F0 mice with 100% ES cell–derived coat color and no chimeras) included the introduction of either inbred 129 and C57BL/6 (B6) ES cells or hybrid 129/B6 F1 (129SvEv/B6) into B6 or C57BL/6Tyrc-2J (B6 albino) host embryos (Table 1). These combinations worked similarly well when using the parental ES cell lines or their targeted subclones (Table 1). Other combinations that have proven traditionally more challenging using the blastocyst approach (such as inbred B6 or hybrid F1 ES cells into outbred Swiss Webster (SW) hosts, or BALB/c ES cells into B6 hosts) were in all cases markedly superior using the eight-cell approach, though these more challenging combinations yielded occasional chimeras in addition to the mice with 100% ES cell–derived coat color (Table 1). Surprisingly, when we injected VGF1 parental cells and targeted clones into SW host embryos, we observed a better outcome from the clones. Although we do not have a definitive explanation for this result, one possibility is that the parental ES cell line is composed of a broad population of cells with variability in its ICM colonization. Thus, the data that we show from specific targeted clones may have been derived from the better performing cells in the parental population. Regardless of how the ES cell clones perform, they always contribute more to the F0 mouse with the eight-cell method as compared with conventional blastocyst injection.
Eight-cell method markedly enhances germline transmission
The striking improvements in ES cell contribution to coat color using the eight-cell method suggested that this approach might similarly result in increased ES cell contribution to other tissues, which should also have the benefit of resulting in higher rates of germline transmission by these mice. F0 generation male mice generated from a sampling of the above injections were bred to test for their transmission of both the ES cell–derived coat color as well as the modified allele in the targeted clones (Supplementary Table 1 online). Only 66% (72 of 109) of the chimeras produced by the blastocyst injections exhibited 100% germline transmission. In contrast, 96% (81 of 84) of the F0 mice generated by the eight-cell method exhibited 100% germline transmission. The remaining 4% (3 of 84) of these mice failed to sire any litters. In agreement with our experience, this rate of nonbreeding males is similar to the breeding behavior of normal males from 129 and B6 strains. Taken together, these results suggest that the male ES cells administered by the eight-cell method completely convert the host embryo gender, and fully contribute to the germ cell lineage.
In an even more stringent test of the eight-cell method, we made use of female (39,XO) ES cell subclones generated from male (40,XY) ES cell clones that have spontaneously lost their Y chromosome. Converting the gender of male embryos requires nearly complete colonization of the host genital ridge17. Female mice with a naturally occurring 39,XO genotype are normal, healthy and fertile. We observe the same phenotype in female F0 mice generated from the injection of 39,XO ES cells into eight-cell embryos. Female (39,XO) ES cells produced complete gender conversion as well as 100% germline transmission by the eight-cell approach, but not by blastocyst injection (Supplementary Tables 2 and 3 online). These results demonstrate that the eight-cell method is able to trigger nearly complete contribution of the injected ES cells to the genital ridge in both male and female host embryos. As expected, an intercross between these 39,XO female F0 mice and F0 male mice, derived from the parental XY ES cell clones by the eight-cell method, produces homozygous mutant mice in the first generation (data not shown). Our finding that the eight-cell method enables female ES cells to completely convert the gender of injected embryos offers an accelerated path to homozygous mutant mice.
Though only part of an initial survey, these results suggest that the eight-cell method may prove useful for many different ES cell/host embryo combinations, including those that contain female XO ES cells and others that have been difficult to obtain. The apparent flexibility of the eight-cell method in tolerating previously unproductive ES cell/host embryo combinations may also provide distinct biological advantages, and substantial cost savings for the production of genetically modified mice in defined genetic backgrounds.
Sensitive assays do not detect host contamination
Our observations of complete ES cell coat contribution, 100% germline transmission and complete gender conversion suggested that the eight-cell method might markedly enhance ES cell contribution to multiple tissues as compared to the conventional blastocyst injection approach. To test this, we used three sensitive and distinct assay approaches to determine the presence of host embryo contribution in F0 mice and embryos produced from blastocyst and eight cell–embryo injections: (i) a quantitative genotyping assay, (ii) a lacZ reporter gene expression assay in adult tissues and (iii) an eGFP fluorescence assay in embryonic tissues. For the quantitative genotyping assay approach, we made use of an XY ES cell line (VGF1.1202) with a targeted disruption of the X-linked—and thus single-copy—gene, Il2rg, which encodes the gamma chain of interleukin-2 receptor (IL-2Rγ). Because these targeted ES cells lack a wild-type version of this gene, any detection of Il2rg DNA in F0 mice derived from injections of the VGF1.1202 line would necessarily indicate the extent of host embryo contamination (Fig. 2a). We detected Il2rg DNA using a TaqMan-based quantitative PCR assay in which threshold cycle (CT) values >40 correspond to a host cell contribution of <1 cell in 1,000 (0.1%). Six tissues (lung, liver, heart, kidney, skeletal muscle and tail) were analyzed from each F0 mouse (Fig. 2). As expected, a conventional knockout mouse for this gene (Fig. 2a, far right) has an average CT value for these six tissues >45, consistent with complete lack of this gene, whereas a wild-type mouse has an average CT value of ∼27 (Fig. 2a, far left). The same assay carried out on seven chimeras generated by conventional blastocyst injections and having >90% ES cell coat color contribution yielded average CT values of ∼27–29, indicating substantial amounts of host cell contamination in the examined tissues. In contrast, analysis of five mice generated with the eight-cell method all yielded average CT values >45, indistinguishable from knockout mice produced by conventional breeding, and indicating that host contamination could not be detected. Assays for two nontargeted reference genes produced similar average CT values in all the samples (data not shown). These results support the notion that the laser-assisted eight-cell method yields F0 generation mice that are substantially, if not completely, ES-cell derived (that is, >99.9% ES-cell derived).
