Abstract
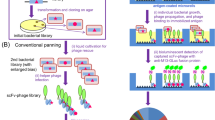
In vitro affinity maturation for evolving catalytic antibodies has been demonstrated by generating a diverse repertoire of the appropriate complementarity-determining regions on a phage surface. Phage display is followed by a selection based on binding to an altered antigen that was not used at the time of immunization, and provides variants with new catalytic activity and substrate specificity. This library format reduces the time needed to isolate the desired catalytic antibody fragments to under 2 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Lerner, A.R., Benkovic, S.J., and Schultz, P.G. 1991. At the crossroads of chemistry and immunology: Catalytic antibodies. Science 252: 659–667.
Schultz, P.G. and Lerner, R.A. 1995. From molecular diversity to catalysis: Lessons from the immune system. Science 269: 1835–1842.
Patten, P.A., Gray, N.S., Yang, P.L., Mark, C.B., Wedemayer, G.J., Boniface, J.J. et al. 1996 The immunological evolution of catalysis. Science 271: 1086–1091.
Hawkins, R.E., Russell, S.J., and Winter, G. 1992. Selection of phage antibodies by binding affinity mimicking affinity maturation. J. Med. Biol. 226: 889–896.
Barbas III, C.F., Bain, J.D., Hoekstra, D.M., and Lerner, R.A. 1992. Semisynthetic combinatorial antibody libraries: A chemical solution to the diversity problem. Proc. Natl. Acad. Sci. USA 89: 4457–4461.
Marks, J.D., Hoogenboom, H.H., Griffiths, A.D., and Winter, G. 1992. Molecular evolution of proteins on filamentous phage. J. Biol. Chem. 267: 16007–16010.
Jackson, J.R., Sathe, G., Rosenberg, A., and Sweet, R. 1995. In vitro antibody maturation. J. Immunol. 154: 3310–3319.
Barbas, C.F. and Burton, D.R. 1996. Selection and evolution of high-affinity human anti-viral antibodies. TIBTECH 14: 230–234.
Iwabuchi, Y., Miyashita, H., Tanimura, R., Kinoshita, K., Kikuchi, M., and Fujii, I. 1994. Regio- and stereoselective deprotection of acylated carbohydrates via catalytic antibodies. J. Am. Chem. Soc. 116: 771–772.
Schmidt, R.R. 1986. New methods for the synthesis of glycosides and oligosaccharides-Are there alternatives to the Koenigs-Knorr method?. Angew. Chem. Int. Ed. Engl. 25: 212–234, and references cited therein.
Jackson, D.Y., Prudent, J.R., Baldwin, E.P., and Schultz, P.G. 1991. A mutage-nesis study of a catalytic antibody. Proc. Natl. Acad Sci. USA 88: 58–62.
Roberts, V.A., Stewart, J., Benkovic, S.J., and Getzoff, E.D. 1994. Catalytic antibody model and mutagenesis implicate arginine in transition-state stabilization. J. Mol. Biol. 235: 1098–1116.
Miyashita, H., Hara, T., Tanimura, R., Fukurama, S., Cagnon, C., Kohara, A., and Fujii, I. 1997. Site-directed mutagenesis of active site contact residues in a hydrolytic abzyme: Evidence for an essential histidine involved in transition state stabilization. J. Mol. Biol. 267: 1247–1257.
Chothia, C. and Lesk, A. 1987. Canonical structures for the hypervariable regions of immunoglobulins. J. Mol Biol. 196: 901–917.
Chothia, C., Lesk, A.M., Tramontano, A., Levitt, M., Smith-Gill, S.J., Air, G. et al. 1989. Conformations of immunoglobulin hypervariable regions. Nature 342: 877–883.
Kabat, E.A., Wu, T.T., Perry, H.M., Gottes-man, K.S., and Foeller, C. 1991 Sequences of proteins of immunological interest, 5th ed. United States Department of Health and Human Services, National Institutes of Health, Bethesda, MD.
