Abstract
We describe an engineered photoactivatable Cas9 (paCas9) that enables optogenetic control of CRISPR-Cas9 genome editing in human cells. paCas9 consists of split Cas9 fragments and photoinducible dimerization domains named Magnets. In response to blue light irradiation, paCas9 expressed in human embryonic kidney 293T cells induces targeted genome sequence modifications through both nonhomologous end joining and homology-directed repair pathways. Genome editing activity can be switched off simply by extinguishing the light. We also demonstrate activation of paCas9 in spatial patterns determined by the sites of irradiation. Optogenetic control of targeted genome editing should facilitate improved understanding of complex gene networks and could prove useful in biomedical applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Jinek, M. et al. RNA-programmed genome editing in human cells. Elife 2, e00471 (2013).
Dow, L.E. et al. Inducible in vivo genome editing with CRISPR-Cas9. Nat. Biotechnol. 33, 390–394 (2015).
González, F. et al. An iCRISPR platform for rapid, multiplexable, and inducible genome editing in human pluripotent stem cells. Cell Stem Cell 15, 215–226 (2014).
Zetsche, B., Volz, S.E. & Zhang, F. A split-Cas9 architecture for inducible genome editing and transcription modulation. Nat. Biotechnol. 33, 139–142 (2015).
Kim, S., Kim, D., Cho, S.W., Kim, J. & Kim, J.-S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Zuris, J.A. et al. Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat. Biotechnol. 33, 73–80 (2015).
Ramakrishna, S. et al. Gene disruption by cell-penetrating peptide-mediated delivery of Cas9 protein and guide RNA. Genome Res. 24, 1020–1027 (2014).
Laplante, M. & Sabatini, D.M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Brieke, C., Rohrbach, F., Gottschalk, A., Mayer, G. & Heckel, A. Light-controlled tools. Angew. Chem. Int. Edn Engl. 51, 8446–8476 (2012).
Lee, H., Larson, D. & Lawrence, D. Illuminating the chemistry of life: design, synthesis, and applications of “caged” and related photoresponsive compounds. ACS Chem. Biol. 4, 409–427 (2009).
DeRose, R., Miyamoto, T. & Inoue, T. Manipulating signaling at will: chemically-inducible dimerization (CID) techniques resolve problems in cell biology. Pflugers Arch. 465, 409–417 (2013).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014).
Kennedy, M.J. et al. Rapid blue-light–mediated induction of protein interactions in living cells. Nat. Methods 7, 973–975 (2010).
Bugaj, L.J., Choksi, A.T., Mesuda, C.K., Kane, R.S. & Schaffer, D.V. Optogenetic protein clustering and signaling activation in mammalian cells. Nat. Methods 10, 249–252 (2013).
Kawano, F., Suzuki, H., Furuya, A. & Sato, M. Engineered pairs of distinct photoswitches for optogenetic control of cellular proteins. Nat. Commun. 6, 6256 (2015).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Kim, H. et al. Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nat. Methods 8, 941–943 (2011).
Ramakrishna, S. et al. Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nuclease-induced mutations. Nat. Commun. 5, 3378 (2014).
Qi, L.S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Voon, D.C. et al. Use of mRNA- and protein-destabilizing elements to develop a highly responsive reporter system. Nucleic Acids Res. 33, e27 (2005).
Lin, Y. et al. Rapidly reversible manipulation of molecular activity with dual chemical dimerizers. Angew. Chem. Int. Edn Engl. 52, 6450–6454 (2013).
Shen, Z. et al. Conditional knockouts generated by engineered CRISPR-Cas9 endonuclease reveal the roles of coronin in C. elegans neural development. Dev. Cell 30, 625–636 (2014).
Nihongaki, Y., Yamamoto, S., Kawano, F., Suzuki, H. & Sato, M. CRISPR-Cas9-based photoactivatable transcription system. Chem. Biol. 22, 169–174 (2015).
Polstein, L.R. & Gersbach, C.A. A light-inducible CRISPR-Cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200 (2015).
Acknowledgements
We would like to thank J. Galipon for technical assistance. This work was supported by Platform for Dynamic Approaches to Living System from the Ministry of Education, Culture, Sports, Science and Technology, Japan (MEXT) and by grants from Japan Society for the Promotion of Science (JSPS).
Author information
Authors and Affiliations
Contributions
Y.N. and M.S. conceived the project. Y.N., F.K., T.N. and M.S. designed the experiments. Y.N. performed experiments and analyzed data. Y.N. and M.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Construction of rapamycin-inducible split-Cas9.
(a) Candidate 18 split sites of Streptococcus pyogenes Cas9 protein. (b) Constructs of rapamycin-inducible Cas9. N- and C- fragments of Cas9 were fused with FRB and FKBP, respectively.
Supplementary Figure 2 Screening of split-Cas9 fragments.
