Abstract
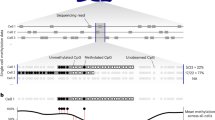
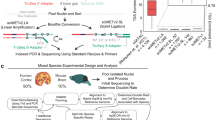
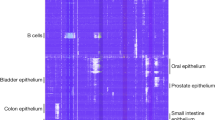
We present a highly scalable assay for whole-genome methylation profiling of single cells. We use our approach, single-cell combinatorial indexing for methylation analysis (sci-MET), to produce 3,282 single-cell bisulfite sequencing libraries and achieve read alignment rates of 68 ± 8%. We apply sci-MET to discriminate the cellular identity of a mixture of three human cell lines and to identify excitatory and inhibitory neuronal populations from mouse cortical tissue.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Varley, K.E. et al. Genome Res. 23, 555–567 (2013).
Lister, R. et al. Nature 462, 315–322 (2009).
Smallwood, S.A. et al. Nat. Methods 11, 817–820 (2014).
Farlik, M. et al. Cell Rep. 10, 1386–1397 (2015).
Farlik, M. et al. Cell Stem Cell 19, 808–822 (2016).
Angermueller, C. et al. Nat. Methods 13, 229–232 (2016).
Clark, S.J. et al. Nat. Protoc. 12, 534–547 (2017).
Luo, C. et al. Science 357, 600–604 (2017).
Amini, S. et al. Nat. Genet. 46, 1343–1349 (2014).
Adey, A. et al. Genome Res. 24, 2041–2049 (2014).
Vitak, S.A. et al. Nat. Methods 14, 302–308 (2017).
Cao, J. et al. Science 357, 661–667 (2017).
Cusanovich, D.A. et al. Science 348, 910–914 (2015).
Adey, A. et al. Genome Biol. 11, R119 (2010).
Karimzadeh, M. et al. bioRxiv (2017).
Zerbino, D.R., Wilder, S.P., Johnson, N., Juettemann, T. & Flicek, P.R. Genome Biol. 16, 56 (2015).
Roadmap Epigenomics Consortium. Nature 518, 317–330 (2015).
The ENCODE Project Consortium. Nature 489, 57–74 (2012).
Lister, R. et al. Science 341, 1237905 (2013).
Guo, J.U. et al. Nat. Neurosci. 17, 215–222 (2014).
Adey, A. & Shendure, J. Genome Res. 22, 1139–1143 (2012).
Krueger, F. & Andrews, S.R. Bioinformatics 27, 1571–1572 (2011).
Lee, D. & Seung, S. Nature 401, 778 (1999).
Ester, M. et al. Proc. 2nd Int. Conf. Knowl. Discov. Data Min. 226–231 (AAAI,1996).
Heinz, S. et al. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We would like to thank B. DeRosa for culturing the primary fibroblast cell line for this project (Department of Molecular & Medical Genetics, Oregon Health & Science University, Portland, Oregon, USA). We would like to thank other members of the Adey laboratory for helpful suggestions and dialog pertaining to this work, particularly S. Vitak. We also thank G. Mandel for providing the mice used in this study and for helpful discussion and comments on the manuscript (Vollum Institute, Oregon Health & Science University, Portland, Oregon, USA). J.R.S. is supported by the Rett Syndrome Research Trust. A.C.A. is supported by an R35 from NIGMS (1R35GM124704-01), and the Knight Cardiovascular Institute. B.J.O. is supported a fellowship from the Sloan Foundation.
Author information
Authors and Affiliations
Contributions
A.C.A. and R.M.M. conceived the sci-MET assay. R.M.M. carried out all sci-MET preparations with contributions from A.J.F. A.C.A., R.M.M., F.J.S., D.P., and S.N. designed the sci-MET adaptors and primers and reduced the assay to practice. R.M.M., F.J.S., D.P., and S.N. carried out all sequencing. R.M.M. led the data analysis. D.S. and Z.X. performed the NMF-tSNE analysis. K.A.T. provided additional analyses. J.R.S. performed mouse cortex dissection. F.J.S., J.S., C.T., and B.J.O. contributed to analysis design and edited the manuscript. A.C.A. supervised all aspects of the study. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.P., S.N., and F.J.S. are all employees of Illumina Inc. F.J.S., D.P., S.N., A.C.A., R.M.M., and J.S. all have one or more patents pertaining to one or more aspects of the technologies described here.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–25 (PDF 5847 kb)
Supplementary Tables
Supplementary tables 1–5 (PDF 158 kb)
Supplementary Notes
Supplementary Notes 1–2 (PDF 646 kb)
Supplementary Code
Supplementary Code: Custom scripts used to demultiplex fastq format files generated based on custom indexing strategy. Files included are text based barcodes, and perl scripts for demultiplexing and deduplicated reads. (ZIP 5 kb)
Supplementary Dataset 1
sciMET Transposase-loaded Oligos (5′-3′) design. (XLSX 13 kb)
Supplementary Dataset 2
sci-MET on Human Cell Line Mix metadata, summary statistics, and quality control metrics. (XLSX 183 kb)
Supplementary Dataset 3
sci-MET on Mouse Cortex metadata, summarystatistics and quality control metrics. (XLSX 83 kb)
Supplementary Dataset 4
Non-binary CG enrichment across genomic annotations and transcription factor binding sites. Pearson's paired chisquared test was performed between non-binary and binary sites per feature per collapsed cluster. (XLSX 77 kb)
Rights and permissions
About this article
Cite this article
Mulqueen, R., Pokholok, D., Norberg, S. et al. Highly scalable generation of DNA methylation profiles in single cells. Nat Biotechnol 36, 428–431 (2018). https://doi.org/10.1038/nbt.4112
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4112
This article is cited by
-
sciMET-cap: high-throughput single-cell methylation analysis with a reduced sequencing burden
Genome Biology (2024)
-
Advances in single-cell omics and multiomics for high-resolution molecular profiling
Experimental & Molecular Medicine (2024)
-
Joint single-cell profiling resolves 5mC and 5hmC and reveals their distinct gene regulatory effects
Nature Biotechnology (2024)
-
Simultaneous single-cell analysis of 5mC and 5hmC with SIMPLE-seq
Nature Biotechnology (2024)
-
Combinatorial quantification of 5mC and 5hmC at individual CpG dyads and the transcriptome in single cells reveals modulators of DNA methylation maintenance fidelity
Nature Structural & Molecular Biology (2024)