Abstract
The neurovascular unit (NVU) regulates metabolic homeostasis as well as drug pharmacokinetics and pharmacodynamics in the central nervous system. Metabolic fluxes and conversions over the NVU rely on interactions between brain microvascular endothelium, perivascular pericytes, astrocytes and neurons, making it difficult to identify the contributions of each cell type. Here we model the human NVU using microfluidic organ chips, allowing analysis of the roles of individual cell types in NVU functions. Three coupled chips model influx across the blood–brain barrier (BBB), the brain parenchymal compartment and efflux across the BBB. We used this linked system to mimic the effect of intravascular administration of the psychoactive drug methamphetamine and to identify previously unknown metabolic coupling between the BBB and neurons. Thus, the NVU system offers an in vitro approach for probing transport, efficacy, mechanism of action and toxicity of neuroactive drugs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Banks, W.A. From blood-brain barrier to blood-brain interface: new opportunities for CNS drug delivery. Nat. Rev. Drug Discov. 15, 275–292 (2016).
Itoh, Y. & Suzuki, N. Control of brain capillary blood flow. J. Cereb. Blood Flow Metab. 32, 1167–1176 (2012).
Magistretti, P.J. & Allaman, I. A cellular perspective on brain energy metabolism and functional imaging. Neuron 86, 883–901 (2015).
Lecrux, C. & Hamel, E. The neurovascular unit in brain function and disease. Acta Physiol. (Oxf.) 203, 47–59 (2011).
Vasilopoulou, C.G., Margarity, M. & Klapa, M.I. Metabolomic analysis in brain research: opportunities and challenges. Front. Physiol. 7, 183 (2016).
Warren, M.S. et al. Comparative gene expression profiles of ABC transporters in brain microvessel endothelial cells and brain in five species including human. Pharmacol. Res. 59, 404–413 (2009).
Naik, P. & Cucullo, L. In vitro blood-brain barrier models: current and perspective technologies. J. Pharm. Sci. 101, 1337–1354 (2012).
Booth, R. & Kim, H. Characterization of a microfluidic in vitro model of the blood-brain barrier (μBBB). Lab Chip 12, 1784–1792 (2012).
Prabhakarpandian, B. et al. SyM-BBB: a microfluidic blood brain barrier model. Lab Chip 13, 1093–1101 (2013).
Griep, L.M. et al. BBB on chip: microfluidic platform to mechanically and biochemically modulate blood-brain barrier function. Biomed. Microdevices 15, 145–150 (2013).
Brown, J.A. et al. Recreating blood-brain barrier physiology and structure on chip: A novel neurovascular microfluidic bioreactor. Biomicrofluidics 9, 054124 (2015).
Brown, J.A. et al. Metabolic consequences of inflammatory disruption of the blood-brain barrier in an organ-on-chip model of the human neurovascular unit. J. Neuroinflammation 13, 306 (2016).
Ballabh, P., Braun, A. & Nedergaard, M. The blood-brain barrier: an overview: structure, regulation, and clinical implications. Neurobiol. Dis. 16, 1–13 (2004).
Banks, W.A. et al. Passage of amyloid beta protein antibody across the blood-brain barrier in a mouse model of Alzheimer's disease. Peptides 23, 2223–2226 (2002).
Easton, A.S. & Fraser, P.A. Variable restriction of albumin diffusion across inflamed cerebral microvessels of the anaesthetized rat. J. Physiol. (Lond.) 475, 147–157 (1994).
Zhao, Z., Nelson, A.R., Betsholtz, C. & Zlokovic, B.V. Establishment and dysfunction of the blood-brain barrier. Cell 163, 1064–1078 (2015).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Jacques, P.F. et al. Relation between folate status, a common mutation in methylenetetrahydrofolate reductase, and plasma homocysteine concentrations. Circulation 93, 7–9 (1996).
Ridley, A.J. Rho GTPases and cell migration. J. Cell Sci. 114, 2713–2722 (2001).
