Abstract
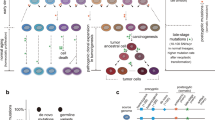
The power of genetics for the elucidation of gene function has been clearly established. In mammalian systems, the limited scope of available methods for the genetic manipulation of cultured cells has discouraged the widespread application of genetic approaches for the study of many current issues in molecular biology. Several recent technological developments hold out the promise of facilitating the manipulation of genes in mammalian cells. Three particular areas are emphasized in this review: 1) mutational analysis using nonsense suppression; 2) gene regulation by rapid, controlled amplification; and 3) gene targeting by homologous recombination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Horowitz, N.H. and Leupold, U. .1951. Some recent studies bearing on the one gene-one enzyme hypothesis. Cold Spring Harbor Symp. Quant. Biol. 16:65–72.
Campbell, A. .1961. Sensitive mutants of bacteriophage lambda. Virology 14:22–32.
Basilico, C. .1977. Temperature-sensitive mutations in animal Cells. Adv. Cancer Res. 24:223–266.
Marcus, M., Fainsod, A., and Diamond, G. 1985. The genetic analysis of mammalian Cell-cycle mutants. Ann. Rev. Genet. 19:389–421.
Dermody, J.J., Wojcik, B.E., Du, H., and Ozer, H.L. .1986. Identification of temperature-sensitive DNA− mutants of Chinese hamster Cells affected in Cellular and viral DNA synthesis. Mol. Cell. Biol. 6:4594–4601.
Benzer, S. and Champe, S.P. 1962. A change from nonsense to sense in the genetic code. Proc. Natl. Acad. Sci. USA 48:1114–1121.
Epstein, R.H., Bolle, A., Steinberg, C.M., Kellenberger, E. Boy de la Tour, E., Chevalley, R. Edgar, R.S., Slisman, M., Denhardt, G.H., and Lielausis, A. 1963. Physiological studies of conditional-lethal mutants of phage T4D. Cold Spring Harbor Symp. Quant. Biol. 28:375–392.
Celis, J.E. and Smith, J.D. 1979. Nonsense Mutations and tRNA Suppressors. Academic Press, London.
Miller, J.H., Coulondre, C., Hofer, M., Schmeissner, U., Sommer, H., and Schmitz, A. 1979. Genetic studies of the lac represser. IX. The generation of altered proteins by the suppression of nonsense mutations. J. Mol. Biol. 131:191–222.
Sedivy, J.M., Capone, J.P., RajBhandary, U.L., and Sharp, P.A. 1987. An inducible mammalian amber suppressor: propagation of a poliovirus mutant. Cell 50:379–389.
Laski, F.A., Belagaje, R., RajBhandary, U.L., and Sharp, P.A. 1982. An amber suppressor tRNA gene derived by site specific mutagenesis: cloning and expression in mammalian Cells. Proc. Natl. Acad. Sci. USA 79:5813–5817.
Temple, G.F., Dozy, A.M., Roy, K.L., and Kan, Y.W. 1982. Construction of a functional human suppressor tRNA gene: an approach to gene therapy of thalassemia. Nature 296:537–540.
Laski, F.A., Belagaje, R., Hudziak, R.M., Capecchi, M.R., Palese, P., RajBhandary, U.L., and Sharp, P.A. 1984. Synthesis of an ochre suppressor transfer RNA gene and expression in mammalian Cells. EMBO J. 3:2445–2452.
Capone, J.P., Sharp, P.A., and RajBhandary, U.L. 1985. Amber, ochre and opal suppressor tRNA genes derived from a human serine tRNA gene. EMBO J. 4:213–221.
Hudziak, R.M., Laski, F.A., RajBhandary, U.L., Sharp, P.A., and Capecchi, M.R. 1982. Establishment of mammalian Cell lines containing multiple nonsense mutations and functional suppressor transfer RNA genes. Cell 31:137–146.
Young, J.F., Capecchi, M.R., Laski, F.A., RajBhandary, U.L., Sharp, P.A., and Palese, P. 1983. Measurement of suppressor transfer RNA activity. Science 221:873–875.
Capone, J.P., Sedivy, J.M., Sharp, P.A., and RajBhandary, U.L. 1986. Introduction of UAG, UAA, and UGA nonsense mutations at a specific site in the Escherichia coli chloramphenicol acetyltransferase gene: use in measurement of amber, ochre, and opal suppression in mammalian Cells. Mol. Cell. Biol. 6:3059–3067.
DePamphilis, M.L. and Bradley, M.K. 1986. Replication of SV40 and polyoma virus chromosomes. In: The Papovaviridae, Volume 1. N. P. Salzman, (Ed.). Plenum Press, NY. p.99–246
Brockman, W.W. and Nathans, D. 1974. The isolation of simian virus 40 variants with specifically altered genomes. Proc. Natl. Acad. Sci. USA 71:942–946.
Tooze, J. 1980. The Molecular Biology of Tumor Viruses. DNA Tumor Viruses, 2nd ed. Cold Spring Harbor Laboratory, NY.
