Abstract
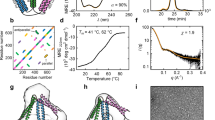
Polypeptides and polynucleotides are natural programmable biopolymers that can self-assemble into complex tertiary structures. We describe a system analogous to designed DNA nanostructures in which protein coiled-coil (CC) dimers serve as building blocks for modular de novo design of polyhedral protein cages that efficiently self-assemble in vitro and in vivo. We produced and characterized >20 single-chain protein cages in three shapes—tetrahedron, four-sided pyramid, and triangular prism—with the largest containing >700 amino-acid residues and measuring 11 nm in diameter. Their stability and folding kinetics were similar to those of natural proteins. Solution small-angle X-ray scattering (SAXS), electron microscopy (EM), and biophysical analysis confirmed agreement of the expressed structures with the designs. We also demonstrated self-assembly of a tetrahedral structure in bacteria, mammalian cells, and mice without evidence of inflammation. A semi-automated computational design platform and a toolbox of CC building modules are provided to enable the design of protein cages in any polyhedral shape.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Douglas, S.M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Seeman, N.C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
He, Y. et al. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature 452, 198–201 (2008).
Veneziano, R. et al. Designer nanoscale DNA assemblies programmed from the top down. Science 352, 1534 (2016).
Benson, E. et al. DNA rendering of polyhedral meshes at the nanoscale. Nature 523, 441–444 (2015).
Lupas, A.N. & Alva, V. Ribosomal proteins as documents of the transition from unstructured (poly)peptides to folded proteins. J. Struct. Biol. 198, 74–81 (2017).
Demain, A.L. & Vaishnav, P. Production of recombinant proteins by microbes and higher organisms. Biotechnol. Adv. 27, 297–306 (2009).
Taylor, W.R., Chelliah, V., Hollup, S.M., MacDonald, J.T. & Jonassen, I. Probing the “dark matter” of protein fold space. Structure 17, 1244–1252 (2009).
Kuhlman, B. et al. Design of a novel globular protein fold with atomic-level accuracy. Science 302, 1364–1368 (2003).
King, N.P. et al. Computational design of self-assembling protein nanomaterials with atomic level accuracy. Science 336, 1171–1174 (2012).
Doyle, L. et al. Rational design of α-helical tandem repeat proteins with closed architectures. Nature 528, 585–588 (2015).
Regan, L. & DeGrado, W.F. Characterization of a helical protein designed from first principles. Science 241, 976–978 (1988).
Woolfson, D.N. The design of coiled-coil structures and assemblies. Adv. Protein Chem. 70, 79–112 (2005).
Gradišar, H. et al. Design of a single-chain polypeptide tetrahedron assembled from coiled-coil segments. Nat. Chem. Biol. 9, 362–366 (2013).
Plückthun, A. Designed ankyrin repeat proteins (DARPins): binding proteins for research, diagnostics, and therapy. Annu. Rev. Pharmacol. Toxicol. 55, 489–511 (2015).
Grove, T.Z., Cortajarena, A.L. & Regan, L. Ligand binding by repeat proteins: natural and designed. Curr. Opin. Struct. Biol. 18, 507–515 (2008).
Bella, J., Hindle, K.L., McEwan, P.A. & Lovell, S.C. The leucine-rich repeat structure. Cell. Mol. Life Sci. 65, 2307–2333 (2008).
Pinheiro, A.V., Han, D., Shih, W.M. & Yan, H. Challenges and opportunities for structural DNA nanotechnology. Nat. Nanotechnol. 6, 763–772 (2011).
Kočar, V. et al. Design principles for rapid folding of knotted DNA nanostructures. Nat. Commun. 7, 10803 (2016).
Drobnak, I., Gradišar, H., Ljubetič, A., Merljak, E. & Jerala, R. Modulation of Coiled-Coil Dimer Stability through Surface Residues while Preserving Pairing Specificity. J. Am. Chem. Soc. 139, 8229–8236 (2017).
Götze, M. et al. StavroX--a software for analyzing cross-linked products in protein interaction studies. J. Am. Soc. Mass Spectrom. 23, 76–87 (2012).