Bonneh-Barkay, D. & Wiley, C.A. Brain extracellular matrix in neurodegeneration. Brain Pathol. 19, 573–585 (2009).
Zamanian, J.L. et al. Genomic analysis of reactive astrogliosis. J. Neurosci. 32, 6391–6410 (2012).
Frey, D., Laux, T., Xu, L., Schneider, C. & Caroni, P. Shared and unique roles of CAP23 and GAP43 in actin regulation, neurite outgrowth, and anatomical plasticity. J. Cell Biol. 149, 1443–1454 (2000).
Northrop, N.A. & Yamamoto, B.K. Methamphetamine effects on blood-brain barrier structure and function. Front. Neurosci. 9, 69 (2015).
Turowski, P. & Kenny, B.A. The blood-brain barrier and methamphetamine: open sesame? Front. Neurosci. 9, 156 (2015).
Zhang, X., Banerjee, A., Banks, W.A. & Ercal, N. N-acetylcysteine amide protects against methamphetamine-induced oxidative stress and neurotoxicity in immortalized human brain endothelial cells. Brain Res. 1275, 87–95 (2009).
Fujimoto, Y. et al. The pharmacokinetic properties of methamphetamine in rats with previous repeated exposure to methamphetamine: the differences between Long-Evans and Wistar rats. Exp. Anim. 56, 119–129 (2007).
Frankel, P.S., Alburges, M.E., Bush, L., Hanson, G.R. & Kish, S.J. Brain levels of neuropeptides in human chronic methamphetamine users. Neuropharmacology 53, 447–454 (2007).
Bélanger, M., Allaman, I. & Magistretti, P.J. Brain energy metabolism: focus on astrocyte-neuron metabolic cooperation. Cell Metab. 14, 724–738 (2011).
Wishart, D.S. et al. The human cerebrospinal fluid metabolome. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 871, 164–173 (2008).
Jaeger, C. et al. The mouse brain metabolome: region-specific signatures and response to excitotoxic neuronal injury. Am. J. Pathol. 185, 1699–1712 (2015).
Berman, S.M. et al. Changes in cerebral glucose metabolism during early abstinence from chronic methamphetamine abuse. Mol. Psychiatry 13, 897–908 (2008).
Zheng, T. et al. The metabolic impact of methamphetamine on the systemic metabolism of rats and potential markers of methamphetamine abuse. Mol. Biosyst. 10, 1968–1977 (2014).
Vargas, M.R., Pehar, M., Cassina, P., Beckman, J.S. & Barbeito, L. Increased glutathione biosynthesis by Nrf2 activation in astrocytes prevents p75NTR-dependent motor neuron apoptosis. J. Neurochem. 97, 687–696 (2006).
Zanelli, S.A., Solenski, N.J., Rosenthal, R.E. & Fiskum, G. Mechanisms of ischemic neuroprotection by acetyl-L-carnitine. Ann. NY Acad. Sci. 1053, 153–161 (2005).
Verleysdonk, S., Martin, H., Willker, W., Leibfritz, D. & Hamprecht, B. Rapid uptake and degradation of glycine by astroglial cells in culture: synthesis and release of serine and lactate. Glia 27, 239–248 (1999).
Fredholm, B.B., Chen, J.F., Cunha, R.A., Svenningsson, P. & Vaugeois, J.M. Adenosine and brain function. Int. Rev. Neurobiol. 63, 191–270 (2005).
Schousboe, A. & Sonnewald, U. The Glutamate/GABA-Glutamine Cycle (Springer, 2016).
Culic, O., Gruwel, M.L. & Schrader, J. Energy turnover of vascular endothelial cells. Am. J. Physiol. 273, C205–C213 (1997).
Turner, D.A. & Adamson, D.C. Neuronal-astrocyte metabolic interactions: understanding the transition into abnormal astrocytoma metabolism. J. Neuropathol. Exp. Neurol. 70, 167–176 (2011).
