Abstract
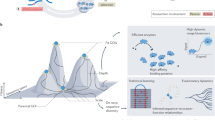
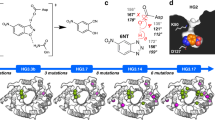
Optimization of a protein's pharmaceutical properties is usually carried out by rational design and/or directed evolution. Here we test an alternative approach based on ancestral sequence reconstruction. Using available genomic sequence data on coagulation factor VIII and predictive models of molecular evolution, we engineer protein variants with improved activity, stability, and biosynthesis potential and reduced inhibition by anti-drug antibodies. In principle, this approach can be applied to any protein drug based on a conserved gene sequence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Doering, C.B. et al. Thromb. Haemost. 88, 450–458 (2002).
Doering, C.B., Healey, J.F., Parker, E.T., Barrow, R.T. & Lollar, P. J. Biol. Chem. 277, 38345–38349 (2002).
Zakas, P.M. et al. PLoS One 7, e49481 (2012).
Zakas, P.M., Vanijcharoenkarn, K., Markovitz, R.C., Meeks, S.L. & Doering, C.B. J. Thromb. Haemost. 13, 72–81 (2015).
Sabatino, D.E. et al. Blood 114, 4562–4565 (2009).
Doering, C.B., Healey, J.F., Parker, E.T., Barrow, R.T. & Lollar, P. J. Biol. Chem. 279, 6546–6552 (2004).
Parker, E.T., Doering, C.B. & Lollar, P. J. Biol. Chem. 281, 13922–13930 (2006).
Zuckerkandl, E. & Pauling, L. J. Theor. Biol. 8, 357–366 (1965).
Merkl, R. & Sterner, R. Biol. Chem. 397, 1–21 (2016).
Risso, V.A., Gavira, J.A., Mejia-Carmona, D.F., Gaucher, E.A. & Sanchez-Ruiz, J.M. J. Am. Chem. Soc. 135, 2899–2902 (2013).
Ivarsson, Y., Mackey, A.J., Edalat, M., Pearson, W.R. & Mannervik, B. J. Biol. Chem. 278, 8733–8738 (2003).
Harms, M.J. et al. Proc. Natl. Acad. Sci. USA 110, 11475–11480 (2013).
Kratzer, J.T. et al. Proc. Natl. Acad. Sci. USA 111, 3763–3768 (2014).
Wilson, C. et al. Science 347, 882–886 (2015).
Gaucher, E.A., Govindarajan, S. & Ganesh, O.K. Nature 451, 704–707 (2008).
Brown, H.C., Gangadharan, B. & Doering, C.B. J. Biol. Chem. 286, 24451–24457 (2011).
Pipe, S.W., Eickhorst, A.N., McKinley, S.H., Saenko, E.L. & Kaufman, R.J. Blood 93, 176–183 (1999).
Leong, L. et al. Blood 125, 392–398 (2015).
Markovitz, R.C., Healey, J.F., Parker, E.T., Meeks, S.L. & Lollar, P. Blood 121, 2785–2795 (2013).
Esmon, C.T. & Lollar, P. J. Biol. Chem. 271, 13882–13887 (1996).
Barrow, R.T., Parker, E.T., Krishnaswamy, S. & Lollar, P. J. Biol. Chem. 269, 26796–26800 (1994).
Healey, J.F. et al. J. Thromb. Haemost. 5, 512–519 (2007).
Meeks, S.L., Healey, J.F., Parker, E.T., Barrow, R.T. & Lollar, P. Blood 110, 4234–4242 (2007).
Bi, L. et al. Nat. Genet. 10, 119–121 (1995).
Brown, H.C. et al. Mol. Ther. Methods Clin. Dev. 1, 14036 (2014).
Gaucher, E.A., Thomson, J.M., Burgan, M.F. & Benner, S.A. Nature 425, 285–288 (2003).
Huelsenbeck, J.P., Ronquist, F., Nielsen, R. & Bollback, J.P. Science 294, 2310–2314 (2001).
Yang, Z. Mol. Biol. Evol. 24, 1586–1591 (2007).
Lollar, P., Parker, E.T. & Fay, P.J. J. Biol. Chem. 267, 23652–23657 (1992).
Healey, J.F. et al. Thromb. Haemost. 102, 35–41 (2009).
Barrow, R.T. & Lollar, P. J. Thromb. Haemost. 4, 2223–2229 (2006).
Larsen, J.E., Lund, O. & Nielsen, M. Immunome Res. 2, 2 (2006).
Spencer, H.T. et al. Mol. Ther. 19, 302–309 (2011).
Parker, E.T. & Lollar, P. Thromb. Haemost. 89, 480–485 (2003).
Dixon, W.J. Neurosci. Biobehav. Rev. 15, 47–50 (1991).
McIntosh, J. et al. Blood 121, 3335–3344 (2013).
Acknowledgements
This work was supported by funding from the National Institutes of Health/National Heart, Lung, and Blood Institute for the Translational Research Centers in Thrombotic and Hemostatic Disorders (U54 HL112309 to H.T.S., S.L.M., and C.B.D.), the Bayer Hemophilia Awards Program, Bayer HealthCare (C.B.D.) as well as a research partnership between Children's Healthcare of Atlanta and the Georgia Institute of Technology (C.B.D. and E.A.G.). We also thank E.T. Parker (Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Emory University, Atlanta, Georgia, USA) for technical assistance with the in vivo mouse studies.
Author information
Authors and Affiliations
Contributions
P.M.Z. designed and performed experiments, analyzed the data, and drafted the manuscript. H.C.B. performed gene transfer experiments and edited the manuscript. K.K. performed experiments. S.L.M. contributed reagents, designed experiments, analyzed data and edited the manuscript. H.T.S. conceived the project, designed experiments, analyzed data and edited the manuscript. E.A.G. performed ASR and edited the manuscript. C.B.D. conceived the project, designed experiments, analyzed data and drafted and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
C.B.D., E.A.G., H.T.S. and P.M.Z. are inventors on a patent application describing ancestral FVIII technology filed by Emory University/Children's Healthcare of Atlanta and the Georgia Institute of Technology. C.B.D. and H.T.S. are co-founders of Expression Therapeutics, LLC, and own equity in the company. Expression Therapeutics owns the intellectual property associated with ET3 and has plans to commercially develop technology used in the research described in this paper. The terms of this arrangement have been reviewed and approved by Emory University in accordance with its conflict of interest policies.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1–3 and Supplementary Note (PDF 2791 kb)
Rights and permissions
About this article
Cite this article
Zakas, P., Brown, H., Knight, K. et al. Enhancing the pharmaceutical properties of protein drugs by ancestral sequence reconstruction. Nat Biotechnol 35, 35–37 (2017). https://doi.org/10.1038/nbt.3677
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3677
This article is cited by
-
Extant Sequence Reconstruction: The Accuracy of Ancestral Sequence Reconstructions Evaluated by Extant Sequence Cross-Validation
Journal of Molecular Evolution (2024)
-
Design, structure and plasma binding of ancestral β-CoV scaffold antigens
Nature Communications (2023)
-
Evolution of CRISPR-associated endonucleases as inferred from resurrected proteins
Nature Microbiology (2023)
-
Enzymatic upgrading of nanochitin using an ancient lytic polysaccharide monooxygenase
Communications Materials (2022)
-
Bypassing evolutionary dead ends and switching the rate-limiting step of a human immunotherapeutic enzyme
Nature Catalysis (2022)