Lawrence, M.S., Phillips, K.J. & Liu, D.R. Supercharging proteins can impart unusual resilience. J. Am. Chem. Soc. 129, 10110–10112 (2007).
Plaxco, K.W., Simons, K.T. & Baker, D. Contact order, transition state placement and the refolding rates of single domain proteins. J. Mol. Biol. 277, 985–994 (1998).
Kočar, V. et al. TOPOFOLD, the designed modular biomolecular folds: polypeptide-based molecular origami nanostructures following the footsteps of DNA. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 7, 218–237 (2015).
Negron, C. & Keating, A.E. A set of computationally designed orthogonal antiparallel homodimers that expands the synthetic coiled-coil toolkit. J. Am. Chem. Soc. 136, 16544–16556 (2014).
Doig, A.J. & Baldwin, R.L. N- and C-capping preferences for all 20 amino acids in alpha-helical peptides. Protein Sci. 4, 1325–1336 (1995).
Fijavž, G., Pisanski, T. & Rus, J. Strong traces model of self-assembly polypeptide structures. MATCH Commun. Math. Comput. Chem. 71, 199–212 (2014)<.
Noel, J.K. et al. SMOG 2: a versatile software package for generating structure-based models. PLoS Comput. Biol. 12, e1004794 (2016).
Englander, S.W. & Mayne, L. The nature of protein folding pathways. Proc. Natl. Acad. Sci. USA 111, 15873–15880 (2014).
Agyemang, A.F., Harrison, S.R., Siegel, R.M. & McDermott, M.F. Protein misfolding and dysregulated protein homeostasis in autoinflammatory diseases and beyond. Semin. Immunopathol. 37, 335–347 (2015).
Jahn, K. et al. Functional patterning of DNA origami by parallel enzymatic modification. Bioconjug. Chem. 22, 819–823 (2011).
Padilla, J.E., Colovos, C. & Yeates, T.O. Nanohedra: using symmetry to design self assembling protein cages, layers, crystals, and filaments. Proc. Natl. Acad. Sci. USA 98, 2217–2221 (2001).
Lai, Y.-T. et al. Designing and defining dynamic protein cage nanoassemblies in solution. Sci. Adv. 2, e1501855 (2016).
King, N.P. et al. Accurate design of co-assembling multi-component protein nanomaterials. Nature 510, 103–108 (2014).
Hsia, Y. et al. Design of a hyperstable 60-subunit protein icosahedron. Nature 535, 136–139 (2016).
Fletcher, J.M. et al. Self-assembling cages from coiled-coil peptide modules. Science 340, 595–599 (2013).
Wen, A.M. & Steinmetz, N.F. Design of virus-based nanomaterials for medicine, biotechnology, and energy. Chem. Soc. Rev. 45, 4074–4126 (2016).
Brunette, T.J. et al. Exploring the repeat protein universe through computational protein design. Nature 528, 580–584 (2015).
Kanekiyo, M. et al. Rational design of an Epstein-Barr virus vaccine targeting the receptor-binding site. Cell 162, 1090–1100 (2015).
López-Sagaseta, J., Malito, E., Rappuoli, R. & Bottomley, M.J. Self-assembling protein nanoparticles in the design of vaccines. Comput. Struct. Biotechnol. J. 14, 58–68 (2016).
Kushnir, N., Streatfield, S.J. & Yusibov, V. Virus-like particles as a highly efficient vaccine platform: diversity of targets and production systems and advances in clinical development. Vaccine 31, 58–83 (2012).
Correia, B.E. et al. Proof of principle for epitope-focused vaccine design. Nature 507, 201–206 (2014).
Kanekiyo, M. et al. Self-assembling influenza nanoparticle vaccines elicit broadly neutralizing H1N1 antibodies. Nature 499, 102–106 (2013).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci. 50, 2.9.1–2.9.31 (2007).
Pettersen, E.F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
McGibbon, R.T. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Perez, F. & Granger, B.E. IPython: a system for interactive scientific computing. Comput. Sci. Eng. 9, 21–29 (2007).
Ivankov, D.N. et al. Contact order revisited: influence of protein size on the folding rate. Protein Sci. 12, 2057–2062 (2003).
