Abstract
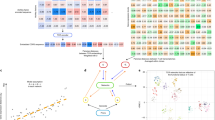
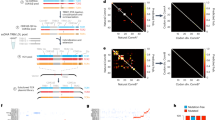
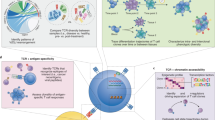
Although each T lymphocyte expresses a T-cell receptor (TCR) that recognizes cognate antigen and controls T-cell activation, different T cells bearing the same TCR can be functionally distinct. Each TCR is a heterodimer, and both α- and β-chains contribute to determining TCR antigen specificity. Here we present a methodology enabling integration of information about TCR specificity with information about T cell function. This method involves sequencing of TCRα and TCRβ genes, and amplifying functional genes characteristic of different T cell subsets, in single T cells. Because this approach retains information about individual TCRα-TCRβ pairs, TCRs of interest can be expressed and used in functional studies, for antigen discovery, or in therapeutic applications. We apply this approach to study the clonal ancestry and differentiation of T lymphocytes infiltrating a human colorectal carcinoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
14 January 2015
In the version of this article initially published, the concentration of the V-region primers in the Online Methods section was given as 0.6 μM. The correct concentration is 0.06 μM. The error has been corrected in the HTML and PDF versions of the article.
References
Wills, Q.F. et al. Single-cell gene expression analysis reveals genetic associations masked in whole-tissue experiments. Nat. Biotechnol. 31, 748–752 (2013).
Newell, E.W., Sigal, N., Bendall, S.C., Nolan, G.P. & Davis, M.M. Cytometry by time-of-flight shows combinatorial cytokine expression and virus-specific cell niches within a continuum of CD8+ T cell phenotypes. Immunity 36, 142–152 (2012).
Shapiro, E., Biezuner, T. & Linnarsson, S. Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat. Rev. Genet. 14, 618–630 (2013).
Bendall, S.C., Nolan, G.P., Roederer, M. & Chattopadhyay, P.K. A deep profiler's guide to cytometry. Trends Immunol. 33, 323–332 (2012).
Spurgeon, S.L., Jones, R.C. & Ramakrishnan, R. High throughput gene expression measurement with real time PCR in a microfluidic dynamic array. PLoS ONE 3, e1662 (2008).
Wu, A.R. et al. Quantitative assessment of single-cell RNA-sequencing methods. Nat. Methods 11, 41–46 (2014).
Newell, E.W. & Davis, M.M. Beyond model antigens: high-dimensional methods for the analysis of antigen-specific T cells. Nat. Biotechnol. 32, 149–157 (2014).
Krogsgaard, M. & Davis, M.M. How T cells 'see' antigen. Nat. Immunol. 6, 239–245 (2005).
Murphy, K., Travers, P., Walport, M. & Janeway, C. Janeway's Immunobiology 8th Edn. (Garland Science, 2012).
Newell, E.W. et al. Combinatorial tetramer staining and mass cytometry analysis facilitate T-cell epitope mapping and characterization. Nat. Biotechnol. 31, 623–629 (2013).
Birnbaum, M.E. et al. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157, 1073–1087 (2014).
Hinrichs, C.S. & Restifo, N.P. Reassessing target antigens for adoptive T-cell therapy. Nat. Biotechnol. 31, 999–1008 (2013).
Han, A. et al. Dietary gluten triggers concomitant activation of CD4+ and CD8+ alphabeta T cells and gammadelta T cells in celiac disease. Proc. Natl. Acad. Sci. USA 110, 13073–13078 (2013).
Kim, S.M. et al. Analysis of the paired TCR alpha- and beta-chains of single human T cells. PLoS ONE 7, e37338 (2012).
Dash, P. et al. Paired analysis of TCRalpha and TCRbeta chains at the single-cell level in mice. J. Clin. Invest. 121, 288–295 (2011).
Gascoigne, N.R. & Alam, S.M. Allelic exclusion of the T cell receptor alpha-chain: developmental regulation of a post-translational event. Semin. Immunol. 11, 337–347 (1999).
Malissen, M. et al. Regulation of TCR alpha and beta gene allelic exclusion during T-cell development. Immunol. Today 13, 315–322 (1992).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Glanville, J. et al. Precise determination of the diversity of a combinatorial antibody library gives insight into the human immunoglobulin repertoire. Proc. Natl. Acad. Sci. USA 106, 20216–20221 (2009).
De Rosa, S.C., Herzenberg, L.A., Herzenberg, L.A. & Roederer, M. 11-color, 13-parameter flow cytometry: identification of human naive T cells by phenotype, function, and T-cell receptor diversity. Nat. Med. 7, 245–248 (2001).
Yanagi, Y., Chan, A., Chin, B., Minden, M. & Mak, T.W. Analysis of cDNA clones specific for human T cells and the alpha and beta chains of the T-cell receptor heterodimer from a human T-cell line. Proc. Natl. Acad. Sci. USA 82, 3430–3434 (1985).
Nakamura, K. et al. Sequence-specific error profile of Illumina sequencers. Nucleic Acids Res. 39, e90 (2011).
