Abstract
Recent technical developments have enabled the transcriptomes of hundreds of cells to be assayed in an unbiased manner, opening up the possibility that new subpopulations of cells can be found. However, the effects of potential confounding factors, such as the cell cycle, on the heterogeneity of gene expression and therefore on the ability to robustly identify subpopulations remain unclear. We present and validate a computational approach that uses latent variable models to account for such hidden factors. We show that our single-cell latent variable model (scLVM) allows the identification of otherwise undetectable subpopulations of cells that correspond to different stages during the differentiation of naive T cells into T helper 2 cells. Our approach can be used not only to identify cellular subpopulations but also to tease apart different sources of gene expression heterogeneity in single-cell transcriptomes.
Similar content being viewed by others
Main
Single-cell measurements of gene expression, using imaging techniques such as RNA-FiSH (fluorescence in situ hybridization), have provided important insights into the kinetics of transcription and cell-to-cell variation in gene expression1,2,3. However, such approaches can examine the expression of only a small number of genes in each experiment, thus restricting our ability to examine co-expression patterns and to robustly identify subpopulations of cells. Protocols have been developed to overcome these limitations by amplifying small quantities of mRNA4,5, which, in combination with microfluidics approaches for isolating individual cells6,7, have been used to analyze the co-expression of tens to hundreds of genes in single cells8,9. These protocols also allow the entire transcriptome of large numbers of single cells to be assayed in an unbiased way. This was initially done using microarrays10,11 but is more often now done using next-generation sequencing12,13,14,15. Such approaches have been used to model early embryogenesis in the mouse16 and to investigate bimodality in gene expression patterns of differentiating immune cell types17.
After the generation of single-cell RNA-sequencing (RNA-seq) profiles from hundreds of cells, one goal to identify subpopulations that share a common gene-expression profile. Some of these subpopulations may represent previously unidentified cell types. Additionally, by studying patterns of gene expression in different single cells, insights into the regulatory landscape of each cell population can be obtained.
However, methods for identifying subpopulations of cells and modeling their gene regulatory landscapes are only now beginning to emerge18,19. To fully exploit single-cell RNA-seq data, we have to account for the random noise inherent to such data sets20 and, equally important, to account for different hidden factors that might result in gene expression heterogeneity. Although the importance of accounting for unobserved factors is well established in bulk RNA-seq studies21,22,23, robust approaches to detect and account for confounding factors in single-cell RNA-seq studies remain to be developed. Here, we describe a computational approach that uses latent variable models to reconstruct such hidden factors from the observed data. We validate our scLVM using a population of staged mouse embryonic stem cells (mESCs), before applying it to study T helper 2 (TH2) cell differentiation. We show that scLVM facilitates the identification of physiologically meaningful subpopulations of cells, which cannot otherwise be found.
Results
Cell cycle variation affects global gene expression
Single-cell RNA-seq is now commonly used to study cell differentiation15,24. Here, we reanalyzed data from a single-cell RNA-seq experiment that was originally designed to study the differentiation of naive T cells into TH2 cells25. Briefly, a population of naive Cd4+ T helper cells were activated and polarized with interleukin (IL)-4 to induce differentiation toward a TH2 subtype. At 4.5 d post-stimulation, cells were sorted into a G4P group (fourth generation, IL-13–GFP+ cells) and a G2N group (second generation, IL-13–GFP− cells). Subsequently, these two groups of cells were pooled in equal proportions. From this pool, a set of 96 asynchronously dividing cells (including both fully and partially differentiated cells) was captured using the Fluidigm C1 system, and sequencing libraries were prepared and processed. After quality control and accounting for technical noise, RNA-seq data for 81 cells and 7,073 genes with variation in their expression level above technical noise were considered for analysis (Supplementary Fig. 1).
The cell cycle is known to have wide-ranging effects on cellular physiology26,27 and can modulate both differentiation and gene expression profiles28 (Fig. 1a). Cells that are analyzed during development are likely to be in different stages of the cell cycle28. When we examined sets of genes whose expression is known to be associated with different cell-cycle stages, we observed that their expression levels varied considerably among single cells (Supplementary Fig. 1). Although variation in gene expression that is linked to the cell cycle can provide important biological insights, in many contexts such variation might mask other more physiologically important differences in gene expression between cells.
(a) The observed expression profile of differentiation marker genes (upper panel) is the result of the differentiation process of interest together with the effects of the cell cycle and other confounding sources of variation. After accounting for cell-cycle effects (middle panel), one can uncover gene expression signatures that contribute to the continuous differentiation process more clearly (lower panel). (b) scLVM two-stage procedure. First, in the fitting stage, the cell-to-cell covariance matrix that corresponds to the cell cycle is inferred from the gene expression profiles of genes with cell-cycle annotation (upper panel). The learnt covariance is then used in downstream analyses, including the detection of substructure, the detection of gene-to-gene correlations and the analysis of variance (lower panel). Biol. var., biological variance; Tech. var., technical variance.
Importantly, variation in gene expression that is linked to the cell cycle is not restricted to well-annotated cell-cycle marker genes. When we examined a set of moderately to highly variable genes that have not previously been associated with the cell cycle, we observed that 2,881 genes (44%) showed a significant correlation of gene expression with at least one cell-cycle gene (P < 0.05, Bonferroni adjusted; Supplementary Fig. 2). Therefore, merely removing the set of annotated cell-cycle genes before performing downstream analyses is likely to be unsuccessful because it would not enable all effects independent of the cell cycle to be detected.
Development of scLVM to account for effects of the cell cycle
We used scLVM to address the confounding effects of the cell cycle. In this type of computational approach, one first reconstructs the cell-cycle state (or other unobserved factors) and then uses this information to infer 'corrected' gene expression levels. This two-step approach enables the effect of unobserved factors on gene expression heterogeneity to be accounted for in downstream analyses, thereby allowing us to study variation in gene expression levels that is independent of the cell cycle. Moreover, for each gene whose expression is analyzed, our method allows the relative contribution of any reconstructed factors that affect cell-to-cell variation in expression to be determined. A schematic overview of the approach is shown in Figure 1b.
