Abstract
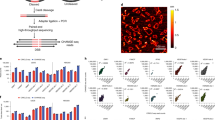
CRISPR-Cas9 genome editing technology holds great promise for discovering therapeutic targets in cancer and other diseases. Current screening strategies target CRISPR-Cas9–induced mutations to the 5′ exons of candidate genes1,2,3,4,5, but this approach often produces in-frame variants that retain functionality, which can obscure even strong genetic dependencies. Here we overcome this limitation by targeting CRISPR-Cas9 mutagenesis to exons encoding functional protein domains. This generates a higher proportion of null mutations and substantially increases the potency of negative selection. We also show that the magnitude of negative selection can be used to infer the functional importance of individual protein domains of interest. A screen of 192 chromatin regulatory domains in murine acute myeloid leukemia cells identifies six known drug targets and 19 additional dependencies. A broader application of this approach may allow comprehensive identification of protein domains that sustain cancer cells and are suitable for drug targeting.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
18 May 2015
In the version of this article initially published online, in the legend of Figure 1e, the time span of the experiment has been stated incorrectly. It should be “GFP+ cells (d2 GFP% divided by d10 GFP%)” instead of “GFP+ cells (d2 GFP% divided by d8 GFP%).” Also in the abstract Cas9 was misspelled as CaS9 in two instances. The errors have been corrected for the print, PDF and HTML versions of the article.
References
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Koike-Yusa, H., Li, Y., Tan, E.P., Velasco-Herrera Mdel, C. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014).
Zhou, Y. et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014).
Doench, J.G. et al. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014).
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Zuber, J. et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478, 524–528 (2011).
Zuber, J. et al. Toolkit for evaluating genes required for proliferation and survival using tetracycline-regulated RNAi. Nat. Biotechnol. 29, 79–83 (2011).
McJunkin, K. et al. Reversible suppression of an essential gene in adult mice using transgenic RNA interference. Proc. Natl. Acad. Sci. USA 108, 7113–7118 (2011).
Shi, J. et al. Role of SWI/SNF in acute leukemia maintenance and enhancer-mediated Myc regulation. Genes Dev. 27, 2648–2662 (2013).
Wang, E. et al. Histone H2B ubiquitin ligase RNF20 is required for MLL-rearranged leukemia. Proc. Natl. Acad. Sci. USA 110, 3901–3906 (2013).
Shi, J. et al. The Polycomb complex PRC2 supports aberrant self-renewal in a mouse model of MLL-AF9;Nras(G12D) acute myeloid leukemia. Oncogene 32, 930–938 (2013).
Mertz, J.A. et al. Targeting MYC dependence in cancer by inhibiting BET bromodomains. Proc. Natl. Acad. Sci. USA 108, 16669–16674 (2011).
Dawson, M.A. et al. Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature 478, 529–533 (2011).
Shi, J. & Vakoc, C.R. The mechanisms behind the therapeutic activity of BET bromodomain inhibition. Mol. Cell 54, 728–736 (2014).
Findlay, G.M., Boyle, E.A., Hause, R.J., Klein, J.C. & Shendure, J. Saturation editing of genomic regions by multiplex homology-directed repair. Nature 513, 120–123 (2014).
Xu, B. et al. Selective inhibition of EZH2 and EZH1 enzymatic activity by a small molecule suppresses MLL-rearranged leukemia. Blood 125, 346–357 (2015).
Daigle, S.R. et al. Selective killing of mixed lineage leukemia cells by a potent small-molecule DOT1L inhibitor. Cancer Cell 20, 53–65 (2011).
Harris, W.J. et al. The histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 21, 473–487 (2012).
Lehnertz, B. et al. The methyltransferase G9a regulates HoxA9-dependent transcription in AML. Genes Dev. 28, 317–327 (2014).
Kim, W. et al. Targeted disruption of the EZH2-EED complex inhibits EZH2-dependent cancer. Nat. Chem. Biol. 9, 643–650 (2013).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Morita, S., Kojima, T. & Kitamura, T. Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther. 7, 1063–1066 (2000).
Acknowledgements
Y. Jin for assistance with pooled sgRNA screen analysis. C.R.V. is supported by National Institutes of Health NIH CA174793, Burroughs-Wellcome Fund Career Award for Medical Scientists, Alex's Lemonade Stand Foundation 'A' Award and the National Cancer Institute Cancer Center Support Grant Development Funds CA45508. J.B.K. is supported by the Simons Center for Quantitative Biology at Cold Spring Harbor Laboratory.
Author information
Authors and Affiliations
Contributions
J.S. and C.R.V. designed experiments; J.S., E.W. and J.P.M. carried out experiments; J.S. analyzed experimental results. J.B.K. analyzed sequencing data and developed analysis tools. Z.W. assisted with Illumina sequencing. J.S., J.B.K. and C.R.V. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Negative selection analysis of sgRNAs targeting all Brd4 exons comparing day 2 to day 10 time points.