Testa, O.D., Moutevelis, E. & Woolfson, D.N. CC+: a relational database of coiled-coil structures. Nucleic Acids Res. 37, D315–D322 (2009).
Grigoryan, G., Reinke, A.W. & Keating, A.E. Design of protein-interaction specificity gives selective bZIP-binding peptides. Nature 458, 859–864 (2009).
Gradišar, H. & Jerala, R. De novo design of orthogonal peptide pairs forming parallel coiled-coil heterodimers. J. Pept. Sci. 17, 100–106 (2011).
Zhao, X., Ghaffari, S., Lodish, H., Malashkevich, V.N. & Kim, P.S. Structure of the Bcr-Abl oncoprotein oligomerization domain. Nat. Struct. Biol. 9, 117–120 (2002).
Oshaben, K.M., Salari, R., McCaslin, D.R., Chong, L.T. & Horne, W.S. The native GCN4 leucine-zipper domain does not uniquely specify a dimeric oligomerization state. Biochemistry 51, 9581–9591 (2012).
Wood, C.W. et al. CCBuilder: an interactive web-based tool for building, designing and assessing coiled-coil protein assemblies. Bioinformatics 30, 3029–3035 (2014).
Phillips, J.C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Abraham, M.J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2, 1–7 (2015).
Chen, Y.H., Yang, J.T. & Chau, K.H. Determination of the helix and beta form of proteins in aqueous solution by circular dichroism. Biochemistry 13, 3350–3359 (1974).
Schrödinger, LLC. The PyMOL Molecular Graphics System, Version∼1.8. (2015).
Drobnak, I., Vesnaver, G. & Lah, J. Model-based thermodynamic analysis of reversible unfolding processes. J. Phys. Chem. B 114, 8713–8722 (2010).
Press, W.H., Teukolsky, S.A., Vetterling, W.T. & Flannery, B.P. Numerical Recipes in C++: The Art of Scientific Computing (Cambridge University Press, 2002).
Gough, B., ed. GNU Scientific Library Reference Manual (Network Theory Ltd., 2009).
Konarev, P., Volkov, V., Sokolova, A., Koch, M. & Svergun, D. PRIMUS - a Windows-PC based system for small-angle scattering data analysis. J. Appl. Cryst. 36, 1277–1282 (2003).
Förster, S., Apostol, L. & Bras, W. Scatter: software for the analysis of nano-and mesoscale small-angle scattering. J. Appl. Cryst. 43, 639–646 (2010).
Franke, D. & Svergun, D.I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J. Appl. Cryst. 42, 342–346 (2009).
Hura, G.L. et al. Comprehensive macromolecular conformations mapped by quantitative SAXS analyses. Nat. Methods 10, 453–454 (2013).
Schneidman-Duhovny, D., Hammel, M. & Sali, A. FoXS: a web server for rapid computation and fitting of SAXS profiles. Nucleic Acids Res. 38, W540–W544 (2010).
Bakan, A., Meireles, L.M. & Bahar, I. ProDy: protein dynamics inferred from theory and experiments. Bioinformatics 27, 1575–1577 (2011).
Petoukhov, M.V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Shevchenko, A., Tomas, H., Havliš, J., Olsen, J.V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2007).
de la Rosa-Trevín, J.M. et al. Scipion: a software framework toward integration, reproducibility and validation in 3D electron microscopy. J. Struct. Biol. 195, 93–99 (2016).
Abrishami, V. et al. A pattern matching approach to the automatic selection of particles from low-contrast electron micrographs. Bioinformatics 29, 2460–2468 (2013).
Sorzano, C.O.S. et al. A clustering approach to multireference alignment of single-particle projections in electron microscopy. J. Struct. Biol. 171, 197–206 (2010).
Goddard, T.D., Huang, C.C. & Ferrin, T.E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Wang, Y. et al. Activation of ATF6 and an ATF6 DNA binding site by the endoplasmic reticulum stress response. J. Biol. Chem. 275, 27013–27020 (2000).
Hornung, V. et al. Silica crystals and aluminum salts activate the NALP3 inflammasome through phagosomal destabilization. Nat. Immunol. 9, 847–856 (2008).