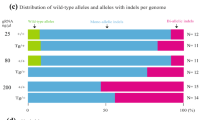
(a) Quantitative PCR of the host embryo-specific Il2rg gene DNA in tissues obtained from high-quality coat-color chimeras (>90%) resulting from the injection of targeted XY ES cell clone VGF1.T1202A-D8 containing a deletion of the X-linked Il2rg gene into B6 blastocysts (B1-B7) or from F0 mice produced from B6 eight-cell embryos (V1-V5). Each bar represents the mean CT value of quadruplicate assays for each of six tissues (lung, heart, liver, skeletal muscle, kidney and spleen). Error bars represent the s.e.m. A CT value of 40, indicated by the dashed red line, is the approximate limit of detection for the assay and represents the detection of one wild-type (host) Il2rg allele in 1,000 male genomic equivalents. Bars labeled WT and KO represent assays of a single tissue from a wild-type and Il2rg null mouse. (b) Quantitative PCR of the host embryo-specific albino tyrosinase alleles in tissues obtained from F0 mice resulting from the injection of parental B6 ES cells (VGB6.2) or targeted VGB6.2 ES cell clones into host B6 albino blastocysts or B6 albino and SW eight-cell embryos. The bar labeled blastocyst control represents the assay for a chimera generated by VGB6.2 cells injected into a B6 albino blastocyst. The tissue replicate assays were performed as in a. A CT value of 40, indicated by the dashed red line, is the approximate limit of detection for the assay and represents the detection of one mutant tyrosinase (host) allele in 2,000 genomic equivalents. The wild-type (B6) tyrosinase allele was detected (average CT = 28) in all of the samples, indicating B6 ES cell contribution to the F0 mice (data not shown). (c) Comparison of X-gal stained whole mounts of skin hair follicles, and tissue sections of liver, and brain from a representative high-quality (>90%) coat-color chimera from a blastocyst injection (top panels) and a representative mouse produced by an eight cell–embryo injection (bottom panels). Host embryos heterozygous for a lacZ gene knocked into the Tmem97 locus were injected with a targeted VGF1 ES cell clone devoid of lacZ gene expression. The scale bars represent 3.0 mm for skin (left panels), liver (left-middle panels) and brain (middle-right panels), 0.43 mm for liver (middle panels) and 0.38 mm for brain (far-right panels). (d) Comparison of representative eGFP fluorescence in E15.5 embryos and extra-embryonic tissues (Ex) produced from blastocyst injections (left panels) and from eight-cell injections (right panels). Host embryos heterozygous for an eGFP transgene knocked into the ubiquitously expressed Gt(ROSA)26Sor gene were injected with a VGF1 ES cell clone lacking an eGFP gene. The extra-embryonic tissues, the vast majority of which are known not to be ES cell–derived, exhibit host embryo contribution from both blastocyst and eight cell–embryo injections, serving as an internal control for eGFP fluorescence.
We repeated this analysis to see if fully ES cell–derived mice could be produced by injecting inbred parental B6 ES cells or targeted B6 ES cell clones into albino B6 hosts or outbred albino SW hosts. In these analyses, contaminating host cell contribution is indicated by the detection of the mutant tyrosinase gene present in either albino host (Fig. 2b). Whereas a 90% coat color chimera derived from a conventional blastocyst injection had an average CT value for the six tissues analyzed of ∼33 (F0 chimera in Fig. 2b, left bar), indicating readily detectable host-embryo contribution, F0 mice from eight cell–embryo injections exhibited complete B6 ES cell–derived coat color and had no detectable host contribution by the tyrosinase assay (CT values >40, corresponding to host cell contribution <1 in 2,000 cells, or <0.05%).
In the second approach for determining host contamination, we attempted an even more sensitive screen that could detect a single contaminating host cell (Fig. 2c). We used host embryos that contained a sensitive lacZ reporter knocked into the Tmem97 locus, resulting in robust and stereotypic reporter expression in hair follicles, liver and brain (data not shown). When ES cells lacking this lacZ reporter are introduced into these reporter-marked host embryos, any detectable reporter staining would reflect host contamination. In 6-week-old chimeric mice derived from conventional blastocyst injections and with a high percentage of ES cell–coat color, host-derived reporter expression was widely observed, indicating substantial host contamination (Fig. 2c, upper panels). In contrast, no reporter expression was detected in 24 F0 mice derived from the eight cell–stage approach (e.g., Fig. 2c, lower panels). Because we have shown that this reporter approach can detect even single cells, these results indicate that the eight-cell method yields mice that rarely, if ever, are contaminated by host cells.