To validate our method, we generated single-cell RNA-seq data from mESCs using the Fluidigm C1 protocol, where the cell-cycle status of each cell is known a priori. We assayed the transcriptional profile of 182 ESCs that had been staged for cell-cycle phase (G1, S and G2M) based on sorting of the Hoechst 33342-stained cell area of a flow cytometry (FACS) distribution. In the fitting stage, scLVM uses the expression profiles of a relatively small set of 892 annotated cell-cycle genes (Supplementary Table 1) to recover a covariance matrix that accounts for cell-to-cell heterogeneity due to the cell cycle (Supplementary Fig. 3). Using alternative annotations for cell-cycle genes (Supplementary Table 1) yielded very similar results (Supplementary Figs. 3, 4, 5). Subsequently, for all remaining genes, we used scLVM to estimate the proportion of variance in expression across cells that is explained by technical noise, biological variability and cell cycle. This approach can also be used to create a 'corrected' gene expression data set, in which the effect of the identified factor(s) is removed, which can be used as the input for existing analysis methods. scLVM is related to approaches for modeling variability in bulk mRNA expression studies21,22 and to methods used in genome-wide association studies in which the relatedness between individuals is inferred from genotype29 and/or expression levels30 and then accounted for in downstream analyses using linear mixed models.
As the cell-cycle stage of each cell is known in our data set, we can compare the scLVM estimates of the proportion of variance explained by the cell cycle with the gold standard values obtained when using the annotation of individual cells based on the Hoechst staining (FACS). We observed a striking correlation (r2 = 0.91) between our scLVM estimates and the gold standard values, providing confidence in the efficacy of our approach (Fig. 2a). The model fit and these estimates for the variance explained by the cell cycle were consistent when a much smaller gene set containing only tens of genes was used to train the model (Supplementary Fig. 5a–g) and when alternative metrics were applied to quantify the proportion of variation explained by the cell cycle (Supplementary Fig. 5h). This suggests that scLVM can be used to robustly recover and estimate the variance due to unobserved factors from relatively small gene sets that annotate these factors. Additionally, we examined how many pairs of genes had significantly correlated patterns of expression across cells (i) without cell-cycle correction, (ii) with the scLVM correction and (iii) with an ideal correction using the gold standard cell-cycle state. The set of significant gene-gene correlations obtained with the scLVM correction was much more consistent with a gene correlation network based on the experimental staging than the set generated under the no-correction model (Supplementary Fig. 6), with the number of false-positive correlations reduced by three orders of magnitude (from 72,117 to 77). Finally, we compared the scLVM correction to a basic removal strategy, in which cell cycle–annotated genes (892 genes, Supplementary Table 1) were omitted from the analysis. A nonlinear principal component analysis (PCA)31 on the data set from which cell-cycle annotated genes were removed yielded a clear separation of cells according to cell-cycle stage (Fig. 2b). In contrast, when repeating the analysis using scLVM-corrected gene expression levels, the same separation of cells was not observed, showing that the cell cycle–related expression signature was effectively removed (Fig. 2c). Further, to show that scLVM is specific in removing the effects of cell cycle–related variation, we considered a noncycling cell type (terminally differentiated neurons) as a negative control. Reassuringly, scLVM attributed more than 30% of variation to the cell cycle for only 27 genes, and the maximum proportion of variation attributed to the cell cycle for any single gene was 37%. In comparison, when we applied scLVM to cycling T cells, for 1,895 genes, more than 30% of variation was attributed to the cell cycle, with the maximum proportion for any single gene being 79% (Fig. 3a and Supplementary Fig. 7). These results give additional confidence that the variance estimates are accurately inferred. Finally, we repeated the validation of scLVM using a second previously published data set of 35 mESCs staged for the cell cycle, but prepared for sequencing with an alternative protocol (Quartz-Seq)32 and cultured under different media conditions that are known to induce reduced variability in expression of cell-cycle genes33. Again, direct comparison of variance estimates from scLVM with the gold standard derived from the staging information of individual cells yielded good agreement (Supplementary Figs. 8 and 9). To assess the consistency of the expression signatures that are used by scLVM to infer the cell-to-cell covariance, we projected the 35 mESCs from this published data set onto the larger mESC validation data set discussed above. This analysis revealed that the expression signatures due to the cell cycle are robust across sequencing protocols, studies and experimental batch (Supplementary Fig. 10). In sum, these analyses provide confidence that our scLVM approach effectively accounts for latent factors such as the cell cycle.
(a) Comparison of the estimated proportion of variability in the expression of each gene across cells due to the cell cycle as inferred using scLVM (x axis) or with gold standard estimates of the cell-cycle stage derived from the Hoechst staining (y axis). The scatter plot compares the proportion of variance explained by either approach, revealing striking concordance (Pearson's r2 = 0.91). (b,c) Nonlinear PCA based on genes not annotated as cell cycle (neither GO nor Cyclebase) (b) and the same nonlinear PCA process carried out using scLVM-corrected gene expression data (c). Cell-cycle annotation of individual cells according to the Hoechst staining is color coded. In the uncorrected expression data, the PCA analysis separates cells according to their cell-cycle stage, even when omitting cell-cycle genes. This clear separation is lost when using scLVM-corrected expression levels, showing that scLVM effectively removes gene expression signatures that are only associated with cell-cycle effects.
