Abstract
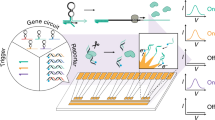
A prototype biosensor array has been assembled from engineered RNA molecular switches that undergo ribozyme-mediated self-cleavage when triggered by specific effectors. Each type of switch is prepared with a 5′-thiotriphosphate moiety that permits immobilization on gold to form individually addressable pixels. The ribozymes comprising each pixel become active only when presented with their corresponding effector, such that each type of switch serves as a specific analyte sensor. An addressed array created with seven different RNA switches was used to report the status of targets in complex mixtures containing metal ion, enzyme cofactor, metabolite, and drug analytes. The RNA switch array also was used to determine the phenotypes of Escherichia coli strains for adenylate cyclase function by detecting naturally produced 3′,5′- cyclic adenosine monophosphate (cAMP) in bacterial culture media.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schena, M., Shalon, D., Davis, R.W. & Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Lockhart, D.J. & Winzeler, E.A. Genomics, gene expression and DNA arrays. Nature 405, 827–836 (2000).
MacBeath, G. & Schreiber, S.L. Printing proteins as microarrays for high-throughput function determination. Science 289, 1760–1763 (2000).
de Wildt, R.M.T., Mundy, C.R., Gorick, B.D. & Tomlinson, I.M. Antibody arrays for high-throughput screening of antibody–antigen interactions. Nat. Biotechnol. 18, 989–994 (2000).
Zhu, H. et al. Analysis of yeast protein kinases using protein chips. Nat. Genet. 26, 283–289 (2000).
Pandey, A. & Mann, M. Proteomics to study genes and genomes. Nature 405, 837–846 (2000).
Marshall, K.A. & Ellington, A.D. Training ribozymes to switch. Nat. Struct. Biol. 6, 992–994 (1999).
Gold, L. Polisky, B. Uhlenbeck, O. & Yarus, M. Diversity of oligonucleotide functions. Annu. Rev. Biochem. 64, 763–797 (1995).
Osborne, S.E. & Ellington, A.D. Nucleic acid selection and the challenge of combinatorial chemistry. Chem. Rev. 97, 349–370 (1997).
Wilson, D.S. & Szostak, J.W. In vitro selection of functional nucleic aicds. Annu. Rev. Biochem. 68, 611–647 (1999).
Li, Y. & Breaker, R.R. Deoxyribozymes: new players in the ancient game of biocatalysis. Curr. Opin. Struct. Biol. 9, 315–323 (1999).
Symons, R.H. Small catalytic RNAs. Annu. Rev. Biochem. 61, 641–671 (1992).
Gesteland, R.F. Cech, T.R. & Atkins, J.F. (eds) The RNA world (Cold Spring Harbor LaboratoryPress, Cold Spring Harbor, NY; 1999).
Soukup, G.A. & Breaker, R.R. Nucleic acid molecular switches. Trends Biotechnol. 17, 469–476 (1999).
Soukup, G.A. & Breaker, R.R. Allosteric nucleic acid catalysts. Curr. Opin. Struct. Biol. 10, 318–325 (2000).
Tang, J. & Breaker, R.R. Rational design of allosteric ribozymes. Chem. Biol. 4, 453–459 (1997).
Tang, J. & Breaker, R.R. Mechanism for allosteric inhibition of an ATP-sensitive ribozyme. Nucleic Acids Res. 26, 4214–4221 (1998).
Soukup, G.A. & Breaker, R.R. Design of allosteric hammerhead ribozymes activated by ligand-induced structure stabilization. Structure 7, 783–791 (1999).
Breaker, R.R. In vitro selection of catalytic polynucleotides. Chem. Rev. 97, 371–390 (1997).
Robertson, M.P. & Ellington, A.D. Design and optimization of effector-activated ribozyme ligases. Nucleic Acids Res. 28, 1751–1759 (2000).
Koizumi, M., Soukup, G.A. Kerr, J.Q. & Breaker, R.R. Allosteric selection of ribozymes that respond to the second messengers cGMP and cAMP. Nat. Struct. Biol. 6, 1062–1071 (1999).
Soukup, G.A., DeRose, E.C. & Breaker, R.R. Generating new ligand-binding RNAs by affinity maturation and disintegration of allosteric ribozymes. RNA, in press.
Soukup, G.A. & Breaker, R.R. Engineering precision RNA molecular switches. Proc. Natl. Acad. Sci. USA 96, 3584–3589 (1999).
Araki, M., Okuno, Y., Hara, Y. & Sugiura, Y. Allosteric regulation of a ribozyme activity through ligand-induced conformational change. Nucleic Acids Res. 26, 3379–3384 (1998).