We performed a final assay using a broadly expressed reporter (that is, the eGFP transgene targeted to be under control of the ubiquitously expressed ROSA26 promoter), which would allow for a more comprehensive assessment of host cell contamination by the extent of green fluorescence detectable at embryonic age E15.5 (Fig. 2d). Embryos derived from conventional blastocyst injections showed prominent eGFP fluorescence (Fig. 2d, left panels), indicating significant host contamination. In contrast, embryos derived from eight-cell injections showed no fluorescence (Fig. 2d, right panels). For both methods of injection we observed green fluorescence in the extra-embryonic tissues, consistent with previous results using tetraploid approaches in which host contribution to extra-embryonic tissues is substantial because of a limited ability of epiblast-committed ES cells to colonize these tissues18.
F0 mice directly used for phenotyping studies
Phenotyping traditionally requires breeding of F0 generation chimeras to generate F1 and F2 mice that are uniformly heterozygous or homozygous for the mutation of interest. We explored whether F0 mice generated via the eight-cell method, that appear substantially—if not completely—ES cell–derived, could be used directly for phenotypic analyses. We selected three different mutations that based on their well-described qualitative and quantitative phenotypic features might easily indicate nonspecific abnormalities as well as host embryo contamination and contribution. The first mutation involves heterozygous deletion of the gene encoding insulin-like growth factor 1 (IGF-1), which has been described as having a subtle but clear-cut effect on body weight19. The second mutation involves deletion of the X-linked gene encoding IL2Rγ, a requisite common subunit of the receptors for IL-2, IL-4, IL-7, IL-9 and IL-15. Because IL2Rγ is absolutely essential for immune development, null mutations cause severe combined immunodeficiency (SCID) in both humans20 and mice21,22. The quantitative nature of the reported defects in the B lymphocyte, T lymphocyte and natural killer (NK) cell compartments of mutant male mice, as well as the ability of small numbers of injected wild-type ES cells to completely rescue SCID phenotypes by repopulating the missing compartments23, make this an ideal model for evaluating the suitability of the eight cell–embryo method for direct immune phenotyping of F0 mice, as well as for ruling out even minute host contamination in these F0 mice. The final phenotyping analysis was performed on mice derived from ES cells harboring a homozygous mutation of the gene for delta-like ligand-4 (Dll4) that has previously been shown to result in uniform embryonic lethality due to dramatic vascular defects24.
In all three phenotyping cases, no unexpected abnormalities or lethalities were present in the F0 cohorts generated using the eight-cell method, and the observed phenotypes were indistinguishable from those noted in the literature as well as in mutant mice produced contemporaneously by conventional breeding. In the IGF-1 phenotyping example, we targeted an inbred B6 ES line (VGB6) to create an ES cell clone (VGB6.2T734C-D5) containing a heterozygous null allele with a lacZ reporter gene in the place of the Igf1 coding region. These targeted ES cells were then introduced into eight-cell SW embryos, and the injected embryos were implanted into surrogate SW mothers along with embryos injected with the parental VGB6 line to serve as wild-type control littermates. The effect of heterozygous deletion of the gene for IGF-1 on postnatal growth deficiency in these F0 mice matches previous reports19, and furthermore is remarkably similar to the growth deficiency noted in conventional F1 heterozygotes (compared to their wild-type littermates) produced by breeding the F0 Igf1+/− mice to B6 females (Fig. 3). Thus, in this example, the F0 mice produced by the eight-cell method are suitable for direct phenotyping and obviate the need for further breeding to obtain F1 generation mice. To independently verify that these F0 Igf1+/− mice were fully ES-cell derived, we performed the tyrosinase gene assay specific for the SW host embryo strain as a measure of host contamination, and determined that it was undetectable in five tissue DNA samples from six independent F0 Igf1+/− mice (data not shown).
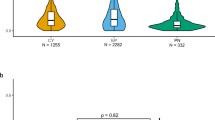
An Igf1+/−ES cell clone, VGB6.2.T734C-D5, and the parental VGB6.2 ES cells were separately injected into Swiss Webster (SW) eight cell–stage embryos and then combined for transfer into SW recipient females to obtain Igf1+/− and Igf1+/+ littermates. Tail biopsies were taken at postpartum day 8 (P8) for genotyping of the Igf1 allele. (a) Postnatal growth curves of Igf1 +/− F0 mice (open circles, n = 7) and Igf1+/+ littermates (closed circles, n = 6) obtained from body weight measurements beginning at P4 and taken twice weekly. F0 mice were weaned at P28. Only measurements of litters containing a minimum of two Igf1+/− and two Igf1+/+ mice were included in the analysis. (b) The F0 Igf1 heterozygous mice from a were bred to B6 females to produce Igf1+/− (open triangles, n = 3) and Igf1+/+ F1 offspring (closed triangles, n = 4). To more accurately compare their postnatal growth with that of the F0 mice, these F1 mice were fostered at birth to SW mothers and weighed on the same postnatal schedule. Error bars indicate s.e.m.