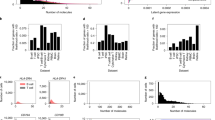
(a) For each gene, scLVM was used to estimate the proportion of variance explained by the cell cycle, technical noise and residual biological variance. Genes were binned by the total variance explained by factors other than technical noise; the bars show average variance contributions for genes in a particular bin. (b) Gene-to-gene variation without cell-cycle correction. (c) Gene-to-gene variation with cell-cycle correction. Shown are −log10 P values from a correlation test between pairs of genes (pv). Without cell-cycle correction, widespread gene-gene correlations were observed. The scLVM correction greatly reduced this background correlation structure (622,769 versus 17,389 correlations, involving 2,053 versus 143 genes; P < 0.05, Bonferroni adjusted). GO analysis revealed that unlike the gene-gene correlations without correction, the significant correlations after scLVLM correction were enriched for plausible functional categories (Supplementary Table 3). (d) Nonlinear PCA applied to the expression data set without cell-cycle correction. The color overlaid on each cell denotes the log10 expression of Gata3 in that cell. (e,f) Nonlinear PCA applied to the expression data set with cell-cycle correction. The color overlaid on each cell denotes the cell cycle–corrected log10 expression of Gata3 in that cell. The corrected data set revealed two distinct subclusters of cells, between which Gata3 (e) and, other factors important for proper TH2 differentiation including receptor genes, cytokines and transcription factors (f) were differentially expressed (all P values < 0.001; Supplementary Fig. 13).
Application of scLVM to identify cell populations in differentiating TH2 cells
We next applied our scLVM approach to study a population of asynchronously differentiating TH2 cells that have previously been profiled using single-cell RNA-seq20,25. We observed that the cell cycle contributed markedly to gene expression variability, in particular for the set of genes with medium to high overall nontechnical variability (Fig. 3a). Genome-wide, for 1,895 (27%) of these variable genes, the cell cycle accounted for more than 30% of the variance in expression across cells (Fig. 3a, Supplementary Figs. 11 and 12, and Supplementary Table 2), suggesting that the expression of many genes is affected by the cell cycle. When comparing the expression signatures of cell-cycle genes in the TH2 cell type with those found in the mESC validation data set, we found striking agreement of the main axes of variation (PC1, r2 = 0.998, Supplementary Fig. 10). This result strongly suggests that scLVM robustly captures cell cycle effects in the TH2 data set.
Turning next to the question of whether pairs of genes show patterns of correlation between cells (gene-gene correlations), we observed a striking decrease in significant correlations after accounting for the cell cycle (P < 0.05, Bonferroni adjusted; Fig. 3b,c and Supplementary Fig. 11). This suggests that many of the gene-gene correlations observed in the initial data were driven by cell-cycle stage. Notably, the much smaller set of genes with significant correlation patterns after correction was enriched for genes involved in glycolysis34 and for genes that mark the cellular response to IL-4 stimulation (Supplementary Table 3a), both of which are key processes in TH2 cell differentiation. In contrast, gene-gene variation obtained using uncorrected data yielded no enrichment for variation in expression of genes involved in glycolysis but instead identified genes that were enriched for cell cycle–related categories (Supplementary Table 3b), again indicating that cell cycle, if not accounted for, is a major confounder of gene-gene correlations.
Next, we examined whether the cell-cycle correction facilitated by the scLVM model enabled a more reliable identification of subpopulations of cells. Without correction, a nonlinear PCA31 revealed little structure in the data, with no obvious subgroups of the cells identified (Fig. 3d and Supplementary Fig. 11). A similar lack of structure was observed when other clustering algorithms, including hierarchical and k-means clustering, were applied (data not shown). However, when applying the same nonlinear PCA approach to the cell cycle–corrected data, two clear subpopulations of cells were identified (Fig. 3e; see Supplementary Data 1 for assignment of cells to clusters).
To investigate whether these two populations correspond to physiologically distinct subsets, we studied the set of 401 genes with significant differences in expression between the clusters (P < 0.05, Bonferroni adjusted; Supplementary Table 4). This set was heavily enriched for genes that have important roles in TH2 cell differentiation—Il4ra35, Gata3 (ref. 36), Stat3 (ref. 37), Klf13 (ref. 38), Batf (ref. 39) (P < 0.0001, Bonferroni adjusted) and Il24 (ref. 40) (P = 0.01) are all upregulated in the right-hand cluster (Fig. 3e), suggesting that cells contained in that group represent fully differentiated TH2 cells, whereas the left-hand population of cells correspond to a group that is only partially differentiated (Fig. 3e,f and Supplementary Fig. 13).
Consistent with this observation, an analysis of 122 manually curated 'TH2 signature' genes (Supplementary Table 5) revealed a significant enrichment in the set of 401 genes that were differentially expressed between the identified clusters (P = 0.001, Hypergeometric Test). Further, Gene Ontology (GO) enrichment analysis showed that the differentially expressed genes contained statistically significant enrichments of genes involved in glycolysis, cellular response to IL-4 stimulation and positive regulation of B-cell proliferation (Supplementary Table 6). To establish whether the genes distinguishing the two clusters act in a coordinated manner, we studied their interactions using the STRING database41. This yielded a densely connected network with three major hubs, which were highly enriched for glycolysis, translational elongation and T-cell activation, respectively (Supplementary Fig. 14). With glycolysis being a hallmark for T-cell activation42 and T-cell activation being linked to increased translational activity43, this provides further evidence that the two clusters contain cells at different positions along the trajectory to becoming fully differentiated TH2 cells.
Importantly, the cell-cycle correction afforded by scLVM not only enabled identification of two cell populations, but was also required for characterizing the two clusters. Testing for differential expression between the two identified populations of cells using the uncorrected data yielded only 7 genes whose transcription differed significantly between clusters (compared to 401 with the correction).