Bröer, S. & Brookes, N. Transfer of glutamine between astrocytes and neurons. J. Neurochem. 77, 705–719 (2001).
Wu, G., Haynes, T.E., Li, H. & Meininger, C.J. Glutamine metabolism in endothelial cells: ornithine synthesis from glutamine via pyrroline-5-carboxylate synthase. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 126, 115–123 (2000).
Bhatia, S.N. & Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 32, 760–772 (2014).
Prantil-Baun, R. et al. Physiologically based pharmacokinetic and pharmacodynamic analysis enabled by microfluidically linked organs-on-chips. Annu. Rev. Pharmacol. Toxicol. 58, 37–64 (2018).
Uchida, Y. et al. Quantitative targeted absolute proteomics of human blood-brain barrier transporters and receptors. J. Neurochem. 117, 333–345 (2011).
Hawrylycz, M.J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Malik, N. et al. Comparison of the gene expression profiles of human fetal cortical astrocytes with pluripotent stem cell derived neural stem cells identifies human astrocyte markers and signaling pathways and transcription factors active in human astrocytes. PLoS One 9, e96139 (2014).
Urich, E., Lazic, S.E., Molnos, J., Wells, I. & Freskgård, P.O. Transcriptional profiling of human brain endothelial cells reveals key properties crucial for predictive in vitro blood-brain barrier models. PLoS One 7, e38149 (2012).
Procaccini, C. et al. Role of metabolism in neurodegenerative disorders. Metabolism 65, 1376–1390 (2016).
Sims, N.R. & Muyderman, H. Mitochondria, oxidative metabolism and cell death in stroke. Biochim. Biophys. Acta 1802, 80–91 (2010).
Giza, C.C. & Hovda, D.A. The neurometabolic cascade of concussion. J. Athl. Train. 36, 228–235 (2001).
Li, S. et al. Predicting network activity from high throughput metabolomics. PLOS Comput. Biol. 9, e1003123 (2013).
Li, S. et al. Constructing a fish metabolic network model. Genome Biol. 11, R115 (2010).
The, M., MacCoss, M.J., Noble, W.S. & Käll, L. Fast and accurate protein false discovery rates on large-scale proteomics data sets with percolator 3.0. J. Am. Soc. Mass Spectrom. 27, 1719–1727 (2016).
Eichler, G.S., Huang, S. & Ingber, D.E. Gene Expression Dynamics Inspector (GEDI): for integrative analysis of expression profiles. Bioinformatics 19, 2321–2322 (2003).
Liebermeister, W. et al. Visual account of protein investment in cellular functions. Proc. Natl. Acad. Sci. USA 111, 8488–8493 (2014).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Szklarczyk, D. et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Orth, J.D., Thiele, I. & Palsson, B.O. What is flux balance analysis? Nat. Biotechnol. 28, 245–248 (2010).
Varma, A. & Palsson, B.O. Metabolic capabilities of Escherichia coli: I. synthesis of biosynthetic precursors and cofactors. J. Theor. Biol. 165, 477–502 (1993).
Lewis, N.E. et al. Large-scale in silico modeling of metabolic interactions between cell types in the human brain. Nat. Biotechnol. 28, 1279–1285 (2010).
Duarte, N.C. et al. Global reconstruction of the human metabolic network based on genomic and bibliomic data. Proc. Natl. Acad. Sci. USA 104, 1777–1782 (2007).
Schellenberger, J. et al. Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox v2.0. Nat. Protoc. 6, 1290–1307 (2011).
Groves, P.M. & Rebec, G.V. Introduction to Biological Psychology (W.C. Brown, 1988).