In the second phenotyping example (Fig. 4), we used hybrid 129/B6 F1 ES cells (VGF1) targeted to carry a null allele of the X-linked—and thus single copy—gene encoding IL2Rγ. Two independently targeted VGF1 ES cell clones (A-B6 and A-F2), as well as the parental VGF1 cells as a control, were separately injected into eight-cell SW host embryos and then transferred to SW foster mothers. Our quantitative genotyping assay on the resulting F0 mice (all male, as expected) for the SW-associated tyrosinase mutation confirmed lack of detectable host contamination, whereas quantitative genotyping on the F0 male mice confirmed complete loss of the Il2rg gene (Fig. 4a). Consistent with this, analyses by flow cytometry of surface expression of the IL2Rγ receptor subunit on spleen and thymus revealed a total absence of IL2Rγ-expressing cells in the F0 mice carrying the disrupted allele (Fig. 4b). As would be expected based on the absence of IL2Rγ-expressing cells, these F0 mice exhibited immune defects identical to those previously reported in conventional IL2Rγ null mice, that is, a total absence of IgM+ B220+ B cells in all compartments (Fig. 4c), a slight increase in CD4+ T cells and a lack of CD11b+ DX5+ NK cells (data not shown). Because of the ability of rare wild-type cells to repopulate missing cellular compartments in SCID models, the phenotype observed is consistent with a lack of contaminating host embryo cells in the mice generated via the eight-cell method. We have confirmed this phenotype using the same mutation targeted into V17, a commercially available 129/BALBc hybrid F1 ES cell line (data not shown).
Two VGF1 XY clones, VGF1.T5057A-B6 (A-B6) and VGF1.T5057B-F2 (B-F2), were injected into B6 eight-cell embryos to produce F0 generation mice. (a) Quantitative PCR genotyping screen for host embryo contamination of F0 mice. DNA was obtained from heart (H), skeletal muscle (M), lung (L), kidney (K) and liver (Li) from three F0 mice from each clone and screened for the host-specific Il2rg allele. Each bar represents the mean CT value of quadruplicate assays for each of the five tissues. A CT value of 40, indicated by the dashed line, is the approximate limit of detection for the assay and represents the detection of one wild-type (host) Il2rg allele in 1,000 male genomic equivalents. Compared to a wild-type (WT) mouse, F0 Il2rg−/y mice had no detectable wild-type (host) Il2rg allele. (b) Surface levels of IL-2Rγ in spleen and thymus in wild-type and Il2rg−/y F0 mice from the A-B6 and B-F2 clones as measured by flow cytometry analysis show no detectable presence of IL-2Rγ (y-axis, cell counts; x-axis, fluorescence intensity). (c) Effect of the Il2rg −/y null mutation on B-cell populations in bone marrow, and spleen of F0 mice from the A-B6 and B-F2 clones compared with a wild-type mouse analyzed for the B cell markers B220-APC and IgM-PE (both axes, fluorescence intensity).
In the final phenotyping example (Fig. 5), we used hybrid VGF1 ES cells homozygous for a targeted deletion of the entire coding sequence of Dll4 (ref. 25) to produce F0 generation mice by the eight-cell method. F0 embryos generated from the injection of these Dll4Lz/Lz ES cells into eight-cell B6 host embryos exhibited the expected lethality, with 100% penetrance, due to severe vascular defects (Fig. 5), as previously described for a similar Dll4 allele in which the first three coding exons were deleted24.
A VGF1 ES cell clone, VGF1.T540E-A5, containing a Dll4−/− homozygous null mutation, was injected into B6 eight cell–stage embryos and E9.5 F0 embryos were harvested. The panels show Anti-PECAM1 immunohistochemistry results as follows: (a) a yolk sac from a Dll4+/+ embryo; (b) a Dll4+/+ embryo; (c) a yolk sac from a Dll4−/− embryo; (d) a Dll4−/− embryo; (e) a yolk sac from a Dll4−/− embryo; (f) a Dll4−/− embryo. Green arrows point to normally developing vasculature in the yolk sac and embryonic head. Red arrows indicate the absence of normal vasculature. Scale bars, 0.31mm.
Discussion
Approaches to assigning gene function, including conventional blastocyst-based approaches as well as tetraploid-based approaches, have significant limitations, and cannot support large-scale functional genomic efforts. Previous work14 showed that injection of hybrid ES cells into eight-cell embryos resulted in greater ES cell coat color contribution and germline transmission as compared to conventional blastocyst injections, but did not quantitatively assess the level of ES cell contribution to the skin or other tissues, and did not assess whether the mice were suitable for direct phenotypic study. Here we report a method by which laser-assisted injection of ES cells into eight cell–stage embryos can efficiently result in viable and healthy F0 generation mice that exhibit 100% germline transmission, contain <0.1% host contamination, and may in fact be completely ES cell–derived. In addition to producing mice that are largely, if not completely ES cell–derived, the eight-cell method results in numbers of healthy weaned mice that are similar to those obtained with the blastocyst approach. Most importantly, the resulting F0 mice can be used directly for phenotypic analyses, yielding phenotypes indistinguishable from those observed in conventionally derived F1 and F2 generation mice.