Accounting for more than one factor
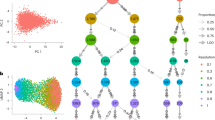
The scLVM approach can be applied to account for the effects of other factors, provided that an informative gene set is available. As an example, we extended the analysis of the TH2 cells by simultaneously modeling the cell-cycle state and the TH2 differentiation process as distinct factors. We used a set of 122 manually curated TH2 signature genes (Supplementary Table 5), introduced earlier, to fit a TH2 differentiation factor after removing the effects of the cell cycle. Although, in general, inference of multiple factors is statistically challenging, the much stronger effect of the cell-cycle factor helps to ensure that inference results are robust when considering different approaches (Supplementary Fig. 15; see Online Methods for a discussion of practical challenges). The joint analysis with both factors offered a more fine-grained decomposition of expression variability, attributing expression variation of individual genes to cell-cycle effects, TH2 differentiation and interactions between both factors (Fig. 4a). The interaction component allows genes that are associated with TH2 differentiation in a cell-cycle-stage-specific manner to be identified (Fig. 4b). Although the overall variance due to these interaction effects was small, a set of 375 genes with strong interactions (explained variance >5%; Supplementary Table 7) contained prominent candidates for effectors of the interplay between the cell cycle and TH2 differentiation. Several TH2 differentiation markers, including Batf and Il2ra, were among these genes (Fig. 4b), and this set was enriched for positive cell proliferation and negative regulation of apoptosis (Supplementary Tables 8 and 9). This finding is consistent with the known link between differentiation and cell proliferation in T helper cells44,45.
(a) For each gene, the proportion of variance explained by the cell cycle, TH2 differentiation, a multiplicative interaction between cell cycle and TH2 differentiation, as well as technical noise and residual biological variance was estimated. Genes were binned by the total variance explained by factors other than technical noise; the bars show average variance contributions for genes in a particular bin. (b) Visualization of the identified interaction between factors for cell cycle and TH2 differentiation for the gene Batf. Shown is the expression level of Batf (y axis) as a function of the inferred cell-cycle stage (x axis), where the level of the TH2 factor is encoded in color. The interaction between the cell cycle factor and the TH2 factor can be viewed as the conditional correlation between cell cycle and Batf expression. For fully differentiated cells (high TH2 factor), there is a strong correlation between cell cycle and gene expression (red dashed line, steep slope). In contrast, for partially differentiated cells (negative TH2 factor) this observed correlation is much weaker (dashed blue line, shallow slope). (c) Effect of accounting for different hidden factors on gene-gene correlations. The number of significant edges in the gene-gene correlation network (P < 0.05, Bonferroni adjusted) decreased by over an order of magnitude after correcting for cell cycle; subsequently accounting for TH2 differentiation resulted in a similar reduction of gene-gene correlations. Finally, accounting for the interaction between TH2 and cell cycle yielded an additional reduction of almost 50% of the remaining gene-gene correlations, suggesting that cell cycle and TH2 differentiation are the predominant source of gene-gene correlations in this data set.
Additionally, we investigated the relevance of the TH2 factor when testing for gene-gene correlation networks (Fig. 4c). The number of significant gene-gene correlations decreased markedly when including the additional TH2-related factors (from 17,389 to 2,077), suggesting that the cell cycle, TH2 differentiation and their interactions are the predominant sources of variation in this population of cells.
In summary, accounting for cell cycle–related variation by using scLVM is necessary both for identification and characterization of distinct populations of T cells that are at different stages of differentiation into mature TH2 cells. We also applied scLVM to other single-cell RNA-seq data sets, including 34 human embryonic stem cells and a set of 90 cells from human preimplantation embryos15, which confirmed that the cell cycle explains substantial proportions of the variability in other contexts. Moreover, correcting for cell cycle as a confounder revealed otherwise hidden structure that might correlate with different cell populations in these independently generated, single-cell RNA-seq data sets (Supplementary Figs. 16 and 17).
Discussion
We have shown how heterogeneity in gene expression in single cells due to factors such as the cell cycle can compromise the interpretation of single-cell RNA-seq experiments. To overcome this problem we present a computational approach that effectively accounts for these confounding factors. This method (Fig. 1) builds on existing approaches for modeling gene expression heterogeneity in bulk data22,30, which we here adapt to single-cell transcriptomics. We have validated our method using a large mESC data set in which the cell-cycle stages of individual cells are known a priori (Fig. 2) and demonstrated the utility of our approach by applying it to obtain insights into TH2 cell differentiation (Fig. 3). We treated the cell cycle as a confounding variable in our study, but cycling-related processes may be of high interest in other contexts. This is exemplified in the analysis of the interaction between the effects of the TH2 differentiation factor and the cell cycle (Fig. 4). More generally, scLVM allows the user to model and account for latent factors of other predefined sets of genes, enabling the sources of variation in a wide range of single-cell RNA-seq experiments to be studied. Our analysis of the TH2 differentiation process uses a nonlinear PCA approach to uncover the differentiation structure. More generally, scLVM can be used to remove variation due to the cell cycle and other confounding factors before applying alternative downstream analytic strategies, such as Monocle18.
One important challenge when multiple confounding factors are considered is to ensure that the model remains statistically identifiable, such that the effect of each individual factor can be robustly estimated. This may be of particular concern if multiple weak and nonindependent factors are present. Finally, we note that there remain open questions regarding the best way to process single-cell RNA-seq data46. In particular, our scLVM approach could be refined in several ways. For example, statistics to formally test for the presence of a particular factor might be warranted and scLVM could also be coupled with methods to reconstruct pseudo-temporal trajectories18. Also, comprehensive methods to properly normalize RNA-seq data within and across multiple independent single-cell transcriptome experiments are an important area of future work.
Methods
Data sets and processing.
Mouse ESC data. A detailed description of cell culture, Hoechst staining, single-cell capture and mRNA sequencing as well as quality control can be found in the Supplementary Notes and Supplementary Figure 18. In brief, Rex1-GFP–expressing mESCs (Rex1-GFP mESCs) were cultured on gelatin-coated dishes using serum-free NDiff 227 medium (Stem Cells Inc.) supplemented with 2i inhibitors. Hoechst staining (Hoechst 33342; Invitrogen) was optimized for Rex1-GFP mESC, and cells were sorted using FACS (MO-FLO XDP; Beckmann Coulter) for respective cell-cycle fractions (G1, S and G2M phase). Single-cell RNA-seq was done using the C1 Single Cell Auto Prep System (Fluidigm; 100-7000). After normalization and estimation of technical noise using ERCC spike-ins (see RNA-seq normalization and estimation of technical noise), we retained a set of 9,571 genes for analysis with variation above the technical background level (FDR <0.1; Supplementary Data 1).