Hafner-Bratkovič, I., Benčina, M., Fitzgerald, K.A., Golenbock, D. & Jerala, R. NLRP3 inflammasome activation in macrophage cell lines by prion protein fibrils as the source of IL-1β and neuronal toxicity. Cell. Mol. Life Sci. 69, 4215–4228 (2012).
Acknowledgements
This research was supported by the Slovenian Research Agency, program P4-0176, projects N4-0037 and J4-5528 (R.J.), L4-6812 (H.G.), and J3-7034 and BI-US/17-18-051 (M.B.); the ERANET SynBio project Bioorigami (ERASYNBIO1-006 to R.J.); COST actions CM1304 (R.J. and A.L.) and CM1306 (R.J. and J.A.); a grant from ICGEB (CRP/SLO14-03) to H.G.; ESRF, for making available its facility for performing SAXS measurements; MSC-ETN 642157 Tollerant H2020 (R.J. and F.L.). This work has been supported by iNEXT, PID1771 (R.J.), PID2706 (R.J.), PID1824 (H.G.), VID3987 (H.G.), funded by the Horizon 2020 Programme of the EU; NVIDIA Corporation for the donation of the Quadro GP100 GPU (J.M.C.); and FP7 project FCUB ERA (GA No. 256716 to T.Ć.V.) for the use of the proteomics facility. We thank K. Djinović Carugo for useful suggestions and for performing and analyzing the initial SAXS experiments. We thank the staff of the Centre for Laboratory Animals at Biotechnical faculty of the University of Ljubljana, where animal experiments were performed. We would like to thank K. Butina, R. Bremšak, I. Škraba, D. Oven, T. Lončar, S. Božič Abram, T. Doles, S. Grudinin, J. Mihailović, and E. Žagar for their technical support. We also thank E. Žerovnik for granting access to the stopped-flow circular dichroism instrument and C. Wood for building preliminary models of the CC pairs. Plasmid encoding firefly luciferase under the ATF6 control (p5XATF6-GL3) was a gift from R. Prywes (Columbia University, New York, NY, USA). Immortalized mouse bone-marrow-derived macrophages were a gift from K. Fitzgerald (University of Massachusetts Medical School, Worcester, MA, USA).
Author information
Authors and Affiliations
Contributions
A.L., F.L., H.G., I.D., J.A., Ž.S., and R.J. designed the CCPO variants. F.L., H.G., Ž.S. and N.K. cloned, purified, and experimentally characterized the proteins. A.L., I.D., J.A., and T.P. wrote the CoCoPOD platform. A.M. and T.Ć.V. performed the cross-linking experiments. J.A. and A.R. performed the SAXS experiments and SAXS data analysis. I.H.-B. and M.B. performed the experiments on the cells. M.B. and D.L. performed confocal microcopy imaging. D.L. performed the animal experiments. R.M. and J.M.C. performed the EM experiments and data processing. R.J. conceived the study, led the research, and wrote the initial manuscript. All authors discussed the results and reviewed and contributed to the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Discussion, Supplementary Figures 1–22, Supplementary Tables 1–7, and Supplementary Note (PDF 18500 kb)
Supplementary Data
Topologies circular permutations TCO (XLSX 86 kb)
Supplementary Code
Supplementary Source Code (ZIP 1243 kb)
Rights and permissions
About this article
Cite this article
Ljubetič, A., Lapenta, F., Gradišar, H. et al. Design of coiled-coil protein-origami cages that self-assemble in vitro and in vivo. Nat Biotechnol 35, 1094–1101 (2017). https://doi.org/10.1038/nbt.3994
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3994
This article is cited by
-
Enhancing prime editor flexibility with coiled-coil heterodimers
Genome Biology (2024)
-
Blueprinting extendable nanomaterials with standardized protein blocks
Nature (2024)
-
A multiplexed bacterial two-hybrid for rapid characterization of protein–protein interactions and iterative protein design
Nature Communications (2023)
-
Precisely patterned nanofibres made from extendable protein multiplexes
Nature Chemistry (2023)
-
Engineering a scalable and orthogonal platform for synthetic communication in mammalian cells
Nature Communications (2023)