The ability of the eight-cell method to yield largely if not fully ES cell–derived F0 mice is supported by our observations of complete ES cell coat color contribution, 100% germline transmission, complete gender conversion, as well as by the quantitative genotyping and sensitive host-reporter analyses. The somewhat unexpected ability of the eight-cell method to yield nearly if not completely ES cell–derived F0 mice might be partially due to the fact that the laser only minimally affects the integrity of the zona pellucida, the envelope that provides three-dimensional contacts between ES cells and blastomeres. In addition, the injection of individually selected ES cells into eight-cell embryos, rather than standard aggregation of clumped ES cells with eight-cell embryos (ref. 18), may also contribute to the greater success of the eight-cell method. In any case, the data clearly demonstrate that the eight-cell method results in F0 generation mice that are largely if not completely ES cell–derived, and addresses major limitations of other approaches. As with the tetraploid approaches, however, the eight-cell method does not efficiently contribute to extra-embryonic tissues, and thus phenotypes due to these tissues are also likely to be missed with this approach when analyzing F0 mice.
Perhaps the most important implications of the eight-cell method are for high-throughput functional genomics. Although recent advances in ES cell targeting have enabled mutagenesis at a genome-wide scale, with ES cells soon to be available with mutations for every gene in the genome through KOMP9 (http://www.genome.gov/19517927; http://sciencenow.sciencemag.org/cgi/content/full/2006/907/1) and EUCOMM10, production of mice from these targeted ES cells has remained limiting. We demonstrate that the eight-cell method can generate cohorts of F0 mice that are immediately suitable for phenotyping, thus enabling large-scale genome-wide efforts by eliminating generations of extensive breeding that were previously required to detect mutant phenotypes. The eight-cell method is effective with inbred ES lines (such as C57BL/6) that are preferred for phenotyping.
The power of the eight-cell approach might be maximized when combined with the rapid generation of inbred ES cell clones harboring multiple mutations, such as homozygous mutations or more complex compound genetic modifications that enable conditional approaches. Conventional approaches require multiple rounds of breeding to combine mutations within a given mouse, produce the desired mice at low frequencies in the final breeding step and usually result in mice with a mixed genetic background. All these difficulties preclude efficient incorporation of phenotyping approaches into high-throughput genome-wide mouse genetic screens. Importantly, the eight-cell method, when combined with the ability of VelociGene8 to rapidly generate ES clones harboring homozygous or compound mutations, could enable efficient and high-throughput phenotyping screens using cohorts of F0 generation mice generated directly from modified inbred ES cells.
Since the laser-assisted eight-cell injection method can dramatically reduce the time, breeding effort and mouse husbandry resources required to take advantage of genome-wide ES cell targeting initiatives, we refer to it as the VelociMouse method.
Methods
Embryo donor and recipient mice.
C57BL/6 (B6), 129Sv/Ev and SW mice were purchased from Taconic Farms. C57BL/6Tyrc-2j (B6 albino) mice were purchased from The Jackson Labs. All mice MEGAscript, Ambion were housed in isolator units and maintained on a 10-h dark/14-h light cycle. For the production of eight cell–stage embryos, 3-week old females were superovulated with intraperitoneal injections of pregnant mare serum gonadotrophin (PMSG) (5 IU/mouse, Calbiochem) and human chorionic gonadotrophin (hCG) (7.5 IU/mouse, Calbiochem). PMSG was administered 46–48 h, and hCG was administered 2 h before mating. The females were screened for vaginal plugs the next morning (0.5 days post coitum, dpc) and housed until embryos were collected (see below). SW females were mated with vasectomized SW males and used as recipients for injected embryo transfer at 2.5 dpc.
ES cell culture.
VGF1 (129Sv/EvB6 F1), VGB6.1 (B6), VGB6.2 (B6) and VGBalbc (BALB/c) ES cell lines were established by previously published methods26. We used Dulbecco's Minimal Essential Medium (DMEM) with high glucose, 0.1 mM nonessential amino acids, 1 mM sodium pyruvate, 0.1 mM 2-mercaptoethanol, 2 mM L-glutamine, penicillin and streptomycin (50 μg/ml each, Gibco), 15% FBS (Hyclone), 2000 U/ml LIF (Chemicon) to grow and maintain ES cells, except for B6-derived ES cells, which were grown in DMEM with high glucose, 0.1 mM nonessential amino acids, 1 mM sodium pyruvate, 0.1 mM 2-mercaptoethanol, 2 mM L-glutamine, penicillin and streptomycin (50 μg/ml each), 15% KO Serum Replacement (Invitrogen) and a 50% medium conditioned by L-cells (American Type Culture Collection (ATCC) no. CRL-2647) grown according to the instructions provided by the ATCC. Gene targeted clones were generated by the VelociGene method as described previously8. ES cells heterozygous for the Dll4 mutation containing an attenuated neomycin phophotransferase (neor) gene (imparts resistance to G418) under the control of the mouse phosphoglycerol kinase gene promoter25 were passaged twice then plated as single cells and selected in 0.4 mg/ml (four times the normal concentration used to select for targeted clones) of G418 (Sigma). Colonies were picked after 10 days and screened by a VelociGene loss of native allele assay8 for the absence of the deleted Dll4 gene sequence and the presence of two copies of the neor gene.