To account for errors in the assignment of a cell-cycle phase using the Hoechst staining (e.g., due to cells cycling after FACS sorting), we performed an additional filtering step based on the ERCC spike-ins. We reasoned that for cells within a cell-cycle phase, the ratio of endogenous reads to total mapped reads—which can be interpreted as a proxy for cell size—should follow a narrow distribution. Therefore, we excluded cells where the difference between this ratio and its median within a cell-cycle phase exceeded one median absolute deviation (Supplementary Fig. 19). This resulted in a filtered set of 59 cells in G1 phase, 58 cells in S phase and 65 cells in G2M phase. Analysis results for the unfiltered data are shown in Supplementary Figure 20 (see Supplementary Notes for full details), leading to consistent overall conclusions.
Mouse ESC data (Quartz-Seq protocol). We used the normalized data and counts from the primary publication32. These data consist of gene expression level estimates, obtained using the Quartz-Seq protocol, for 35 mESCs, where the cell-cycle state of each cell is known a priori (7 S, 8 G2M and 20 G1 cells). FACS sorting the distribution of the Hoechst 33342-stained cell area with gates corresponding to G1, S and G2/M phases was used to establish the cell-cycle state before processing. In this particular data set, technical noise cannot be reliably estimated owing to the lack of spike-ins. Consequently, we estimated the amount of technical (null) noise expected for genes with variable levels of expression using a log-linear fit between the expression mean and the squared coefficient of variation between cells, approximating the typical fitting procedure when spike-ins are available (Supplementary Fig. 1b). This approach yielded a total of 5,546 highly variable genes (FDR< 0.1; see RNA-seq normalization below).
T-cell data. Generation of the T-cell data has been described in detail previously20,25. In brief, untouched Naïve CD4+ cells from spleens of IL-13eGFP Balb/c mice were negatively selected and differentiated toward TH2 in anti-CD3/CD28 coated plates. CellTrace Violet staining was performed according to manufacturer's instructions. After 4.5 d of activation cells were sorted according to presence/absence of GFP and number of cell division. In particular GFP-Negative cells that had undergone 2 cycles of cell division and GFP-Positive cells that had divided 4 times were then pooled in 1:1 ratio and loaded on a C1 machine for capturing. Duplets and cells with low yield or poor quality cDNA were removed, yielding 81 cells for analysis. After normalization and estimation of technical noise using ERCC spike-ins (see RNA-seq normalization and estimation of technical noise), we retained a set of 7,073 genes for analysis with variation above the technical background level (FDR <0.1; Supplementary Data 2).
RNA-seq normalization and estimation of technical noise. For the T-cell data, raw read counts were normalized using the approach proposed in DESeq47, deriving size factors for each cell from the ERCC spike-ins. Estimates of the technical variability were also derived using the ERCC spike-ins, adapting the approach in Brennecke et al.20 (Supplementary Fig. 1a). We omitted the normalization for cell size as proposed previously20 because the computational correction by scLVM yielded much better results (Supplementary Fig. 21). This is likely explained by noting that cell size and cell cycle are correlated, thus the normalization proposed by Brennecke et al. reduces the amount of information available for inferring cell-cell correlations due to cell cycle; see also Supplementary Figure 21 and discussion in Supplementary Notes. To determine genes with high biological variability, we followed Brennecke et al.20 and tested against the null hypothesis that the biological coefficient of variation is at most 50% (at 10% FDR, Supplementary Notes). This justifies ignoring Poisson shot noise because of the large proportion of technical noise of genes expressed at low levels (see ref. 20 and details below). For the Quartz-Seq mESC data no spike-ins were available; we therefore used fragments per kilobase of transcript per million fragments mapped (FPKM) expression estimates as provided by the authors. Because there were no spike-ins, we estimated the baseline variability using a log-linear fit to describe the relationship between mean and squared coefficient of variation overall (Supplementary Fig. 1c). All subsequent analyses were carried out on log-transformed normalized count values and log-transformed FPKM estimates for the T-cell and newly generated mESC data and the Quartz-Seq mESC data, respectively.
scLVM method.
The scLVM algorithm is a two-step approach. First, one or more covariance structures are inferred from genes that are annotated to hidden factors such as cell-cycle progression. Subsequently, these covariance structures can be used to account for the hidden factors as random effects in a mixed model, allowing the variance in expression for each gene to be decomposed into a technical, a biological and a separate component for each hidden factor. Additionally, the hidden factors can be accounted for when performing pairwise gene-gene correlation analyses, and further allow 'corrected' residual gene expression data sets to be generated. scLVM is closely related to previous approaches that correct for hidden confounding factors in gene expression data21,48 and the inference employed to fit hidden factors builds on the PANAMA model30.
Briefly, the fitting process uses Gaussian Process Latent variable models (GPLVMs)31, a recent development in machine learning and statistics. The approach resembles a PCA on genes annotated to a hidden factor (such as cell cycle). However, instead of explicitly reconstructing PCA loadings and scores, the GPLVM approach fits a low-rank cell-to-cell covariance to the observed gene expression matrix of these genes. Related approaches have been proposed to account for relatedness between individuals in the context of expression Quantitative Trait Loci (eQTL) studies30, where an individual-to-individual covariance is inferred to explain the heterogeneity in gene expression levels between individuals rather than cells.
More specifically, for any gene g that is annotated to the hidden factor under consideration, its expression profile yg across cells is modeled as

where X represents the hidden factor (such as cell cycle), C corresponds to additional observed covariates (if available) and

denotes the residual variance. Because the same distributional assumptions are shared across a large set of genes in the annotated set, the state of the hidden variables X and the remaining covariance parameters can be robustly inferred by means of standard maximum likelihood approaches (Supplementary Notes). Once X is inferred, we calculate the covariance structure between cells, which is induced by the hidden factor as Σ = XXT .