Acknowledgements
This research was supported by the Wyss Institute for Biologically Inspired Engineering at Harvard University, Defense Advanced Research Projects Agency (DARPA) under Cooperative Agreement Number W911NF-12-2-0036 (D.E.I. & K.K.P.), and Sverige-Amerika Stiftelsen, Carl Trygger Stiftelse, Erik och Edith Fernstrom's stiftelse (A.H.). The views and conclusions contained in this document are those of the authors and should not be interpreted as representing the official policies, either expressed or implied, of DARPA or the US Government. This work also was made possible by access to the microfabrication facilities of the Center for Nanoscale Systems (CNS), a member of the National Nanotechnology Coordinated Infrastructure Network (NNCI), which is supported by the National Science Foundation under NSF award no. 1541959. CNS is part of Harvard University. We also thank T. Ferrante for technical assistance, M. Rosnach and J.P. Ferrier for artwork and technical illustrations, to J.A. Goss for his help with chips design and fabrication, and the Harvard Medical School Neurobiology Imaging Facility (supported in part by NINDS P30 Core Center grant #NS072030) for consultation and instrument availability.
Author information
Authors and Affiliations
Contributions
B.M.M., A.H., E.A.F., T.G., D.E.I. and K.K.P. designed the study. C.V. ran and analyzed the MS samples, and A.R.P. and D.S. performed the flux balance analysis modeling. S.P.S. performed bioinformatic analysis for proteomics and MS. A.H., E.A.F. and T.-E.P. conducted the BBB chip culture and B.M.M., T.G. and s.d. performed brain chip culture and imaging. B.M.M., A.H., E.A.F., T.G., S.D. and R.M. performed confocal imaging, N.B. and B.B. conducted proteomic run and analysis. K.S. performed the COMSOL modeling, B.M.M., T.G. and A.C. conducted sample preparation for brain chip. B.M.M., T.G. and J.C.N. contributed to the brain chip design. B.M.M., A.H., E.A.F., T.G., D.E.I. and K.K.P. prepared illustrations and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.E.I. holds equity in Emulate, Inc., consults for the company, and chairs its scientific advisory board. K.K.P. is a consultant and a member of the Scientific Advisory Board of Emulate, Inc.
Integrated supplementary information
Supplementary Figure 1 Diagram of BBB and brain chips.
(a) Schematic of the BBB Chip demonstrates the 3 parts of the chip, Top PDMS channel, membrane and Bottom PDMS channel; (b) Image of 2 BBB Chips, the red color represents the bottom (Endo) channel while the blue represents the top (Pericytes/Astrocytes) channel, (c) Schematic of the Brain Chip demonstrates the different parts of the chip, (from top to bottom) the manifold which hold the chip and enables perfusion of the chip, Brain Chip, PDMS gasket, Topas substrate and PC base which hold the chip; (d) Image of Brain Chips, top tube represents the inlet and outlet of the Brain Chips. The orange triangles represent the location of the human neuronal culture. (e) The diagram represents the experimental setup. The red line represents the endothelial media (aBlood) whereas the blue line represents the neuronal media (aCSF).
Supplementary Figure 2 Immunohistochemical characterization of human neural cell cultures in the brain chip and the neurovascular cells in the BBB chip.
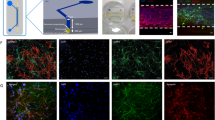
(a, b) Confocal images of astrocytes stained for glial fibrillary acidic protein (GFAP, red), neurons stained for β-III-tubulin (βIII, green) and cell nuclei stained with 4′,6-diamidino-2phenylindole (DAPI, blue). (c, d) Confocal images of neurons (βIII, green), synapses (synaptophysin, red) and cell nuclei (DAPI, blue). (e, f) Confocal images of dopaminergic neurons stained for tyrosine hydroxylase (TH, green), astrocytes (GFAP, red) and cell nuclei (DAPI, blue). (g, h) Confocal images of neurons (βIII, green), GABAergic neurons stained for vesicular gamma-aminobutiric acid transporter (VGAT, red) and cell nuclei (DAPI, blue). (i, j) Confocal micrographs of neurons (βIII, green), glutamatergic neurons stained for glutamate decarboxylase (GAD, red) and GABAergic neurons (VGAT, blue). (a, c, e, g, i) scale bars = 3 mm; (b, d, f, h, j) scale bars = 100 μm. (k, l) Demonstration of different cell types that comprise the BBB: (k) Endothelial cells (ZO-1, green, nuclei, blue), (l) Pericytes (Phalloidin, green) and Astrocytes (GFAP, red) and nuclei, blue. Scale bar = 50 μm.(m) Endothelial cells (Occludin, green, nuclei, blue), (n) Endothelial cells (Claudin-5, green, nuclei, blue). More than 10 cultures have been replicated and characterized with immunofluorescence for each condition with similar results.