Generation of F0 mice.
The introduction of mouse ES cells into mouse blastocysts was done as described16. Eight-cell embryos were flushed from the oviducts of plugged females at 2.5 dpc in standard ES cell medium (without leukemia inhibitory factor, LIF). Embryo injections were performed on eight-cell embryos before obvious compaction using a micromanipulator set-up identical to that for blastocyst injections. An eight-cell embryo, while being restrained by the holding pipette, is prepared for injection by making a perforation in the zona pellucida (ZP) with the delivery of an 800 μs tangential laser pulse (at 100% power) from an XYClone laser system (Hamilton Thorne Bioscience) to the outer margin of the ZP, as it appears in the microscope, in a region that is selected to be as far removed from any blastomeres as possible, guided by a virtual 'aiming reticle' provided by the manufacturer's software. This system allows for the alignment, temperature control and delivery of a pulse precisely to ablate a small portion of the zona pellucida without damaging the embryo. A standard injection needle was used to introduce eight ES cells through the perforation in the ZP and into the perivitelline space. Eight-cell embryos injected with VGF1, 129, VGBalbc and 129/Balbc ES cells were cultured overnight at 37 °C/5% CO2 in KSOM (Chemicon), while those injected with B6 ES cells were cultured in DMEM (Invitrogen) containing L-cell conditioned medium. We have found that the laser-assisted injection of eight cell–stage embryos is easily learned by anyone competent in standard blastocyst injection technology. In fact, the same injectors originally trained for blastocyst injection have approximately twice the throughput, for the reasons stated below. The method utilizes the identical injection equipment except for the addition of the laser system. We harvest eight cell–stage embryos from oviducts at 2.5 dpc and inject them before obvious compaction. Because the eight-cell method requires the injection of only 7–9 ES cells rather than the 12–15 ES cells that are normally injected into blastocysts, the injection of eight cell–stage embryos allows for roughly twice the number of injections in the same amount of time. We have found that the injection of fewer than six ES cells more often results in chimeric mice. The injection of more than nine cells has no added benefit because the excess cells tend to leak out through the opening in the ZP. When we injected morula-stage embryos, the yield of F0 births was greatly reduced, and the technique was more difficult. We have not attempted to inject embryos younger than the eight-cell stage because of the timing of these embryos. Only healthy embryos that have progressed from the eight-cell stage during overnight culture are transferred to the uterine horns of pseudo-pregnant SW females by standard methods. All animals were maintained and processed under the auspices of the Internal Animal Care and Use Committee of Regeneron Pharmaceuticals, Inc.
Genotype analysis.
We used TaqMan (Applied Biosystems) Q-PCR to assay the mouse tail DNA for the eGFP reporter gene, the X-linked Il2rg gene, the mutation in the tyrosinase gene that is responsible for albinism in the SW and B6 Tyrc-2j strains27, the Sry gene to screen for loss of the Y chromosome, and for SOSTDC1 and Wnt2b as reference genes. All DNA samples from mouse tail biopsies were analyzed in quadruplicate in an Applied Biosystems PRISM 7900HT Sequence Detection System, which determined the CT for each reaction, the point in the PCR at which the fluorescence signal reached the preset threshold. PCR primers (Gene Link) and TaqMan probes (5′-FAM-3′- Black Hole Quencher, Biosearch); 5′-FAM- or 5′-VIC-Minor Groove Binder Non-Fluorescent Quencher (MGBNFQ; Applied Biosystems) used were as follows: eGFP forward, 5′-CTACCCCGACCACATGAAGC–3′; eGFP reverse, 5′–TGCGCTCCTGGACGTAGC–3′; eGFP probe, 5′–FAM-TCTTCAAGTCCGCCATGCCCG-BHQ–3′; Il2rg forward, 5′–CACAGACTACACCCAGAGAAAGAAGA–3′; Il2rg reverse, 5′–AGGACTAAGAAGGATCTAGGTGACAATAA–3′; Il2rg probe, 5′–FAM– AGCACCATGTTGAAACT–MGBNFQ–3′; (SW albino assay) tyrosinase forward, 5′–GGCAACTTCATGGGTTTCAAC–3′; (SW albino assay) tyrosinase reverse 5′–GACTCGCTTCTCTGTACAATTTGG–3′; (SW albino assay) tyrosinase WT allele probe, 5′–FAM-ATCCAAACTTACAGTTTC-MGBNFQ–3′; (SW albino assay) tyrosinase mutant allele probe, 5′–VIC-ATCCAAACTTAGAGTTTC-MGBNFQ–3′; (B6 Tyrc-2j albino assay) tyrosinase forward, 5′–GCACCATCTGGACCCTCAGTTC–3′; (B6 Tyrc-2j assay) tyrosinase reverse 5′–GCAGTTGAAACCCATGAAGTTG–3′; (B6 Tyrc-2j albino assay) tyrosinase WT allele probe, 5′–FAM-TGACCGTGAGTCCT-MGBNFQ–3′; (B6 Tyrc-2j albino assay) tyrosinase mutant allele probe, 5′–VIC-TGGATGACCTTGAGTCC-MGBNFQ–3′; Sry forward, 5′–GCAGCTTACCTACTTACTAACAGCTGAC–3′; Sry reverse, 5′–GCTGTGCTGAGGTGCTCCTG–3′; Sry-probe, 5′–FAM–CACTGGTGAGCATACAC–MGBNFQ–3′; SOSTDC1 forward, 5′–TTAAACCTGTCCCGGCACA–3′; SOSTDC1-reverse, 5′–CTGCCTCCATTCCTGGCTT–3′; SOSTDC1 probe, 5′–FAM–CCCAGCAGCAACAGCACCCTGAA–BHQ–3′; Wnt2b forward, 5′–GCAGCTGTGACCCATATACCC–3′; Wnt2b reverse, 5′–CACTACAGCCACCCCAGTCAA–3′; and Wnt2b-probe, 5′–FAM–CGGTCGGCACCATGATCAACGA–BHQ–3′.
