Abstract
The combination of Cas9, guide RNA and repair template DNA can induce precise gene editing and the correction of genetic diseases in adult mammals. However, clinical implementation of this technology requires safe and effective delivery of all of these components into the nuclei of the target tissue. Here, we combine lipid nanoparticle–mediated delivery of Cas9 mRNA with adeno-associated viruses encoding a sgRNA and a repair template to induce repair of a disease gene in adult animals. We applied our delivery strategy to a mouse model of human hereditary tyrosinemia and show that the treatment generated fumarylacetoacetate hydrolase (Fah)-positive hepatocytes by correcting the causative Fah-splicing mutation. Treatment rescued disease symptoms such as weight loss and liver damage. The efficiency of correction was >6% of hepatocytes after a single application, suggesting potential utility of Cas9-based therapeutic genome editing for a range of diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Doudna, J.A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Mali, P., Esvelt, K.M. & Church, G.M. Cas9 as a versatile tool for engineering biology. Nat. Methods 10, 957–963 (2013).
Sander, J.D. & Joung, J.K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Aponte, J.L. et al. Point mutations in the murine fumarylacetoacetate hydrolase gene: animal models for the human genetic disorder hereditary tyrosinemia type 1. Proc. Natl. Acad. Sci. USA 98, 641–645 (2001).
Azuma, H. et al. Robust expansion of human hepatocytes in Fah-/-/Rag2-/-/Il2rg-/- mice. Nat. Biotechnol. 25, 903–910 (2007).
Paulk, N.K. et al. Adeno-associated virus gene repair corrects a mouse model of hereditary tyrosinemia in vivo. Hepatology 51, 1200–1208 (2010).
Schwank, G. et al. Functional repair of CFTR by CRISPR-Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell 13, 653–658 (2013).
Wu, Y. et al. Correction of a genetic disease in mouse via use of CRISPR-Cas9. Cell Stem Cell 13, 659–662 (2013).
Long, C. et al. Prevention of muscular dystrophy in mice by CRISPR-Cas9-mediated editing of germline DNA. Science 345, 1184–1188 (2014).
Ran, F.A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Swiech, L. et al. In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9. Nat. Biotechnol. 33, 102–106 (2015).
Zuris, J.A. et al. Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat. Biotechnol. 33, 73–80 (2015).
Chu, V.T., Weber, T. & Wefers, B. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat. Biotechnol. 33, 543–548 (2015).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Khorsandi, S.E. et al. Minimally invasive and selective hydrodynamic gene therapy of liver segments in the pig and human. Cancer Gene Ther. 15, 225–230 (2008).
Kay, M.A. State-of-the-art gene-based therapies: the road ahead. Nat. Rev. Genet. 12, 316–328 (2011).
Yin, H. et al. Non-viral vectors for gene-based therapy. Nat. Rev. Genet. 15, 541–555 (2014).
Love, K.T. et al. Lipid-like materials for low-dose, in vivo gene silencing. Proc. Natl. Acad. Sci. USA 107, 1864–1869 (2010).
Semple, S.C. et al. Rational design of cationic lipids for siRNA delivery. Nat. Biotechnol. 28, 172–176 (2010).
Chen, D. et al. Rapid discovery of potent siRNA-containing lipid nanoparticles enabled by controlled microfluidic formulation. J. Am. Chem. Soc. 134, 6948–6951 (2012).
Kormann, M.S. et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nat. Biotechnol. 29, 154–157 (2011).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Kauffman, K.J. et al. Optimization of lipid nanoparticle formulations for mRNA delivery in vivo with fractional factorial and definitive screening designs. Nano Lett. 15, 7300–7306 (2015).
Sundararajan, S., Wakamiya, M., Behringer, R.R. & Rivera-Pérez, J.A. A fast and sensitive alternative for β-galactosidase detection in mouse embryos. Development 139, 4484–4490 (2012).
Kim, H. et al. A co-CRISPR strategy for efficient genome editing in Caenorhabditis elegans. Genetics 197, 1069–1080 (2014).
Barzel, A. et al. Promoterless gene targeting without nucleases ameliorates haemophilia B in mice. Nature 517, 360–364 (2015).
Xue, W. et al. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature 514, 380–384 (2014).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2014).
Mahiny, A.J. et al. In vivo genome editing using nuclease-encoding mRNA corrects SP-B deficiency. Nat. Biotechnol. 33, 584–586 (2015).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Li, H. et al. In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature 475, 217–221 (2011).
Wu, X. et al. Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat. Biotechnol. 32, 670–676 (2014).
Chen, H.Z. et al. Canonical and atypical E2Fs regulate the mammalian endocycle. Nat. Cell Biol. 14, 1192–1202 (2012).