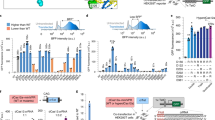
(a) Ligand-induced Cas9 activity measured by luciferase reporter plasmid HDR assay. HEK293T cells were transfected with N-fragment of Cas9 fused with FRB, C-fragment of Cas9 fused with FKBP, stop codon-inserted luciferase reporter and promoter-less luciferase donor vector. (b) Second screening of Cas9 split sites in the vicinity of residues 713 and 714.
Supplementary Figure 3 Schematic of paCas9.
The schematic is based on the crystal structures of Cas9 (PDB ID: 4UN3) (ref. 17) and the light-state dimer of Vivid (PDB ID: 3RH8) (supplementary ref. 1), a precursor protein of Magnets. N713 and C714 fragments are shown in red and blue, respectively. Positive Magnet and negative Magnet are shown in gold and cyan, respectively. sgRNA and targeted DNA are shown in green and yellow, respectively. The positions connected between Cas9 fragments and Magnets are circled in black.
Supplementary Figure 4 Time-course analysis of paCas9-mediated indel mutation in EMX1 locus.
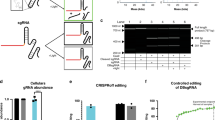
HEK293T cells were transfected with paCas9-1 targeting EMX1. After 20 h incubation, the cells were illuminated by 1.2 W/m2 blue light and genomic DNA was extracted at indicated time points. Indel mutation frequencies were evaluated by mismatch-sensitive T7E1 assay. Data are shown as the mean ± s.e.m. (n=4 from two individual experiments with biological duplicates). Indel frequencies at 0, 1, 3 and 6 h were below the detection limit (1%).
Supplementary Figure 5 paCas9-mediated indel mutation of human EMX1 locus in HeLa cells.
Indel mutation frequencies were evaluated by mismatch-sensitive T7E1 assay.
Supplementary Figure 6 paCas9 nickase can facilitate efficient genome editing using a pair of sgRNAs.
(a) Schematic of light-induced DNA double-strand breaks using a pair of sgRNAs with paCas9 D10A nickase. By introducing D10A mutation in N-terminal fragment of Cas9, paCas9 nuclease can be converted into paCas9 nickase. Using a pair of sgRNAs targeting opposite strands of targeted gene, paCas9 nickase can offer optical control of DNA double-stranded break in targeted locus. (b) Schematic of the human EMX1 locus presenting the target region of a paired sgRNAs indicated by blue lines. PAMs are also shown in red. Red allows indicate putative cleavage site. (c) Representative T7E1 assay result for calculating indel mutation rates induced by paCas9 nickase (mean, n=2 from independent experiments).
Supplementary Figure 7 Spatial surrogate reporter activation by paCas9.
(a) Schematic of surrogate EGFP reporter system (ref. 24, 25). The surrogate reporter consists of mCherry, target sequence of paCas9 (here EMX1 target site) and EGFP. In the dark, while mCherry is constitutively expressed from CMV promoter, EGFP fluorescence is not observed because EGFP gene is out of frame in the absence of Cas9 activity. Upon blue light irradiation, paCas9 is activated and a double-strand break is introduced into the EMX1 target site of the reporter. By NHEJ pathway, this break is repaired with frameshift mutations. This frameshift mutation makes EGFP gene in frame, causing the expression of mCherry-EGFP fusion protein. (b) Slit-patterned activation of paCas9. HEK293T cells were transfected with paCas9-2, surrogate EGFP reporter and sgRNA targeting the surrogate reporter. Twenty hours post transfection, samples were incubated under dark state, global blue light irradiation, and slit-patterned blue light using a photomask for 24 h.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 2 and 3, and Supplementary Notes 1 and 2 (PDF 1694 kb)
Supplementary Table 1
Target sequences of sgRNAs and oligonucleotide sequences for constructing sgRNAs. (XLSX 24 kb)
Rights and permissions
About this article
Cite this article
Nihongaki, Y., Kawano, F., Nakajima, T. et al. Photoactivatable CRISPR-Cas9 for optogenetic genome editing. Nat Biotechnol 33, 755–760 (2015). https://doi.org/10.1038/nbt.3245
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3245
This article is cited by
-
Spatiotemporal control of RNA metabolism and CRISPR–Cas functions using engineered photoswitchable RNA-binding proteins
Nature Protocols (2024)
-
Traceless photodegradable polymer cocoons for universal protein delivery and light-controlled gene editing
Science China Chemistry (2024)
-
A potential paradigm in CRISPR/Cas systems delivery: at the crossroad of microalgal gene editing and algal-mediated nanoparticles
Journal of Nanobiotechnology (2023)
-
Synthetic circuits based on split Cas9 to detect cellular events
Scientific Reports (2023)
-
Design of hypoxia responsive CRISPR-Cas9 for target gene regulation
Scientific Reports (2023)