Abstract
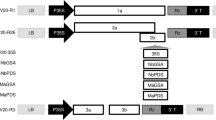
We have established a regeneration protocol for melon (Cucumis melo L.), using cotyledons as explants, in which the overall regeneration frequency reached more than 90 percent, and combined this protocol with Agrobacterium tumefaciens transformation to produce transgenic melon plants. Cotyledon explants were coculti-vated with A. tumefaciens carrying a disarmed Ti plasmid in which a dihydrofo-late reductase (DHFR) coding sequence and a β-glucuronidase (GUS) coding sequence were separately fused to the Cauliflower Mosaic Virus (CaMV) 35S promoter. A second strain carried a plasmid in which the neomycin phosphotransfer-ase II (NPTII) and GUS coding sequences were fused to a bidirectional CaMV 35S promoter. Under optimal conditions, the highest transformation frequency was approximately 6 percent. The transgenic melon plants were normal in morphology and fertile. Segregation analysis of R1 seedlings showed that the transgenes were inherited in a Mendelian manner. The CaMV 35S promoter was found to be active in most but not all tissues of transgenic melon plants, and the expression pattern conferred by the promoter changed during plant development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Trulson, A.J. and Shahin, E.A. 1986. In vitro plant regeneration in the genus Cucumis. Plant Science 47: 35–43.
Kim, S.-G., Chang, J.-R., Cha, H.C. and Lee, K.-W. 1988. Callus growth and plant regeneration in diverse cultivars of cucumber. Plant Cell, Tissue and Organ Culture 12: 67–74.
Srivastava, D.R., Andrianov, V.M. and Piruzian, E.S. 1989. Tissue culture and plant regeneration of watermelon (Citrullus vulgaris Schard. cv. Melitopolski). Plant Cell Reports 8: 300–302.
Niedz, R.P., Smith, S.S., Dunbar, K.B., Stephens, C.T. and Murakishi, H.H. 1989. Factors influencing shoot regeneration from cotyledonary explants of Cucumis melo. Plant Cell, Tissue and Organ Culture 18: 313–319.
Kathal, R., Bhatnagar, S.P. and Bhojwani, S.S. 1988. Regeneration of plants from leaf explants of Cucumis melo cv. Pusa Sharbati. Plant Cell Reports 7: 449–451.
Dirks, R. and van Buggenum, M. 1989. In vitro plant regeneration from leaf and cotyledon explants of Cucumis melo L. Plant Cell Reports 7: 626–627.
Moreno, V., Garcia-Sogo, M., Granell, I., Garcia-Sogo, B. and Roig, L.A. 1985. Plant regeneration from calli of melon (Cucumis melo L., cv. ‘Amarillo Oro’). Plant Cell, Tissue and Organ Culture 5: 139–146.
Trulson, A.J., Simpson, R.B. and Shahin, E.A. 1986. Transformation of cucumber (Cucumis sativus L.) plants with Agrobacterium rhizogenes. Theor. Appl. Genet. 73: 11–15.
Chee, P.P. 1990. Transformation of Cucumis sativus tissue by Agrobacterium tumefaciens and the regeneration of transformed plants. Plant Cell Reports 9: 245–248.
Ott, R.W., Ren, L. and Chua, N.-H. 1990. A bidirectional enhancer cloning vehicle for higher plants. Mol. Gen. Genet. 221: 121–124.
Jia, S.-R., Yang, M.-Z., Ott, R. and Chua, N-H. 1989. High frequency transformation of Kalanchoe laciniata. Plant Cell Reports 8: 336–340.
Koncz, C., Langridge, W.H.R., Olsson, O., Schell, J. and Szalay, A.A. 1990. Bacterial and firefly luciferase genes in transgenic plants: Advantages and disadvantages of a reporter gene. Developmental Genetics 11: 224–232.
deWet, J.R., Wood, K.V., Helsinki, D.R. and DeLuca, M. 1985. Cloning of firefly luciferase cDNA and expression of active luciferase in Escherichia coli. Proc. Natl. Acad. Sci. USA. 82: 7870–7873.
Jefferson, R.A. 1987. Assaying chimeric genes in plants: the β-glucuronidase as a versatile gene fusion marker in higher plants. EMBO J. 6: 3901–3907.
Benfey, P.N., Ren, L. and Chua, N.-H. 1989. The CaMV 35S enhancer contains at least two domains which can confer different developmental and tissue-specific expression patterns. EMBO J. 8: 2195–2202.
Benfey, P.N. and Chua, N.-H. 1989. Regulated genes in transgenic plants. Science 244: 174–181.
Odell, J.T., Nagy, F. and Chua, N.-H. 1985. Identification of DNA sequences required for activity of the cauliflower mosaic virus 35S promoter. Nature 313: 810–812.
Sanders, P.R., Winter, J.A., Zarnason, A.R., Rogers, S.G. and Fraley, R.T. 1987. Comparison of cauliflower mosaic virus 35S promoter and nopaline synthase promoters in transgenic plants. Nucleic Acid Res. 15: 1543–1558.
Kay, B., Chen, A., Daly, M. and McPherson, J. 1987. Duplication of CaMV 35S promoter sequences creates a strong enhancer for plant genes. Science 236: 1299–1302.
Murashige, T. and Skoog, F. 1962. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant. 15: 473–497.
Fraley, R.T., Rogers, S.G., Horsch, R.B., Eichholtz, D.A., Flick, C.L., Hoffman, N.I. and Saunders, P.R. 1985. The SEV system: a new disarmed Ti plasmid vecter system for plant transformation. Bio/Technology 3: 629–635.
Rogers, S.G., Horsch, R.B. and Fraley, R.T. 1986. Gene transfer in plants: Production of transformed plants using Ti plasmid vectors. Methods in Enzymol. 118: 627–640.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Dong, JZ., Yang, MZ., Jia, SR. et al. Transformation of Melon (Cucumis melo L.) and Expression from the Cauliflower Mosaic Virus 35S Promoter in Transgenic Melon Plants. Nat Biotechnol 9, 858–863 (1991). https://doi.org/10.1038/nbt0991-858
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0991-858
This article is cited by
-
Production of transgenic diploid Cucumis melo plants
Plant Cell, Tissue and Organ Culture (PCTOC) (2017)
-
Genetic Improvement of Sugarcane for Drought and Salinity Stress Tolerance Using Arabidopsis Vacuolar Pyrophosphatase (AVP1) Gene
Molecular Biotechnology (2014)
-
Agrobacterium-mediated transformation and shoot regeneration in elite breeding lines of western shipper cantaloupe and honeydew melons (Cucumis melo L.)
Plant Cell, Tissue and Organ Culture (PCTOC) (2012)
-
Agrobacterium-Mediated Genetic Transformation of Pogostemon cablin (Blanco) Benth. Using Leaf Explants: Bactericidal Effect of Leaf Extracts and Counteracting Strategies
Applied Biochemistry and Biotechnology (2012)
-
Histological study of organogenesis in Cucumis melo L. after genetic transformation: why is it difficult to obtain transgenic plants?
Plant Cell Reports (2011)