An important choice when fitting the model is the dimensionality of the hidden factor matrix, X , which corresponds to the rank of the cell-to-cell covariance matrix Σ. In the context of distinct factors such as the cell cycle or TH2 differentiation, we found that a one-dimensional factor (rank one covariance) is commonly sufficient (see also the scree plots in Supplementary Fig. 22 and Choosing the rank of the cell-cycle factor). In general, the P-value distribution of a test statistic on the residual data set48, heuristic selection approaches21 or hierarchical modeling to regularize the effective dimensionality of the hidden factor22,30 can also be employed (Supplementary Notes).
Alternative fitting approaches, including methods to account for multiplicative effects between covariates and hidden factors, are discussed in the Supplementary Notes. Once fitted, the covariance matrix Σ can be used for a range of analyses, using efficient implementations of linear mixed models29,49 to decompose variance, test for gene-gene correlations or produce residuals corrected for the latent factors under consideration.
Analysis of variance. To estimate the components of variance, scLVM employs a linear mixed model that is fitted to the expression levels of each gene, decomposing sources of variation. Contributions from hidden factors such as cell-cycle effects, technical noise and residual biological variation to the observed expression variability of gene g are modeled as random effects:

with

denoting the variance attributable to H hidden factors (see section below for a discussion of estimating multiple hidden factors), residual biological variability (not related to hidden factors) and technical noise/baseline variability respectively. The hidden factor covariance matrices Σh are estimated in the GPLVM step and

is estimated from spike-ins as described above. The parameters

are then estimated by maximum likelihood. Interactions between pairs of factors can be considered by combining their previously estimated covariance matrices; see section above and Supplementary Notes.
Gene-gene correlation analysis. To estimate pairwise correlation coefficients while controlling for hidden factors such as the cell cycle, we introduce an additional fixed effect representing the contribution of another gene j

In this linear mixed model, βij can be interpreted as the pairwise correlation coefficient between genes i and j, and its significance can be assessed by means of a standard likelihood ratio test. Owing to efficient implementations of mixed models in applications to GWAS29,49, these correlation tests are extremely efficient (Supplementary Notes).
Creating residual expression data sets with the effect of hidden factors removed. To facilitate reuse of existing analyses methods, such as clustering, visualization or dimension reduction approaches, scLVM facilitates generation of a corrected expression data set where the effect of one or multiple hidden factors (e.g., the cell cycle) is removed.
For each gene i, the variance component model (see above) implies a predictive distribution of the cell-cycle component with mean ŷi and predictive variance  . Expression levels that are corrected for the effect of hidden factors can then be obtained from the model residuals, that is, yi* = yi − ŷi. These corrected gene expression values can be used in the full range of existing methods, including clustering or nonlinear PCA31. Cell-cycle corrected expression values for the T-cell data and the mESC data are available online (Supplementary Data 1 and 2).
. Expression levels that are corrected for the effect of hidden factors can then be obtained from the model residuals, that is, yi* = yi − ŷi. These corrected gene expression values can be used in the full range of existing methods, including clustering or nonlinear PCA31. Cell-cycle corrected expression values for the T-cell data and the mESC data are available online (Supplementary Data 1 and 2).
Applying scLVM to multiple annotated gene sets. In some circumstances, scLVM can be used to fit more than a single factor, provided that multiple informative gene sets are available. In general, statistical identifiability is a major concern and careful choice of the inference approach is important (Supplementary Notes). These factors can either be considered independently or learned by conditioning on one of them if prior knowledge exists as to which has a stronger effect (as for the cell cycle). As the cell cycle represents the predominant source of variation in our data, the cell-cycle factor can be recovered irrespective of other sources of variation. Therefore, we first learn the cell-cycle factor Xcc as described above. Then we extend the single factor model by conditioning on the inferred factor Xcc and including an interaction term, which we define by a point-wise product (Supplementary Notes). Alternative analysis approaches are discussed below; see Alternative approaches to fit the TH2 factor and Supplementary Figure 15.
Analysis details.
Validation of scLVM using mESC data. We used the annotated cell-cycle state in the ESC data sets to validate the accuracy of the model-based cell-cycle reconstruction carried out by scLVM. Briefly, instead of fitting the scLVM covariance from the data, we used the known grouping of cells into G1, S- and G2/M-phase as covariates and estimated the proportion of variance explained by the sum of all three covariates. In the case of the unfiltered mESC data (this study used the C1 protocol; see Data set and processing), the variance estimates of scLVM were compared to a model with a cell-cycle covariance and an additional factor that explains cell size variation. To estimate cell size we used the ratio of endogenous reads to total mapped reads, thereby capturing large variations in cell size within individual cell-cycle phases in the unfiltered data (Supplementary Notes). These estimates were then compared with the variance estimates using scLVM (Fig. 2a). In the same vein, the covariates can be included in pairwise gene-gene correlation analyses, again comparing the inference results based on the Hoechst staining to estimates obtained using the covariance structure inferred by scLVM. Further details on the estimation procedure using the gold standard are provided in Supplementary Notes.
Assessment of the effect of alternative cell cycle gene annotations. Unless otherwise stated, we considered the union of genes from CycleBase, and GO categories annotated as cell cycle related, resulting in 892 genes. Briefly, we combined all cell cycle–annotated genes (GO:0007049) in the Gene Ontology database along with the 600 top-ranked genes from CycleBase (Supplementary Notes). To assess to what extent the gene set annotation affects the performance of scLVM, we additionally considered either CycleBase genes or the GO annotated genes alone (Supplementary Figs. 3, 4, 5 and Supplementary Table 1), which yielded very similar results. Furthermore, we carried out a subsampling experiment, where random subsets of the full set of 892 genes were used to fit the cell cycle factor (Supplementary Fig. 5a–g). This showed that a relatively small set of 50 genes is sufficient to robustly identify the cell cycle. Finally, estimates for the variance explained by the cell cycle were consistent when alternative metrics were applied to quantify the proportion of variation explained by the cell cycle (Supplementary Fig. 5h and Supplementary Notes).