Beta-galactosidase/anti-PECAM immunohistochemistry.
Six-week old mice were killed by a lethal dose of an anesthetic. Tissues to be used for DNA analysis were removed, and animals were perfusion-fixed and processed for β-galactosidase histochemistry on 100 μm freezing microtome sections as described28. Anti-PECAM immunohistochemistry on embryos and yolk sacs was done as previously described25.
Flow cytometry of bone marrow, spleen and thymus.
Cell suspensions from bone marrow, spleen and thymus were made using standard methods29. Cells were resuspended at 5 × 105 cells/ml in BD Pharmingen FACS staining buffer, blocked with anti-mouse CD16/32 (BD Pharmingen), stained with the appropriate cocktail of antibodies (below) and fixed with BD Cytofix, all according to manufacturer's instructions. Final cell pellets were resuspended in 0.5 ml staining buffer and analyzed using BD FACS Calibur and BD CellQuest Pro software. The following antibody cocktails were used for bone marrow analysis: A, rat anti-mouse IgM-PE (Pharmingen)/anti-mouse CD45R(B220)-APC(Pharmingen); B, rat anti-mouse CD43(S7)-PE (Pharmingen)/anti-mouse CD45R(B220)-APC (Pharmingen). The following antibody cocktails were used for spleen and thymus analysis: A, rat anti-mouse IgM-PE (Pharmingen)/rat anti-mouse CD45R(B220)-APC (Pharmingen); B, rat anti-mouse CD49b(DX5)-PE (Pharmingen)/rat anti-mouseCD11b-APC (Pharmingen)C, rat anti-mouse CD132 (TUGm2)-PE (Pharmin''gen))/rat anti-mouse CD45R (B220)-APC (Pharmingen); D, rat anti-mouse CD4(L3T4)-PE (Pharmingen))/rat anti-mouse CD8a-APC (Pharmingen). All monoclonal antibodies were prepared in a mass dilution/cocktail. All antibodies were added to a final concentration of 0.5 mg/105 cells.
Note: Supplementary information is available on the Nature Biotechnology website.
References
Capecchi, M.R. Generating mice with targeted mutations. Nat. Med. 7, 1086–1090 (2001).
Evans, M.J. The cultural mouse. Nat. Med. 7, 1081–1083 (2001).
Goldstein, J.L. Laskers for 2001: knockout mice and test-tube babies. Nat. Med. 7, 1079–1080 (2001).
Smithies, O. Forty years with homologous recombination. Nat. Med. 7, 1083–1086 (2001).
Cecconi, F. & Gruss, P. From ES cells to mice: the gene trap approach. Methods Mol. Biol. 185, 335–346 (2002).
Chen, W.V. & Soriano, P. Gene trap mutagenesis in embryonic stem cells. Methods Enzymol. 365, 367–386 (2003).
Skarnes, W.C. et al. A public gene trap resource for mouse functional genomics. Nat. Genet. 36, 543–544 (2004).
Valenzuela, D.M. et al. High-throughput engineering of the mouse genome coupled with high-resolution expression analysis. Nat. Biotechnol. 21, 652–659 (2003).
Austin, C.P. et al. The knockout mouse project. Nat. Genet. 36, 921–924 (2004).
Auwerx, J. et al. The European dimension for the mouse genome mutagenesis program. Nat. Genet. 36, 925–927 (2004).
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J.C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl. Acad. Sci. USA 90, 8424–8428 (1993).
Eggan, K. et al. Hybrid vigor, fetal overgrowth, and viability of mice derived by nuclear cloning and tetraploid embryo complementation. Proc. Natl. Acad. Sci. USA 98, 6209–6214 (2001).
Eakin, G.S., Hadjantonakis, A.K., Papaioannou, V.E. & Behringer, R.R. Developmental potential and behavior of tetraploid cells in the mouse embryo. Dev Biol (2005).
