Abstract
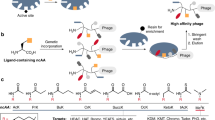
Besides natural peptide ligands, screening of random peptide libraries has yielded novel bioactive peptides for cell surface receptors. A method is described that uses a modified bacteriophage as a detection reagent to monitor the expression of receptor channels in mammalian cells and to probe the molecular interaction between phage-tethered peptides (ΦT-peptides) and specific receptor targets. By taking advantage of a specific multivalent interaction between ΦT-peptides and the receptor target, assays have been developed that use ΦT-peptides specific for the N-methyl-D-aspartate glutamate receptor, an important ligand-gated ion channel in the nervous system, to monitor the receptor expression in cultured mammalian cells. Combining these ΦT-peptide binding assays with fluorescence-activated cell sorting, 104 random glutamate receptor mutants were screened and candidate interaction residues were identified. This dual heterologous expression system offers a powerful approach to the molecular studies of protein-protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Smith, G.P. 1985. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228: 1315–1317.
Geysen, H.M., Rodda, S.J., and Mason, T.J. 1986. A priori delineation of a peptide which mimics a discontinuous antigenic determinant. Mol. Immunol. 23: 709–715.
Cwirla, S.E., Peters, E.A., Barrett, R.W., and Dower, W.J. 1990. Peptides on phage: a vast library of peptides for identifying ligands. Proc. Natl. Acad. Sci. USA 87: 6378–6382.
Devlin, J.J., Panganiban, L.C., and Devlin, P.E. 1990. Random peptide libraries: a source of specific protein binding molecules. Science 249: 404–406.
Scott, J.K. and Smith, G.P. 1990. Searching for peptide ligands with an epitope library. Science 249: 386–390.
Btond-Elguindi, S., Cwirla, S.E., Dower, W.J., Lipshutz, R.J., Sprang, S.R., Sambrook, J.F., et al. 1993. Affinity panning of a library of peptides displayed on bacteriophages reveals the binding specificity of BiP. Cell 75: 717–728.
Dedman, J.R., Kaetzel, M.A., Chan, H.C., Nelson, D.J., and Jamieson, G. Jr. 1993. Selection of targeted biological modifiers from a bacteriophage library of random peptides. The identification of novel calmodulin regulatory peptides. J. Biol. Chem. 268: 23025–23030.
Kay, B.K., Adey, N.B., He, Y.S., Manfredi, J.P., Mataragnon, A.H., and Fowlkes, D.M. 1993. An M13 phage library displaying random 38-amino-acid peptides as a source of novel sequences with affinity to selected targets. Gene 128: 59–65.
Sparks, A.B., Quilliam, L.A., Thorn, J.M., Der, C.J., and Kay, B.K. 1994. Identification and characterization of Src SH3 ligands from phage-displayed random peptide libraries. J. Biol. Chem. 269: 23853–23856.
Wrighton, N., Farrell, F., Chang, R., Kashyap, A., Barbone, F., Mulcahy, L., et al. 1996. Small peptides as potent mimetics of the protein hormone erythropoietin. Science 273: 458–463.
Li, M. Yu, W.F., Chen, C.H., Cwirla, S., Whitehorn, S., Tate, E., et al. 1996. In vitro selection of peptides acting at a new site of NMDA glutamate receptors. Nat. Biotechnol. 14: 986–991.
Whitehorn, E.A., Tate, E., Yanofsky, S.D., Kochersperger, L., Davis, A., Mortensen, R.B., et al. 1995. A generic method for expression and use of “tagged” soluble versions of cell surface receptors. Bio/technology 13: 1215–1219.
Cik, M., Chazot, P.L., and Stephenson, F.A. 1993. Optimal expression of cloned NMDAR1/NMDAR2A heteromeric glutamate receptors: a biochemical characterization. Biochem. J. 296: 877–83.
Rice, G.C., Goeddel, D.V, Cachianes, G., Woronicz, J., Chen, E.Y., Williams, S.R., et al. 1992. Random PCR mutagenesis screening of secreted proteins by direct expression in mammalian cells. Proc. Natl. Acad. Sci. USA 89: 5467–5471.
Barry, M.A., Dower, W.J., and Uohnston, S.A. 1996. Toward cell-targeting gene therapy vectors: selection of cell-binding peptides from random peptide-presenting phage libraries. Nat Med. 2: 299–305.
Pasqualini, R. and Ruoslahti, E. 1996. Organ targeting in vivo using phage display peptide libraries. Nature 380: 364–366.
Leung, D., Chen, E., and Goeddel, D. 1989. A method for random mutagenesis of a defined DNA segment using a modified porymerase chain reaction. Technique 1: 11–15.
Hirt, B. 1967. Selective extraction of polyoma DNA from infected mouse cell cultures. J. Mol. Biol. 26: 365–369.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, M. Use of a modified bacteriophage to probe the interactions between peptides and ion channel receptors in mammalian cells. Nat Biotechnol 15, 559–563 (1997). https://doi.org/10.1038/nbt0697-559
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0697-559
This article is cited by
-
The Kv1.3 K+ channel in the immune system and its “precision pharmacology” using peptide toxins
Biologia Futura (2021)
-
Central Nervous System Embryogenesis and Its Failures
Pediatric and Developmental Pathology (2002)
-
Applications of display technology in protein analysis
Nature Biotechnology (2000)