Identification of subclusters in the ESC and T-cell data. We considered nonlinear PCA31 for the analysis of subclusters in single-cell data sets, which has previously been considered for application to single-cell transcriptomics data50. When correcting for cell cycle, we used the scLVM residual expression data sets (see Choosing the rank of the cell-cycle factor) as input, otherwise we used the preprocessed log expression values.
Choosing the rank of the cell-cycle factor. As described above, scree plots generated for both the T-cell and the mESC data suggested that the largest proportion of variance was explained by the first principal component (Supplementary Fig. 22). Consequently, we used a K = 1 rank covariance matrix to fit the cell-cycle factor in most experiments. When omitting the filtering of cells (quality control, Supplementary Notes), a second component (K = 2) was necessary to fully capture the variation in the data. This second component likely captures intra cell-phase differences in cell size (see also Supplementary Figs. 19–20 and Supplementary Notes).
Alternative approaches to fit the TH2 factor. In order to assess the robustness of the conditional fitting for the TH2 factor as described above (Applying scLVM to multiple annotated gene sets), we compared the results with a conceptually simpler 'iterative approach', where we first regressed out the cell-cycle effects as described above (Gene-gene correlation analysis), before fitting the state of the differentiation factor on cell cycle–corrected expression values.
Reassuringly, the TH2 differentiation factor recovered by either of the approaches was strikingly correlated (Pearson r2 = 0.82, Supplementary Fig. 15c) and was consistent with the subclusters of cells identified by the unsupervised PCA approach (Fig. 3e). In the variance decomposition, the factor determined by the iterative approach yielded a smaller proportion of variance attributable to the TH2 differentiation factor (2.6% versus 5.3%), which can be attributed to the assumption of a common parameter for all genes in the conditional approach (the iterative approach allows a gene-specific contribution of the cell-cycle factor). Critically, the set of genes identified in the interaction component and the GO analysis for the set of genes with a strong interaction effect yielded consistent results (Supplementary Tables 8 and 9).
Accession codes.
mESC data have been deposited at ArrayExpress: E-MTAB-2805. RNA-seq data from the TH2 cells have previously been described20,25 and are available under at ArrayExpress: E-MTAB-2512. Cell cycle–corrected and uncorrected expression values for the T-cell data as well as the mESC data are provided as Supplementary Data 1 and 2. An open source software implementation of scLVM is freely available on GitHub: https://github.com/PMBio/scLVM.
Accession codes
References
Levsky, J.M., Shenoy, S.M., Pezo, R.C. & Singer, R.H. Single-cell gene expression profiling. Science 297, 836–840 (2002).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Raj, A., van den Bogaard, P., Rifkin, S.A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008).
Liu, J., Hansen, C. & Quake, S.R. Solving the “world-to-chip” interface problem with a microfluidic matrix. Anal. Chem. 75, 4718–4723 (2003).
Citri, A., Pang, Z.P., Sudhof, T.C., Wernig, M. & Malenka, R.C. Comprehensive qPCR profiling of gene expression in single neuronal cells. Nat. Protoc. 7, 118–127 (2012).
Wheeler, A.R. et al. Microfluidic device for single-cell analysis. Anal. Chem. 75, 3581–3586 (2003).
Marcus, J.S., Anderson, W.F. & Quake, S.R. Microfluidic single-cell mRNA isolation and analysis. Anal. Chem. 78, 3084–3089 (2006).
Guo, G. et al. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell 18, 675–685 (2010).
Burton, A. et al. Single-cell profiling of epigenetic modifiers identifies PRDM14 as an inducer of cell fate in the mammalian embryo. Cell Reports 5, 687–701 (2013).
Luo, L. et al. Gene expression profiles of laser-captured adjacent neuronal subtypes. Nat. Med. 5, 117–122 (1999).
Chiang, M.K. & Melton, D.A. Single-cell transcript analysis of pancreas development. Dev. Cell 4, 383–393 (2003).
Tang, F. et al. RNA-seq analysis to capture the transcriptome landscape of a single cell. Nat. Protoc. 5, 516–535 (2010).
Islam, S. et al. Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq. Genome Res. 21, 1160–1167 (2011).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Yan, L. et al. Single-cell RNA-seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013).
Tang, F. et al. Tracing the derivation of embryonic stem cells from the inner cell mass by single-cell RNA-seq analysis. Cell Stem Cell 6, 468–478 (2010).
Shalek, A.K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Pollen, A.A. et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat. Biotechnol. 32, 1053–1058 (2014).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Leek, J.T. & Storey, J.D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 3, e161 (2007).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, e1000770 (2010).
Li, S. et al. Detecting and correcting systematic variation in large-scale RNA sequencing data. Nat. Biotechnol. 32, 888–895 (2014).
Xue, Z. et al. Genetic programs in human and mouse early embryos revealed by single-cell RNA sequencing. Nature 500, 593–597 (2013).
Mahata, B. et al. Single-cell RNA sequencing reveals T helper cells synthesizing steroids de novo to contribute to immune homeostasis. Cell Reports 7, 1130–1142 (2014).
Newman, J.R. et al. Single-cell proteomic analysis of S. cerevisiae reveals the architecture of biological noise. Nature 441, 840–846 (2006).
Gold, D., Mallick, B. & Coombes, K. Real-time gene expression: statistical challenges in design and inference. J. Comput. Biol. 15, 611–623 (2008).
Singh, A.M. et al. Cell-cycle control of developmentally regulated transcription factors accounts for heterogeneity in human pluripotent cells. Stem Cell Reports 1, 532–544 (2013).