Barbas, C.F., Kang, A.S., Lerner, R.A., and Benkovic, S.J. 1991. Assembly of combinatorial antibody libraries on phage surfaces: the gene III site. Proc Natl. Acad. Sci. USA 88: 7978–7982.
Jacobs, J.W. 1991. New perspectives on catalytic antibodies. Bio/Technology 9: 258–262.
Stewart, J.D. and Benkovic, S.J. 1995. Transition-state stabilization as a measure of the efficiency of antibody catalysis. Nature 375: 388–391.
Fujii, I., Tanaka, F., Miyashita, H., Tanimura, R., and Kinoshita, K. 1995. Correlation between antigen-combining-site structures and functions within a panel of catalytic antibodies generated against a single transition state analog. J. Am. Chem. Soc. 117: 6199–6209.
Baca, M., Scanlan, T.S., Stephenson, R.C., and Wells, J.A. 1997. Phage display of a catalytic antibody to optimize for transition-state analog binding. Proc. Natl. Acad. Sci. USA 94: 10063–10068.
Zhou, G.W., Guo, J., Huang, W., Fletterick, R.J., and Scanlan, T.S. 1994. Crystal structure of a catalytic antibody with a serine protease active site. Science 265: 1059–1064.
Guo, J., Huang, W., and Scanlan, T.S. 1994. Kinetic and Mechanistic characterization of an efficient hydrolytic antibody: Evidence for the formation of an acyl intermediate. J. Am. Chem. Soc. 116: 6062–6069.
Murphy, D.J. 1995. Revisiting ground-state and transition-state effects, the split-site model, and the “fundamentalist position” of enzyme catalysis. Biochemistry 34: 4507–4510.
Wells, T.N.C. and Fersht, A.R. 1986. Use of binding energy in catalysis analyzed by mutagenesis of the tyrosyl-tRNA synthetase. Biochemistry 25: 1881–1886.
Fersht, A. 1985. Enzyme substrate complementarity and the use of binding energy in catalysis, pp. 324–327 in Enzyme structure and mechanism, 2nd ed. W.H. Freeman and Company, New York.
Wedemayer, G.J., Patten, P.A., Wang, L.H., Schultz, P.G., and Stevens, R.C. 1997. Structural insights into the evolution of an antibody combining site. Science 276: 1665–1669.
Sheriff, S., Silverton, E.W., Padlan, E.A., Cohen, G.H., Smith-Gill, S.J., Finzel, B.C., and Davies, D.R. 1987. Three-dimensional structure of an antibody-antigen complex. Proc. Natl. Acad. Sci. USA 84: 8075–8079.
Desmet, J., De Maeyer, M., Hazes, B., and Lasters, I. 1992. The dead-end elimination theorem and its use in protein side-chain positioning. Nature 356: 539–542.
Tanimura, R., Kidera, A., and Nakamura, H. 1994. Determinants of protein side-chain packing. Protein Sci. 3: 2358–2365.
Morikami, K., Nakai, T., Kidera, A., Saito, M., and Nakamura, H. 1992. PRESTO (Protein Engineering Simulation): a vectorized molecular mechanics program for biopolymers. Comp. Chem. 16: 243–248.
Friguet, B., Chaffotte, A.F., Djavadi-Ohaniance, L., and Goldberg, M.E. 1985. Measurements of the true affinity constant in solution of antigen-antibody complexes by enzyme-linked immunosorbent assay. J. Immunol. Methods 77: 305–319.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fujii, I., Fukuyama, S., Iwabuchi, Y. et al. Evolving catalytic antibodies in a phage-displayed combinatorial library. Nat Biotechnol 16, 463–467 (1998). https://doi.org/10.1038/nbt0598-463
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0598-463
This article is cited by
-
Phage display systems and their applications
Applied Microbiology and Biotechnology (2006)
-
In vitro abzyme evolution to optimize antibody recognition for catalysis
Nature Biotechnology (2001)
-
When blind is better: Protein design by evolution
Nature Biotechnology (1998)
-
Alchemy, enzymes, and the blind-watchmaker
Nature Biotechnology (1998)