Supplementary Figure 3 Flow velocity in the model system
(a) Simulation of flow distribution in the coupled chip setup (COMSOL®). Model illustrates precise control of fluidic flow using our microfluidic platform to accurately mimic the differential flow velocities found throughout the BBB-Brain-BBB Chips. Flow distribution confirms our coupled system follows laminar flow with lower velocities applied to the apical channels of the BBB Chip and Brain Chip with a higher velocity found in the basal or vascular component of the BBB Chip.
Supplementary Figure 4 Permeability measurements in the empty BBB-brain-BBB system.
(a) Mean fluorescence intensity measurement of cascade blue-containing medium at different points of the system: vascular component of influx BBB, inlet of Brain Chip, inlet of the brain side of efflux BBB and outlet of the vascular side of efflux BBB. Values are presented as percentage of cascade blue (CB) fluorescence intensity of medium flowing into the system at the inlet vascular part of the influx BBB. N=5 for Vessel 1 and N= 3 for Perivasc.1, Brain and Vessel 2, N representing independent NVU systems, (p values: SI Notes, Supplementary Table 1b.) (b) Mean fluorescence intensity measurement of Alexa-555 labeled BSA (BSA-555) at different points throughout the system. Values are presented as percentage of the BSA-555 intensity values of the medium flowing into the system at the inlet vascular component of the influx BBB microfluidic device (p values: SI Notes, Supplementary Table 1b.). Error bars are SEM, N=5 for Vessel 1 and N= 3 for Perivasc.1, Brain and Vessel 2, significance calculated with one-way ANOVA, Bonferroni post-test.
Supplementary Figure 5 Protein cluster density maps for GEDI Figure 2 and breakdown of the biological processes associated with NVU compartment proteomes.
(a) Mosaics representing the number of proteins that were assigned to each cluster (i.e. tile) in the mosaic after SOM training. These mosaics illustrate the distribution of proteins across the global expression profile GEDI maps shown in Fig. 2. The data are reduced dimensionality by classifying proteins with similar expression profiles into discrete groups that are organized into distinct two-dimensional mosaics. Each tile of these mosaics represents a cluster of proteins and the mapping of proteins to these tiles is conserved across samples to facilitate comparison. Thus, the tiles represent the amount of proteins that have the same expression profile, ranging from low numbers illustrated in blue and high numbers in red. The protein expression for each of the NVU compartments (Endo, Peri/Astro and Neurons) was compared between the uncoupled and coupled conditions. (b). Proteomaps illustrate the KEGG Orthology biological process terms associated with low and high abundance proteins observed in the expression profiles of fluidically uncoupled chips compartments of endothelial cells, pericyte/astrocyte and neuronal/astrocyte cultures contrasted with expression values observed in coupled BBB-Brain-BBB Chips. In these Voronoi diagrams, each individual polygon represents one protein, wherein polygon area is a function of mass abundance. Each polygon is color-coded according to the KEGG Orthology term associated with it, and polygons representing the same KEGG Orthology term are clustered into larger polygons to form the map (Supplementary Video 3, 4, 5, 6, 7, 8). The percentage of the protein expression profiles represented by each KEGG Orthology term shown in the Proteomaps is presented in Fig. 2b. (c) Principle component analysis (PCA) was used to assess the variability in global protein expression observed in the coupled and uncoupled cell populations for each compartment of the BBB and Brain Chips N=3 representing independent NVU systems.