Lippert, C. et al. FaST linear mixed models for genome-wide association studies. Nat. Methods 8, 833–835 (2011).
Fusi, N., Stegle, O. & Lawrence, N.D. Joint modelling of confounding factors and prominent genetic regulators provides increased accuracy in genetical genomics studies. PLoS Comput. Biol. 8, e1002330 (2012).
Lawrence, N.D. Gaussian process latent variable models for visualisation of high dimensional data. Adv. Neural Inf. Process. Syst. 16, 329–336 (2004).
Sasagawa, Y. et al. Quartz-Seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity. Genome Biol. 14, R31 (2013).
Grün, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014).
Fox, C.J., Hammerman, P.S. & Thompson, C.B. Fuel feeds function: energy metabolism and the T-cell response. Nat. Rev. Immunol. 5, 844–852 (2005).
Nelms, K., Keegan, A.D., Zamorano, J., Ryan, J.J. & Paul, W.E. The IL-4 receptor: signaling mechanisms and biologic functions. Annu. Rev. Immunol. 17, 701–738 (1999).
Zhu, J., Yamane, H., Cote-Sierra, J., Guo, L. & Paul, W.E. GATA-3 promotes TH2 responses through three different mechanisms: induction of TH2 cytokine production, selective growth of TH2 cells and inhibition of Th1 cell-specific factors. Cell Res. 16, 3–10 (2006).
Stritesky, G.L. et al. The transcription factor STAT3 is required for T helper 2 cell development. Immunity 34, 39–49 (2011).
Zhou, M. et al. Kruppel-like transcription factor 13 regulates T lymphocyte survival in vivo. J. Immunol. 178, 5496–5504 (2007).
Betz, B.C. et al. Batf coordinates multiple aspects of B and T cell function required for normal antibody responses. J. Exp. Med. 207, 933–942 (2010).
Sahoo, A. et al. Stat6 and c-Jun mediate TH2 cell-specific IL--24 gene expression. J. Immunol. 186, 4098–4109 (2011).
Jensen, L.J. et al. STRING 8–a global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 37, D412–D416 (2009).
Chang, C.H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Garcia-Sanz, J.A., Mikulits, W., Livingstone, A., Lefkovits, I. & Mullner, E.W. Translational control: a general mechanism for gene regulation during T cell activation. FASEB J. 12, 299–306 (1998).
Bird, J.J. et al. Helper T cell differentiation is controlled by the cell cycle. Immunity 9, 229–237 (1998).
Wilson, C.B., Makar, K.W. & Perez-Melgosa, M. Epigenetic regulation of T cell fate and function. J. Infect. Dis. 185 (suppl. 1), S37–S45 (2002).
Stegle, O., Teichmann, S.A. & Marioni, J.C. Computational and analytical challenges in single-cell transcriptomics. Nat. Rev. Genet. (in the press).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Gagnon-Bartsch, J.A. & Speed, T.P. Using control genes to correct for unwanted variation in microarray data. Biostatistics 13, 539–552 (2012).
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824 (2012).
Buettner, F. & Theis, F.J. A novel approach for resolving differences in single-cell gene expression patterns from zygote to blastocyst. Bioinformatics 28, i626–i632 (2012).
Acknowledgements
We thank S. Anders and A. Baud for helpful discussions. We also thank the Sanger-EBI Single Cell Centre for technical support. We acknowledge support of the European Research Council (Starting grant no. 260507 thSWITCH to S.A.T., Starting Grant LatentCauses to F.J.T., Marie Curie FP7 fellowship 253524 to O.S.), the Sanger-EBI Single Cell Centre (K.N.N. & A.S.) and the European Molecular Biology Organization (short-term fellowship to F.B.).
Author information
Authors and Affiliations
Contributions
F.B. developed the method, performed the analysis and wrote the paper. K.N.N. performed the mESC experiments and contributed to the analysis. F.P.C. and A.S. contributed to method development and analysis. V.P., S.A.T. and F.J.T. helped interpret the biological results. S.A.T. and V.P. designed the mouse TH2 differentiation experiment. J.C.M. and O.S. designed and supervised this study, contributed to the method development and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–22, Supplementary Tables 3, 6, 8–9 and Supplementary Notes (PDF 31436 kb)
Supplementary Table 1
List of genes annotated from cell cycle, either from GO or from CycleBase. (XLSX 61 kb)
Supplementary Table 2
List of contribution of all variance components for the T-cell data. (XLSX 484 kb)
Supplementary Table 4
List of significantly differentially expressed genes between identified cell sub clusters. (XLSX 47 kb)
Supplementary Table 5
Manually curated list of 122 Th2 signature genes. (XLSX 33 kb)
Supplementary Table 7
List of genes with more than 5% of the variance explained by interaction between cell cycle and differentiation. (XLSX 34 kb)
Supplementary Data 1
Corrected and uncorrected expression values for T-cell data. (XLSX 9175 kb)
Supplementary Data 2
Corrected and uncorrected expression values for the newly generated mouse ESC data. (XLSX 37981 kb)
Rights and permissions
About this article
Cite this article
Buettner, F., Natarajan, K., Casale, F. et al. Computational analysis of cell-to-cell heterogeneity in single-cell RNA-sequencing data reveals hidden subpopulations of cells. Nat Biotechnol 33, 155–160 (2015). https://doi.org/10.1038/nbt.3102
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3102
This article is cited by
-
Characteristics of A-type voltage-gated K+ currents expressed on sour-sensing type III taste receptor cells in mice
Cell and Tissue Research (2024)
-
iDESC: identifying differential expression in single-cell RNA sequencing data with multiple subjects
BMC Bioinformatics (2023)
-
On triangle inequalities of correlation-based distances for gene expression profiles
BMC Bioinformatics (2023)
-
Variability of fluorescence intensity distribution measured by flow cytometry is influenced by cell size and cell cycle progression
Scientific Reports (2023)
-
A spectral method for assessing and combining multiple data visualizations
Nature Communications (2023)