Rio, D.C., Clark, S.G., and Tjian, R. 1986. A mammalian host-vector system that regulates expression and amplification of transfected genes by temperature induction. Science 227:23–28.
Botchan, M., Topp, W., and Sambrook, J. 1979. Studies on simian virus 40 excision from Cellular chromosomes. Cold Spring Harbor Symp. Quant. Biol. 43:709–719.
Kern, F.G. and Basilico, C. 1986. An inducible eukaryotic host-vector expression system: amplification of genes under the control of the polyoma late promoter in a Cell line producing a thermolabile large T antigen. Gene 43:237–245.
Portela, A., Melero, J.A., de la Luna, S., and Ortin, J. 1986. Construction of Cell lines that regulate by temperature the amplification and expression of influenza virus non-structural protein genes. EMBO J. 5:2387–2392.
Kaufman, R.D. and Sharp, P.A. 1982. Amplification and expression of sequences cotransfected with a modular dihydrofolate reductase complementary DNA gene. J. Mol. Biol. 159:601–621.
Davie, E.W. and Fujikawa, K. 1975. Basic mechanisms in blood coagulation. Ann. Rev. Biochem. 44:799–829.
Barbacid, M. 1987. Ras genes. Ann. Rev. Biochem. 56:779–828.
Orr-Weaver, T.L., Szostak, J.W., and Rothstein, R.J. 1981. Yeast transformation: a model system for study of recombination. Proc. Natl. Acad. Sci. USA 78:6354–6358.
Rothstein, R. 1983. One-step gene disruption in yeast. Meth. Enzymol. 101:202–211.
Mammalian homologous recombination. 1984. In: Cold Spring Harbor Symp. Quant. Biol. 49:123–198.
Subramani, S. and Berg, P. 1983. Homologous and nonhomologous recombination in monkey Cells. Mol. Cell. Biol. 3:1040–1052.
Lin, F.-L., Sperle, K., and Sternberg, N. 1985. Recombination in mouse L Cells between DNA introduced into Cells and homologous chromosomal sequences. Proc. Natl. Acad. Sci. USA 82:1391–1395.
Smithies, O., Gregg, R.G., Boggs, S.S., Koralewski, M.A., and Kucherlapati, R.S. 1985. Insertion of DNA sequences into the human chromosomal β-globin locus by homologous recombination. Nature 317:230–234.
Thomas, K.R., and Capecchi, M.R. (1987). Site-directed mutagenesis by gene targeting in mouse embryo-derived stem Cells. Cell 51:503–512.
Smithies, O., Koralewski, M.A., Song, K.-Y., and Kucherlapati, R.S. 1984. Homologous recombination with DNA introduced into mammalian Cells. Cold Spring Harbor Symp. Quant. Biol. 48:161–170.
Thomas, K.R., Folger, K.R., and Capecchi, M.R. 1986. High frequency targeting of genes to specific sites in the mammalian genome. Cell 44:419–428.
Song, K.-Y., Schwartz, F., Maeda, N., Smithies, O., and Kucherlapati, R. 1987. Accurate modification of a chromosomal plasmid by homologous recombination in human Cells. Proc. Natl. Acad. Sci. USA 84:6820–6824.
Doetschman, T., Gregg, R.G., Maeda, N., Hooper, M.L., Melton, D.W., Thompson, S., and Smithies, O. 1987. Targeted correction of a mutant HPRT gene in mouse embryonic stem Cells. Nature 330:576–578.
Lin, F.-L., Sperle, K., and Sternberg, N. 1984. Model for homologous recombination during transfer of DNA into mouse L Cells: role for DNA ends in the recombination process. Mol. Cell. Biol. 4:1020–1034.
Kucherlapati, R.S., Eves, E.M., Song, K.-Y., Morse, B.S., and Smithies, O. 1984. Homologous recombination between plasmids in mammalian Cells can be enhanced by treatment of input DNA. Proc. Natl. Acad. Sci.USA 81:3153–3157.
Wake, C.T., Vernaleone, F., and Wilson, J.H. 1985. Topological requirements for homologous recombination among DNA molecules transfected into mammalian cells. Mol. Cell. Biol. 5:2080–2089.
Chang, X.-B. and Wilson, J.H. 1987. Modification of DNA ends can decrease end joining relative to homologous recombination in mammalian cells. Proc. Natl. Acad. Sci. USA 84:4959–4963.
Saiki, R.T., Gelfand, D.H., Stoffel, S., Scharf, S.J., Higuchi, R., Horn, G.T., Mullis, K.B., and Erlich, H.A. 1988. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239:487–491.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Sedivy, J. New Genetic Methods for Mammalian Cells. Nat Biotechnol 6, 1192–1196 (1988). https://doi.org/10.1038/nbt1088-1192
Issue Date:
DOI: https://doi.org/10.1038/nbt1088-1192
This article is cited by
-
DnaK-Mediated Alterations in Human Growth Hormone Protein Inclusion Bodies
Nature Biotechnology (1992)
-
A novel Vero cell line for use as a mammalian host-vector system in serum-free medium
Cytotechnology (1991)
-
Production of Soluble Recombinant Proteins in Bacteria
Nature Biotechnology (1989)