Elliott, B., Richardson, C., Winderbaum, J., Nickoloff, J.A. & Jasin, M. Gene conversion tracts from double-strand break repair in mammalian cells. Mol. Cell. Biol. 18, 93–101 (1998).
Goldberg, A.D. et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140, 678–691 (2010).
Findlay, G.M., Boyle, E.A., Hause, R.J., Klein, J.C. & Shendure, J. Saturation editing of genomic regions by multiplex homology-directed repair. Nature 513, 120–123 (2014).
Cox, D.B., Platt, R.J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Atabai, K. et al. Mfge8 is critical for mammary gland remodeling during involution. Mol. Biol. Cell 16, 5528–5537 (2005).
Gilbert, L.A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Bolukbasi, M.F. et al. DNA-binding-domain fusions enhance the targeting range and precision of Cas9. Nat. Methods 12, 1150–1156 (2015).
Zhu, L.J. et al. ChIPpeakAnno: a Bioconductor package to annotate ChIP-seq and ChIP-chip data. BMC Bioinformatics 11, 237 (2010).
Zhu, L. Overview of guide RNA design tools for CRISPR-Cas9 genome editing technology. Front. Biol. 10, 289–296 (2015).
Zhu, L.J., Holmes, B.R., Aronin, N. & Brodsky, M.H. CRISPRseek: a bioconductor package to identify target-specific guide RNAs for CRISPR-Cas9 genome-editing systems. PLoS One 9, e108424 (2014).
Acknowledgements
We thank M. Grompe, S. Levine, T. Jacks, P. Sharp, E. Sontheimer, C. Mello, P. Zamore, M. Moore, T. Flotte, T. Tammela, F. Sanchez-Rivera, T. Papagiannakopoulos, D. Wang, J. Moore and A. Vegas for discussions and for sharing reagents, S. Hough for technical assistance and K. Cormier for histology. This work is supported by grants from the National Institutes of Health (NIH), 5R00CA169512 and Worcester Foundation (to W.X.). H.Y. is supported by Skoltech Center and 5-U54-CA151884-04 (NIH Centers for Cancer Nanotechnology Excellence and the Harvard-MIT Center of Cancer Nanotechnology Excellence). Y.D. acknowledges support from the National Institute of Biomedical Imaging and Bioengineering for his postdoctoral fellowship 1F32EB017625. V.K. acknowledges support from the Russian scientific fund, grant number 14-34–00017. This work is supported in part by Cancer Center Support (core) grant P30-CA14051 from the NIH. We thank the Swanson Biotechnology Center for technical support. We thank C. Wang at Boston Children's Hospital Viral Core for AAV prep (supported by core grant 5P30EY012196-17). The authors acknowledge the service to the MIT community of the late S. Collier.
Author information
Authors and Affiliations
Contributions
H.Y., W.X. and D.G.A. designed the study. H.Y. and W.X. directed the project. H.Y., C.-Q.S., J.R.D., L.J.Z., Y.L., Q.W., J.Y., S.S., A.B., A.G., M.F.B., A.P., S.W. and R.L.B. performed experiments and analyzed data. G.G., Z.W., Y.D., V.K., S.A.W. and R.L. provided reagents and conceptual advice. H.Y., W.X. and D.G.A. wrote the manuscript with comments from all authors.
Corresponding authors
Ethics declarations
Competing interests
D.G.A., H.Y., J.R.K. and W.X. have applied for patents on the subject matter of this paper. D.G.A. is a scientific co-founder of CRISPR Therapeutics.
Integrated supplementary information
Supplementary Figure 1 Cas9 mRNA nanoparticles characterization.
(a) nano.Cas9 formulation scheme. Cas9 mRNA was mixed with C12-200, DOPE, Cholesterol, C14PEG2000 and arachidonic acid in a microfluidic chamber. (b) nano.Cas9 structure is characterized by cryo-TEM. Scale bar indicates 100nm. (c) Average diameter of nano.Cas9 was measured by dynamic light scattering. The size of nano.Cas9 (d) and the polydispersity index (PDI) (e) were measured 0, 7, 11 or 18 days after formulation and storage at 4˚C.
Supplementary Figure 2 The expression of proteins in mouse liver after mRNA nanoparticles treatment.
(a) C57bl/6 mice were i.v. injected with nanoparticles encapsulated with β-gal (b and c) or Cas9 mRNA (nano.Cas9, d and e), and livers taken. (b) The expression of β-gal protein was measured in liver lysate at 14 hours after injection. (c) The activity of β-gal in liver sections was determined by salmon-gal assay. Scale bar indicates 200 µm. (d) The expression of Cas9 protein was measured in liver lysate 14 hours after injection. 50μg negative control samples mixed with 10, 1 or 0.1ng Cas9 protein served as positive controls. β-actin served as a loading control in (b) and (d). (e) The Cas9 mRNA level in liver was determined by qRT-PCR at 4, 14, and 24 hours after injection (n=3 mice).