Tokunaga, T. & Tsunoda, Y. Efficacious production of viable germ-line chimeras between embryonic stem (ES) cells and eight-cell stage embryos. Dev. Growth Differ. 34, 561–566 (1992).
Yagi, T. et al. A novel ES cell line, TT2, with high germline-differentiating potency. Anal. Biochem. 214, 70–76 (1993).
Papaioannou, V.E. Production of chimeras and genetically defined offspring from targeted ES-cells. in Gene Targeting: A Practical Approach (ed. Joyner, A.) 177–206, (Oxford University Press, Inc., New York, 1999).
McLaren, A. Chimaeras and sexual differentiation. in Chimaeras in Developmental Biology (eds. LeDouarin, N. & McLaren, A.) 381–399, (Academic Press, London, 1984).
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. . Manipulating the Mouse Embryo: a Laboratory Manual. 764 p. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 2003).
Powell-Braxton, L. et al. IGF-I is required for normal embryonic growth in mice. Genes Dev. 7, 2609–2617 (1993).
Noguchi, M. et al. Interleukin-2 receptor gamma chain: a functional component of the interleukin-7 receptor. Science 262, 1877–1880 (1993).
DiSanto, J.P., Muller, W., Guy-Grand, D., Fischer, A. & Rajewsky, K. Lymphoid development in mice with a targeted deletion of the interleukin 2 receptor gamma chain. Proc. Natl. Acad. Sci. USA 92, 377–381 (1995).
Cao, X. et al. Defective lymphoid development in mice lacking expression of the common cytokine receptor gamma chain. Immunity 2, 223–238 (1995).
Chen, J., Lansford, R., Stewart, V., Young, F. & Alt, F.W. RAG-2-deficient blastocyst complementation: an assay of gene function in lymphocyte development. Proc. Natl. Acad. Sci. USA 90, 4528–4532 (1993).
Duarte, A. et al. Dosage-sensitive requirement for mouse Dll4 in artery development. Genes Dev. 18, 2474–2478 (2004).
Gale, N.W. et al. Haploinsufficiency of delta-like 4 ligand results in embryonic lethality due to major defects in arterial and vascular development. Proc. Natl. Acad. Sci. USA 101, 15949–15954 (2004).
Auerbach, W. et al. Establishment and chimera analysis of 129/SvEv- and C57BL/6-derived mouse embryonic stem cell lines. Biotechniques 29, 1024–1028, 1030, 1032 (2000).
Yokoyama, T. et al. Conserved cysteine to serine mutation in tyrosinase is responsible for the classical albino mutation in laboratory mice. Nucleic Acids Res. 18, 7293–7298 (1990).
Adams, N.C. & Gale, Nicholas W. High resolution gene expression analysis in mice using genetically inserted reporter genes. in Mammalian and Avian Transgenesis–New Approaches (ed. Lois, S.P.a.C.) (Springer Verlag, Berlin Heidelberg, 2006).
Stevens, S. et al. Preparation of mouse cell suspensions. in Selected Methods inCellular Immunology (ed. Mishell, B., Shiigim SM) (W. H. Freeman and Company, New York, 1980).
Acknowledgements
We thank the VelociGene team for the targeted ES cells, Yingzi Xue for designing TaqMan assays, Keith Anderson for help with confocal imaging, and Ning Yuan, Yongli Chang, Kathleen Lincoln, Kim Cirillo and James Hiatt, for technical assistance. W.T.P. wants to thank John Poueymirou for his support. We would like to thank Nicole Graiff for help with the manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Most authors are employees of Regeneron Pharmaceuticals, Inc.
Supplementary information
Supplementary Fig. 1
8-cell embryo laser injection and comparison of ES cell contribution to the ICM in both 8-cell and blastocyst injections
Supplementary Table 1
Comparison of germ-line transmission in F0 mice from blastocyst and 8-cell embryo injections
Supplementary Table 2
F0 female mice with 100% ES cell-derived coat color can be obtained efficiently from 39,XO ES cells.
Supplementary Table 3
Germ-line Transmission of F0 Female Mice derived from 39,XO Targeted ES Clones.
Supplementary Video 1
Confocal sections through a blastocyst derived from a blastocyst injection
Supplementary Video 2
Confocal sections through a blastocyst derived from an 8-cell injection
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike license (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar license.
About this article
Cite this article
Poueymirou, W., Auerbach, W., Frendewey, D. et al. F0 generation mice fully derived from gene-targeted embryonic stem cells allowing immediate phenotypic analyses. Nat Biotechnol 25, 91–99 (2007). https://doi.org/10.1038/nbt1263
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1263
This article is cited by
-
Butyrophilin-like 2 regulates site-specific adaptations of intestinal γδ intraepithelial lymphocytes
Communications Biology (2021)
-
Functional biology of the Steel syndrome founder allele and evidence for clan genomics derivation of COL27A1 pathogenic alleles worldwide
European Journal of Human Genetics (2020)
-
GALECTIN-8 Is a Neuroprotective Factor in the Brain that Can Be Neutralized by Human Autoantibodies
Molecular Neurobiology (2019)