Supplementary Figure 6 GluR2 penetration across the barrier after Meth addition, schematic of experiment shown in Figure 3.
(a) The BBB Chipinflux recapitulates tight barrier preventing penetration of an antibody (anti-GluR2, pink y-shaped objects) from the vascular to the perivascular compartment during control conditions. (b) 24 hrs of Meth addition compromises the barrier, allowing the anti-GluR2 to stain the neurons. Due to the low concentration of Meth reaching the BBB Chipefflux, the barrier stayed intact. (c) 24 hrs after Meth withdrawal resulted in recovery of the Chipinflux while anti-GluR2 remains attached to the neurons.
Supplementary Figure 7 Effect of Meth on the barrier properties of the BBB.
(a-b) Meth dose response of TEER values for a BBB Transwell model. Mean values are represented as percentage of control (untreated well). (a) Meth concentrations were applied 24hrs and (b) recovery for 24 hrs after Meth removal. Error bars are SEM, N=2. (c) Effect of Meth on neuronal viability in the Brain Chip. Cell viability of neurons treated with 1 mM Meth. Dead cells were stained with ethidium homodimer while all cells were stained with DAPI and images were analyzed with a custom MATLAB® script. Results are presented as percentage of live and dead cells, (p values: for each group, the p values for the live vs. dead was <0.0001, SI Notes, Supplementary Table 1b.). Error bars are SEM, N=3 neuronal cultures, were 17–24 field of views were taken for each culture, in order to calculate the mean value for each setup. significance calculated with one-way ANOVA, Bonferroni post-test.
Supplementary Figure 8 Mass balance of Meth in the coupled NVU microfluidic system and Meth uptake by cells in well plates.
(a) Characterization of Meth dilution in BBB-Brain-BBB Chips. Mass spectrometry analysis of the concentration of Meth in different compartments throughout the system: vascular component of the influx BBB (Vessel 1), inlet of the Brain Chip (Perivasc. 1), outlet of the CSF (Brain) and vascular part of the efflux BBB (Vessel 2). Mean values are represented as percentage of the Meth dose flowed into the system (vascular part of the influx BBB), (p values <0.0001 SI Notes, Supplementary Table 1b.). N=3 representing independent NVU systems, significance calculated with one-way ANOVA, Bonferroni post-test (b) Mean values of Meth uptake by the neurovascular cell types in well-plates after 24hrs incubation N=3 representing independent NVU systems.
Supplementary Figure 9 Perivasculature response to Meth.
Immunofluorescence microscopic views of the different cell populations within the influx and efflux BBB Chip in the fluidically coupled NVU system. Perivascular cells stained with astrocyte specific glial fibrillary associated protein (GFAP) and general F-actin staining with Phalloidin demonstrated consistent morphology in the influx and efflux BBB. The cell morphology was compared between steady state (a, b), addition of Meth (c, d), and drug removal (e, f). Administration of Meth for 24 hrs demonstrated only a minor change in perivascular cell morphology of mostly GFAP-negative cells (pericytes and GFAP-negative astrocytes) in the influx BBB (c), whereas efflux BBB maintained the control morphology (d). 24 hrs recovery restored the morphology of the influx BBB Chip (e) which is now identical to the efflux BBB (f). The experiment represented by the immunofluorescence micrographs was repeated 3 times with 2–4 individual NVU systems for each repeat.
Supplementary Figure 10 Untargeted metabolism.