Soukup, G.A., Emilsson, G.A.M. & Breaker, R.R. Altering molecular recognition of RNA aptamers by allosteric selection. J. Mol. Biol. 298, 623–632 (2000).
Wang, D.Y. & Sen D. Rationally designed allosteric variants of hammerhead ribozymes responsive to the HIV-1 Tat protein. J. Mol. Biol., in press.
Porta, H. & Lizardi, P.M. An allosteric hammerhead ribozyme. Biotechnology 13, 161–164 (1995).
Kuwabara, T. et al. A novel allosterically trans-activated ribozyme, the maxizyme, with exceptional specificity in vitro and in vivo. Mol. Cell 2, 617–627 (1998).
Robertson, M.P. & Ellington, A.D. In vitro selection of an allosteric ribozyme that transduces analytes to amplicons. Nat. Biotechnol. 17, 62–66 (1999).
Komatsu, Y., Yamashita, S., Kazama, N., Nobuoka, K. & Ohtsuka, E. Construction of new ribozymes requiring short regulator oligonucleotides as a cofactor. J. Mol. Biol. 299, 1231–1243 (2000).
Fedor, M.J. & Uhlenbeck, O.C. Kinetics of intermolecular cleavage by hammerhead ribozymes. Biochemistry 31, 12042–12054 (1992).
Li., Y. & Breaker, R.R. Kinetics of RNA degradation by specific base catalysis of transesterification involving the 2′-hydroxyl group. J. Am. Chem. Soc. 121, 5364–5372 (1999).
Herne, T.M. & Tarlov, M.J. Characterization of DNA probes immobilized on gold surfaces. J. Am. Chem. Soc. 119, 8916–8920 (1997).
Guisan, J.M., Melo, F.V. & Ballesteros, A. Determination of intrinsic properties of immobilized enzymes. 1. Kinetic studies on sepharose–staphylococcal nuclease in the absence of diffusional limitations. Appl. Biochem. Biotechnol. 6, 25 (1981).
Guisan, J.M., Melo, F.V. & Ballesteros, A. Determination of intrinsic properties of immobilized enzymes. 2. Kinetic studies on sepharose–staphylococcal nuclease in the presence of diffusional limitations. Appl. Biochem. Biotechnol. 6, 37 (1981).
Jose, A.M., Soukup, G.A. & Breaker, R.R. Cooperative binding of effectors by an allosteric ribozyme. Nucleic Acids Res., in press.
Potter, K., Chaloner-Larsson, G. & Yamazaki, H. Abnormally high rate of cAMP excretion from an Escherichia coli mutant deficient in cyclic AMP receptor protein. Biochem. Biophys. Res. Commun. 57, 379–385 (1974).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular cloning: a laboratory manual. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY; 1989).
Sabourin, D. & Beckwith, J. Deletion of the Escherichia coli crp gene. J. Bacteriol. 122, 338–340 (1975).
Shah, S. & Peterkofsky, A. Characterization and generation of Escherichia coli adenylate cyclase deletion mutants. J. Bacteriol. 173, 3238–3242 (1991).
Tang, J. & Breaker, R.R. Structural diversity of self-cleaving ribozymes. Proc. Natl. Acad. Sci. USA 97, 5784–5789 (2000).
Hertel, K.J. et al. Numbering system for the hammerhead. Nucleic Acids Res. 20, 3252 (1992).
Acknowledgements
We are grateful to Mark Reed and James Klemic for the production of gold-coated surfaces, and Michael Tarlov for helpful suggestions regarding gold–thiol immobilization. Funding for this work was provided by grants from the National Institutes of Health, the Defense Advanced Research Projects Agency (DARPA), and the Yale Diabetes and Endocrinology Research Center. R.R.B. is the recipient of a Hellman Family Fellowship and a fellowship from the David and Lucile Packard Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Seetharaman, S., Zivarts, M., Sudarsan, N. et al. Immobilized RNA switches for the analysis of complex chemical and biological mixtures. Nat Biotechnol 19, 336–341 (2001). https://doi.org/10.1038/86723
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/86723
This article is cited by
-
Identifying the preferred RNA motifs and chemotypes that interact by probing millions of combinations
Nature Communications (2012)
-
Engineering ligand-responsive gene-control elements: lessons learned from natural riboswitches
Gene Therapy (2009)
-
Recent advances of aptamer sensors
Science in China Series B: Chemistry (2008)
-
Study of thiazole orange in aptamer-based dye-displacement assays
Analytical and Bioanalytical Chemistry (2008)
-
Aptamers as molecular recognition elements for electrical nanobiosensors
Analytical and Bioanalytical Chemistry (2008)