Abstract
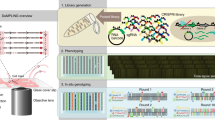
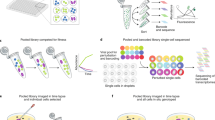
Multiplexed genome engineering approaches can be used to generate targeted genetic diversity in cell populations on laboratory timescales, but methods to track mutations and link them to phenotypes have been lacking. We present an approach for tracking combinatorial engineered libraries (TRACE) through the simultaneous mapping of millions of combinatorially engineered genomes at single-cell resolution. Distal genomic sites are assembled into individual DNA constructs that are compatible with next-generation sequencing strategies. We used TRACE to map growth selection dynamics for Escherichia coli combinatorial libraries created by recursive multiplex recombineering at a depth 104-fold greater than before. TRACE was used to identify genotype-to-phenotype correlations and to map the evolutionary trajectory of two individual combinatorial mutants in E. coli. Combinatorial mutations in the human ES2 ovarian carcinoma cell line were also assessed with TRACE. TRACE completes the combinatorial engineering cycle and enables more sophisticated approaches to genome engineering in both bacteria and eukaryotic cells than are currently possible.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Wang, H.H. et al. Genome-scale promoter engineering by coselection MAGE. Nat. Methods 9, 591–593 (2012).
Alper, H., Miyaoku, K. & Stephanopoulos, G. Construction of lycopene-overproducing E. coli strains by combining systematic and combinatorial gene knockout targets. Nat. Biotechnol. 23, 612–616 (2005).
Lee, J.W. et al. Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat. Chem. Biol. 8, 536–546 (2012).
Wang, B.L. et al. Microfluidic high-throughput culturing of single cells for selection based on extracellular metabolite production or consumption. Nat. Biotechnol. 32, 473–478 (2014).
Pál, C., Papp, B. & Pósfai, G. The dawn of evolutionary genome engineering. Nat. Rev. Genet. 15, 504–512 (2014).
Romero, P.A. & Arnold, F.H. Exploring protein fitness landscapes by directed evolution. Nat. Rev. Mol. Cell Biol. 10, 866–876 (2009).
Wang, H.H. & Church, G.M. Multiplexed genome engineering and genotyping methods applications for synthetic biology and metabolic engineering. Methods Enzymol. 498, 409–426 (2011).
Barrick, J.E. & Lenski, R.E. Genome dynamics during experimental evolution. Nat. Rev. Genet. 14, 827–839 (2013).
Sandoval, N.R. et al. Strategy for directing combinatorial genome engineering in Escherichia coli. Proc. Natl. Acad. Sci. USA 109, 10540–10545 (2012).
Woodruff, L.B. et al. Genome-scale identification and characterization of ethanol tolerance genes in Escherichia coli. Metab. Eng. 15, 124–133 (2013).
Warner, J.R., Reeder, P.J., Karimpour-Fard, A., Woodruff, L.B. & Gill, R.T. Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides. Nat. Biotechnol. 28, 856–862 (2010).
Kosuri, S. & Church, G.M. Large-scale de novo DNA synthesis: technologies and applications. Nat. Methods 11, 499–507 (2014).
Wang, X. et al. Engineering furfural tolerance in Escherichia coli improves the fermentation of lignocellulosic sugars into renewable chemicals. Proc. Natl. Acad. Sci. USA 110, 4021–4026 (2013).
Phillips, P.C. Epistasis—the essential role of gene interactions in the structure and evolution of genetic systems. Nat. Rev. Genet. 9, 855–867 (2008).
Khan, A.I. et al. Negative epistasis between beneficial mutations in an evolving bacterial population. Science 332, 1193–1196 (2011).
Craig, D.W. et al. Identification of genetic variants using bar-coded multiplexed sequencing. Nat. Methods 5, 887–893 (2008).
Tewhey, R. et al. Microdroplet-based PCR enrichment for large-scale targeted sequencing. Nat. Biotechnol. 27, 1025–1031 (2009).
Mamanova, L. et al. Target-enrichment strategies for next-generation sequencing. Nat. Methods 7, 111–118 (2010).
Metzker, M.L. Sequencing technologies—the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Casini, A. et al. One-pot DNA construction for synthetic biology: the Modular Overlap-Directed Assembly with Linkers (MODAL) strategy. Nucleic Acids Res. 42, e7 (2014).
Wetmur, J.G. et al. Molecular haplotyping by linking emulsion PCR: analysis of paraoxonase 1 haplotypes and phenotypes. Nucleic Acids Res. 33, 2615–2619 (2005).