Systematic evaluation of 64 Brd4 sgRNAs in negative selection experiments, targeting each Brd4 exon. The location of each sgRNA relative to the Brd4 protein is indicated along the x-axis. BD1: bromodomain 1, BD2: bromodomain 2, ET: extra-terminal domain, CTM: C-terminal motif. Plotted is the fold change of GFP positivity comparing day 2 and day 10 post-infection, representing the average of three independent biological replicates.
Supplementary Figure 2 SURVEYOR assay and deep sequencing analysis of indel mutations induced by various Brd4 or Smarca4 sgRNAs.
(a) Top panel, location of Brd4 sgRNAs relative to the domain architecture of the Brd4 N-terminus. Bottom panel, SURVEYOR assay of indel mutations at corresponding Brd4 genomic DNA regions. Analysis was performed at day 3 post-infection. sgRNA targeting ROSA26 locus serves as the negative control. The GFP+/sgRNA+ percentages of each sample at day 3 are labeled under the gel image. The indel% was calculated based on the relative intensity of the DNA bands using ImageJ software. The normalized indel% was calculated by dividing the indel% by the GFP%. Representative image of 2 independent experiments is shown. (b) SURVEYOR assay of indel mutations induced by Brd4 sgRNAs at the indicated timepoints post-infection. Mutations induced by e3.3 undergo stronger negative selection than mutations induced by e3.1. (c) Deep sequencing-based analysis of CRISPR-mediated mutagenesis efficiency at the indicated Brd4 sgRNA cut sites performed at various timepoints post-infection. Illumina sequencing was used to quantify indel mutations at the corresponding to the sgRNA cut site. The GFP% at these timepoints was used to determine the overall indel% in transduced cells. ND: Not determined since the GFP % was low due to sever negative selection. (d) SURVEYOR assay of indel mutations induced by Smarca4 sgRNAs at the indicated timepoints post-infection. Mutations induced by e16.1 and e16.2 undergo stronger negative selection than mutations induced by e2.1 and e3.1. M: marker. (e) Analysis of CRISPR-mediated mutagenesis efficiency at the indicated Smarca4 sgRNA cut sites performed at various timepoints post-infection. Illumina sequencing was used to quantify indel mutations at the corresponding to the sgRNA cut site. The GFP% at these timepoints was used to determine the overall indel% in transduced cells. ND: Not determined since the GFP % was low due to severe negative selection.
Supplementary Figure 3 Brd4 BD1 sgRNAs do not exhibit off-target mutagenesis of homologous BD1 domains of Brd2 and Brd3.
Analysis of CRISPR editing efficiency at the indicated BD1 domain-encoding exons of Brd2, Brd3, and Brd4, following transduction with Brd4 BD1 targeting sgRNAs e3.3 and e4.1. Analysis was performed at day 3 post-infection. The indel% was calculated based on the relative intensity of the DNA bands using ImageJ software. Results are representative of two independent biological replicates. M: marker. N.D.: not determined.
Supplementary Figure 4 Deep sequencing analysis of mutation abundance following CRISPR-targeting of different Smarca4 or Rosa26 regions.
(a-c) This analysis was performed on PCR-amplified genomic DNA corresponding to the sgRNA cut site at the indicated timepoints. Indel mutations were categorized into two groups: in-frame (3n) or frameshift (3n+1, 3n+2). Nonsense mutations were also included with the frameshift category, however such mutations were rare in this analysis. Green and red numbers indicate the number of in-frame and frameshift mutants that were tracked, respectively. For a and b, dots of the same color indicate the median normalized abundance at the indicated time point for all mutations within each group; shaded regions indicate the interquartile range of normalized abundance values. For c, the relative abundance of 50 individual ROSA26 indels (indicated as light-gray lines) at indicated timepoints normalized to day 3 abundance. The black line represents the median normalized abundance across all 50 mutations. For a and b, significant differences between the enrichment values of the in-frame and frameshift mutations were assessed using a Mann-Whitney-Wilcoxon test; ** indicates p < 0.01, and *** indicates p < 0.005. The normalized abundance of each tracked mutation was defined as the ratio of the number of observed mutant sequences divided by the number of wild-type sequences, normalized by the value of this same quantity at day 3.
Supplementary Figure 5 Deep-sequencing analysis of in-frame mutation frequency induced by various sgRNAs.
Across 12 different sgRNAs used in this study, deep sequencing analysis of mutations at day 3 indicates an average frequency of in-frame mutations (3n) of 29.4%, with the remaining indel mutations being frameshifts, which matches well the expected ratio and the observations of others
Supplementary Figure 6 A model illustrating the expected genotypes and mutational abundance observed upon CRISPR targeting of different regions of an essential protein.
(a, left) Model for anticipated genotypes upon CRISPR mutagenesis of a 5’ coding exon that lacks a functionally important domain, in which in-frame variants would retain functionality. If 33% of CRISPR mutations are in-frame and 66% are frameshift, then 4/9 of cells would be expected to have biallelic frameshift mutations, which would represent a homozygous null state. 5/9 of cells would carry at least one in-frame indel allele, which would retain functionality. This would render ~56% of cells in the population with a less severe phenotype. (a, right) The anticipated deep-sequencing based analysis of mutational abundance when targeting a 5’ coding exon that lacks a functional domain. Since each in-frame mutation will cause the cell it resides in to be phenotypically unaffected, the prevalence of each in-frame mutation (relative to the wild type allele) will remain constant over time. Each frameshift mutation, on the other hand, has a 1/3 probability of being paired with an in-frame mutation and a 2/3 probability of being paired with another frameshift. Cells will be phenotypically affected more strongly in the latter case. Therefore, the prevalence of each frameshift mutation will first decrease then plateau at a value of 1/3. More precisely, the relative prevalence of in-frame (Pif) and frameshift (Pfs) mutations as a function of time will be
The prevalence of both in-frame and frameshift mutation will decay at rate r.