Potential biochemical pathways associated with significant metabolic changes identified by MS analysis (Compound Discoverer) within cells in the vascular compartment of the influx or efflux BBB Chips (Vessel 1 or 2, respectively), perivascular compartments of the influx or efflux BBB Chips (Perivasc 1 or 2, respectively), or the lower compartment of the Brain Chip (Brain) within the coupled NVU system under control conditions (b) or exposed to Meth for 24 hrs (c). Vessel 1 (Blue), Vessel 2 (Magenta), Perivasc 1 (Green), Perivasc 2 (Yellow) and Brain (Black). (c) IPA was used to identify the significant “Disease Pathways” together with “Biofunction” and their regulation which change due to Meth administration, using Z-score, (Z-score scale: red, high Z-score, pathway is upregulated, blue, low Z-score, pathway is downregulated)
Supplementary Figure 11 Metabolic cycle of labelled glucose.
(a) Exploded illustration of the glycolysis and TCA cycles highlighting the modifications to each carbon entity during all steps in the pathway. In our system, we used all carbons C13 labeled glucose (referred to as 6C13; i.e, when a molecule had 2 carbons C13 it was referred as 2C13) to illustrate the glycolysis processes of each carbon. (b) Distribution of the number of C13-labelled carbons in glucose and glutamate in the fluidically coupled NVU system. (c) Distribution of the number of C13-labelled carbons in glucose and glutamate in the uncoupled Brain Chip as it was perfused with C13-labeled glucose, lactate and pyruvate.
Supplementary Figure 12 Theoretical metabolic flux balance analysis.
Theoretical metabolic flux balance analysis of the GABA production in synaptic cleft as the neurons can uptake exogenous pyruvate and lactate without the astrocytes contribution (while Fig. 6d display the exogenous uptake by the astrocytes and directly supplying to neurons). The GABA exchange is shown as a function of the uptake of glucose or equal amounts of lactate and pyruvate in the presence or absence of metabolites in parentheses. All fluxes are reported as μmol•g wet brain−1•min−1.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 (PDF 2998 kb)
Supplementary Information
Supplementary Tables 1–11 and Supplementary Scripts 1–5 (PDF 1699 kb)
Video 1
Hypoxia conditions in the chips (AVI 1982 kb)
Video 2
Cascade blue diffusion in the system (AVI 10443 kb)
Video 3
Proteomaps of uncoupled BBB Chip endo (WMV 2990 kb)
Video 4
Proteomaps of coupled BBB Chip endo (WMV 2855 kb)
Video 5
Proteomaps of uncoupled BBB Chipastro/peri (WMV 2964 kb)
Video 6
Proteomaps of coupled BBB Chip astro/peri (WMV 2812 kb)
Video 7
Proteomaps of uncoupled Brain Chip (WMV 2848 kb)
Video 8
Proteomaps of coupled Brain Chip (WMV 2896 kb)
Video 9
Live imaging of the Brain Chip (AVI 12823 kb)
Rights and permissions
About this article
Cite this article
Maoz, B., Herland, A., FitzGerald, E. et al. A linked organ-on-chip model of the human neurovascular unit reveals the metabolic coupling of endothelial and neuronal cells. Nat Biotechnol 36, 865–874 (2018). https://doi.org/10.1038/nbt.4226
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4226
This article is cited by
-
Endothelial EGLN3-PKM2 signaling induces the formation of acute astrocytic barrier to alleviate immune cell infiltration after subarachnoid hemorrhage
Fluids and Barriers of the CNS (2024)
-
Brain-on-a-chip: an emerging platform for studying the nanotechnology-biology interface for neurodegenerative disorders
Journal of Nanobiotechnology (2024)
-
Induced pluripotent stem cells (iPSCs): molecular mechanisms of induction and applications
Signal Transduction and Targeted Therapy (2024)
-
Mucus production, host-microbiome interactions, hormone sensitivity, and innate immune responses modeled in human cervix chips
Nature Communications (2024)
-
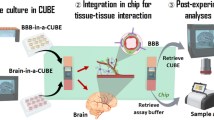
Modular tissue-in-a-CUBE platform to model blood-brain barrier (BBB) and brain interaction
Communications Biology (2024)