Estep, A.L., Palmer, C., McCormick, F. & Rauen, K.A. Mutation analysis of BRAF, MEK1 and MEK2 in 15 ovarian cancer cell lines: implications for therapy. PLoS ONE 2, e1279 (2007).
Williams, R. et al. Amplification of complex gene libraries by emulsion PCR. Nat. Methods 3, 545–550 (2006).
Kanagawa, T. Bias and artifacts in multitemplate polymerase chain reactions (PCR). J. Biosci. Bioeng. 96, 317–323 (2003).
Pääbo, S., Irwin, D.M. & Wilson, A.C. DNA damage promotes jumping between templates during enzymatic amplification. J. Biol. Chem. 265, 4718–4721 (1990).
Volkmer, B. & Heinemann, M. Condition-dependent cell volume and concentration of Escherichia coli to facilitate data conversion for systems biology modeling. PLoS ONE 6, e23126 (2011).
Sawitzke, J.A. et al. Probing cellular processes with oligo-mediated recombination and using the knowledge gained to optimize recombineering. J. Mol. Biol. 407, 45–59 (2011).
Farasat, I. et al. Efficient search, mapping, and optimization of multi-protein genetic systems in diverse bacteria. Mol. Syst. Biol. 10, 731 (2014).
Minty, J.J. et al. Evolution combined with genomic study elucidates genetic bases of isobutanol tolerance in Escherichia coli. Microb. Cell Fact. 10, 18 (2011).
Goodarzi, H. et al. Regulatory and metabolic rewiring during laboratory evolution of ethanol tolerance in E. coli. Mol. Syst. Biol. 6, 378 (2010).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Na, D. et al. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat. Biotechnol. 31, 170–174 (2013).
Yoo, S.M., Na, D. & Lee, S.Y. Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nat. Protoc. 8, 1694–1707 (2013).
Jiang, W. et al. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Datta, S., Costantino, N., Zhou, X. & Court, D.L. Identification and analysis of recombineering functions from Gram-negative and Gram-positive bacteria and their phages. Proc. Natl. Acad. Sci. USA 105, 1626–1631 (2008).
Gevertz, J.L., Dunn, S.M. & Roth, C.M. Mathematical model of real-time PCR kinetics. Biotechnol. Bioeng. 92, 346–355 (2005).
Eklund, A.C., Friis, P., Wernersson, R. & Szallasi, Z. Optimization of the BLASTN substitution matrix for prediction of non-specific DNA microarray hybridization. Nucleic Acids Res. 38, e27 (2010).
Miura, F., Uematsu, C., Sakaki, Y. & Ito, T. A novel strategy to design highly specific PCR primers based on the stability and uniqueness of 3′-end subsequences. Bioinformatics 21, 4363–4370 (2005).
Caporaso, J.G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 108, 4516–4522 (2011).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Acknowledgements
Special thanks to S.A. Lynch and W.C. Grau for support and advice on this work. We would like to thank J. Huntley and S. Gao at the Colorado Biofrontiers Advanced Sequencing Core Facility for assistance with high-throughput sequencing. We would like to thank J. Liddle for generously sharing the ES2 cancer cell line. This work was funded by DOE BER Genomic Sciences Program Award Number DE-SC0008812. We would also like to acknowledge the C2B2/NSF REU Program (NSF#1261303) for funding G.D.D.
Author information
Authors and Affiliations
Contributions
R.I.Z., A.D.G. and R.T.G. conceived this idea. R.I.Z., A.D.G., G.P., T.J.M. and R.T.G. designed experiments. G.D.D. and R.I.Z. performed kinetic modeling. R.I.Z. performed experiments with assistance from G.P., T.Y.G. and N.R.B. R.I.Z. and R.T.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–22 and Supplementary Tables 1–16 (PDF 2221 kb)
Rights and permissions
About this article
Cite this article
Zeitoun, R., Garst, A., Degen, G. et al. Multiplexed tracking of combinatorial genomic mutations in engineered cell populations. Nat Biotechnol 33, 631–637 (2015). https://doi.org/10.1038/nbt.3177
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3177
This article is cited by
-
Synthetic biology advances and applications in the biotechnology industry: a perspective
Journal of Industrial Microbiology and Biotechnology (2018)
-
Upconversion Nanoparticles-Encoded Hydrogel Microbeads-Based Multiplexed Protein Detection
Nano-Micro Letters (2018)
-
Genome-wide analysis of E. coli cell-gene interactions
BMC Systems Biology (2017)
-
Optimizing complex phenotypes through model-guided multiplex genome engineering
Genome Biology (2017)
-
Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering
Nature Biotechnology (2017)