Supplementary Figure 3 Cas9 mRNA nanoparticles are well tolerated.
C57/Bl6 mice were treated with 2mg/kg nano.Cas9, and histology (a), the levels of liver damage markers (b) and plasma cytokines (c) were determined after 24 hours. Scale bar indicates 50μM. (n=4 mice).
Supplementary Figure 4 The time course of sgRNA expression in mouse liver.
Mice were injected with AAV-HDR and livers taken at 0, 3, 7 and 14 days after injection. qRT-PCR was performed to determine sgRNA expression in liver. The expression levels were normalized to Day 3 (n = 4 mice).
Supplementary Figure 5 A PCR approach proves substitution of the correct sequence.
(a) Design of the PCR primers. Blue arrow indicates the reverse PCR primer, which is outside the repair template. The sequence of the forward primer is presented, and “G” and “CC” in the corrected sequence are highlighted. (b) Genomic DNA of the liver tissue was extracted, and PCR was performed using the primers in (a). The predicted size of PCR product is 1.02kb. A representative sample from each group is shown (n = 3 mice). (c) The PCR product from (b) was cloned to a TA cloning vector and Sanger sequenced. The corrected “G” and “CC” are highlighted.
Supplementary Figure 6 Viral delivery of Cas9 does not increase HDR rate compared to mRNA delivery.
(a) Design of AAV-HDR template. Four “G” point mutations resulting in stabilization of β-Catenin are highlighted. Ad.Cas9 is an adenovirus expressing Cas9. (b) β-Catenin IHC. AAV-HDR-Ctnnb1 alone serves as a control. Arrows denote β-Catenin positive hepatocytes. (c-d) The Ctnnb1 locus in the liver total DNA of Ad.Cas9+ AAV-HDR-Ctnnb1 treated mice (n=2) were deep sequenced to measure indels. (e) β-Catenin positive hepatocytes were counted to determine the percentage of HDR. P < 0.01 (n = 3 mice). mRNA delivery of Cas9 yields higher rate of HDR for Fah (>6%).
Supplementary Figure 7 Cas9 mRNA delivery has minimal off-target effects at assayed sites in vivo.
(a) Top ranking off-target sites (OT1, OT3 and OT4) for sgFah and the predicted score (Hsu et al, 2013). Mismatch bases are in red. Score for the wildtype sgFah.2 targets the mutant Fah which has one mismatch with wildtype Fah (wt Fah). (b) Indel frequency is low and is comparable between control mouse and nano-Cas9+ AAV-HDR mouse. OT1, OT3 and OT4 regions were PCR amplified from mouse liver genomic DNA and analyzed by deep sequencing. (c) Surveyor assay did not detect indels at OT1, OT3 and OT4. Predicted size of uncut and cut bands are indicated.
Supplementary Figure 8 Indel rate measured by deep sequencing for GUIDE-Seq off-target sites.
OT1 is the strongest off-target sites identified by GUIDE-Seq. GOT1-11 are additional genomic sites that displayed GUIDE-Seq oligonucleotide insertions. (a) Mouse Hepa1-6 liver cells transfected with pX330.sgFah.2. #1 and #2 are replicates. (b) Mouse livers treated with nano.Cas9 and AAV-HDR (treated) or control treated (control). Fah is the on-target site. See Table S9 for details.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–6 and Supplementary Sequences (PDF 3895 kb)
Supplementary Table 7
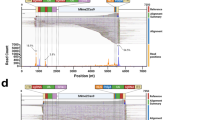
sgRNA2 GUIDE-seq +&- strand peaks (PDF 40329 kb)
Supplementary Table 8
sgRNA2 GUIDE-seq merged peaks (PDF 75 kb)
Supplementary Table 9
Deep sequencing of off-target sites (PDF 192 kb)
Rights and permissions
About this article
Cite this article
Yin, H., Song, CQ., Dorkin, J. et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat Biotechnol 34, 328–333 (2016). https://doi.org/10.1038/nbt.3471
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3471
This article is cited by
-
Targeted Gene Insertion: The Cutting Edge of CRISPR Drug Development with Hemophilia as a Highlight
BioDrugs (2024)
-
In vivo LNP-CRISPR Approaches for the Treatment of Hemophilia
Molecular Diagnosis & Therapy (2024)
-
A potential paradigm in CRISPR/Cas systems delivery: at the crossroad of microalgal gene editing and algal-mediated nanoparticles
Journal of Nanobiotechnology (2023)
-
Peptide-encoding mRNA barcodes for the high-throughput in vivo screening of libraries of lipid nanoparticles for mRNA delivery
Nature Biomedical Engineering (2023)
-
Combinatorial design of nanoparticles for pulmonary mRNA delivery and genome editing
Nature Biotechnology (2023)