(b, left) Model for anticipated genotypes upon CRISPR mutagenesis of an exon that encodes a functionally important domain, in which both in-frame and frameshift mutations will disable the protein. Nearly every cell in which both alleles are mutagenized will therefore lose the functionality of this protein and thus be phenotypically affected (b, right). The anticipated deep-sequencing based analysis of mutational abundance when targeting an exon encoding a functionally important domain. The prevalence of both in-frame and frameshift mutation will decay at rate r. This decay will ultimately plateau at a value of f, where f is the failure rate of CRISPR mutagenesis, due to CRISPR either not mutagenizing both alleles within the cell or producing a non-disruptive mutation in the unobserved allele. Specifically,
Supplementary Figure 7 Deacetylase domain-focused CRISPR-Cas9 screen in murine MLL-AF9/NrasG12D acute myeloid leukemia cells.
Summary of negative selection experiments with sgRNAs targeting the indicated domains plotted as fold change in GFP-positivity. Each bar represents the mean value of three independent biological replicates for an independent sgRNA targeting the indicated domain. The two deacetylase domains of HDAC6 are indicated as a1 and a2.
Supplementary Figure 8 Pooled sgRNA screen targeting lysine methyltransferase domains leads to similar results as analysis of individual sgRNAs using GFP reporters.
(a) Results of the pooled sgRNA screen evaluating lysine methyltransferase dependencies. The pooled library of sgRNAs was transduced into RN2c cells at a representation of ~500 transduced cells per sgRNA, followed by collection of genomic DNA at day 2 and day 12 post-infection. The sgRNA cassette was PCR-amplified from these samples and subjected to Illumina sequencing to measure the abundance of individual sgRNAs over time. The fold change in sgRNA abundance was calculated and plotted as the average of two independent biological replicates. Results were normalized to ROSA26 sgRNA. Red indicates the known drug targets within this class of regulators. The results closely match the findings obtained by scoring sgRNAs individually, shown in Figure 3. (b) Scatter plot that compares the fold change measurements between the two independent replicates.
Supplementary Figure 9 Lysine methyltransferase sgRNA screen performed in Cas9+ 38B9 cells (murine B-cell progenitor line) and in Cas9+ NIH3T3 cells (immortalized fibroblasts).
Cell lines were transduced with MSCV-Cas9-PGK-Puro followed by puromycin selection, prior to transduction with U6-sgRNA-EFS-GFP lentivirus. Summary of negative selection experiments with sgRNAs targeting the indicated domains plotted as fold change in GFP-positivity. A 20-fold cutoff was applied for visualization purposes.
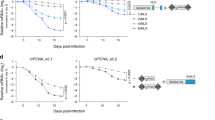
Supplementary Figure 10 Deep sequencing analysis of indel mutations induced by various Ezh2 or Dot1l sgRNAs.
Analysis of CRISPR-mediated mutagenesis efficiency at the indicated either Ezh2 or Dot1l sgRNA cut sites performed at various timepoints post-infection. Ezh2_e2.1 and Dot1l_e1.1 sgRNAs target 5’ coding exons. Ezh2_e19.2, Dot1l_e7.1, and Dot1l_e11.2 sgRNAs target methyltransferase domains. Illumina sequencing was used to quantify the CRISPR-induced indel mutations at the corresponding sgRNA cut site. The GFP% at these timepoints was used to determine the overall indel% in transduced cells. ND: Not determined since the GFP % was low due to sever negative selection.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Discussion (PDF 624 kb)
Supplementary Table 1
Supplementary Table 1 (XLSX 116 kb)
Rights and permissions
About this article
Cite this article
Shi, J., Wang, E., Milazzo, J. et al. Discovery of cancer drug targets by CRISPR-Cas9 screening of protein domains. Nat Biotechnol 33, 661–667 (2015). https://doi.org/10.1038/nbt.3235
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3235
This article is cited by
-
METTL16 promotes liver cancer stem cell self-renewal via controlling ribosome biogenesis and mRNA translation
Journal of Hematology & Oncology (2024)
-
TAZ2 truncation confers overactivation of p300 and cellular vulnerability to HDAC inhibition
Nature Communications (2023)
-
Chromatin complex dependencies reveal targeting opportunities in leukemia
Nature Communications (2023)
-
PAX3-FOXO1 uses its activation domain to recruit CBP/P300 and shape RNA Pol2 cluster distribution
Nature Communications (2023)
-
KDM6B protects T-ALL cells from NOTCH1-induced oncogenic stress
Leukemia (2023)