Law, J.P. et al. The importance of Foxp3 antibody and fixation/permeabilization buffer combinations in identifying CD4+CD25+Foxp3+ regulatory T cells. Cytometry A 75, 1040–1050 (2009).
Vahedi, G., Kanno, Y., Sartorelli, V. & O'Shea, J.J. Transcription factors and CD4 T cells seeking identity: masters, minions, setters and spikers. Immunology 139, 294–298 (2013).
Oestreich, K.J. & Weinmann, A.S. Master regulators or lineage-specifying? Changing views on CD4+ T cell transcription factors. Nat. Rev. Immunol. 12, 799–804 (2012).
Wilson, C.B., Rowell, E. & Sekimata, M. Epigenetic control of T-helper-cell differentiation. Nat. Rev. Immunol. 9, 91–105 (2009).
Collins, A., Littman, D.R. & Taniuchi, I. RUNX proteins in transcription factor networks that regulate T-cell lineage choice. Nat. Rev. Immunol. 9, 106–115 (2009).
Assenmacher, M., Lohning, M. & Radbruch, A. Detection and isolation of cytokine secreting cells using the cytometric cytokine secretion assay. Curr. Protoc. Immunol. 46, 6.27 (2002).
Anderson, P. Post-transcriptional control of cytokine production. Nat. Immunol. 9, 353–359 (2008).
Fontenot, J.D., Gavin, M.A. & Rudensky, A.Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Ribas, A. Tumor immunotherapy directed at PD-1. N. Engl. J. Med. 366, 2517–2519 (2012).
Sliwkowski, M.X. & Mellman, I. Antibody therapeutics in cancer. Science 341, 1192–1198 (2013).
Pagès, F. et al. Effector memory T cells, early metastasis, and survival in colorectal cancer. N. Engl. J. Med. 353, 2654–2666 (2005).
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Gerlinger, M. et al. Ultra-deep T-cell receptor sequencing reveals the complexity and intratumour heterogeneity of T-cell clones in renal cell carcinomas. J. Pathol. 231, 424–432 (2013).
Sherwood, A.M. et al. Tumor-infiltrating lymphocytes in colorectal tumors display a diversity of T cell receptor sequences that differ from the T cells in adjacent mucosal tissue. Cancer immunol. immunother. 62, 1453–1461 (2013).
Sasada, T. & Suekane, S. Variation of tumor-infiltrating lymphocytes in human cancers: controversy on clinical significance. Immunotherapy 3, 1235–1251 (2011).
deLeeuw, R.J., Kost, S.E., Kakal, J.A. & Nelson, B.H. The prognostic value of FoxP3+ tumor-infiltrating lymphocytes in cancer: a critical review of the literature. Clin. Cancer Res. 18, 3022–3029 (2012).
Scurr, M., Gallimore, A. & Godkin, A. T cell subsets and colorectal cancer: discerning the good from the bad. Cell. Immunol. 279, 21–24 (2012).
Tosolini, M. et al. Clinical impact of different classes of infiltrating T cytotoxic and helper cells (Th1, th2, treg, th17) in patients with colorectal cancer. Cancer Res. 71, 1263–1271 (2011).
Ladoire, S., Martin, F. & Ghiringhelli, F. Prognostic role of FOXP3+ regulatory T cells infiltrating human carcinomas: the paradox of colorectal cancer. Cancer immunol. immunother. 60, 909–918 (2011).
Blatner, N.R. et al. Expression of RORgammat marks a pathogenic regulatory T cell subset in human colon cancer. Sci. Transl. Med. 4, 164ra159 (2012).
Gounaris, E. et al. T-regulatory cells shift from a protective anti-inflammatory to a cancer-promoting proinflammatory phenotype in polyposis. Cancer Res. 69, 5490–5497 (2009).
Miyara, M. et al. Functional delineation and differentiation dynamics of human CD4+ T cells expressing the FoxP3 transcription factor. Immunity 30, 899–911 (2009).
Zhou, L., Chong, M.M. & Littman, D.R. Plasticity of CD4+ T cell lineage differentiation. Immunity 30, 646–655 (2009).
Acknowledgements
We thank members of the Davis laboratory and the Y.-H. Chien laboratory for helpful discussions. We thank E. Newell for critical reading of the manuscript and helpful suggestions. We thank C. Bolen for assistance with data analysis. We thank X. Ji for deep sequencing. Tissue was obtained through the Stanford University Tissue Bank. The Stanford Shared FACS Facility provided use of equipment and the Stanford Functional Genomics Facility provided deep sequencing services. A.H. is supported by a grant from the Simons Foundation. L.H. is supported by a fellowship from the German Research Foundation (D.F.G.). M.M.D. is funded by the US National Institutes of Health and is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.H. conceived the project and experiments, designed methodology and primers, performed experiments, analyzed data, assisted in the optimization of the software pipeline, generated figures and wrote the manuscript. J.G. designed the software, analyzed data and generated figures. L.H. designed and performed experiments, analyzed data and generated figures. M.M.D. conceived the project and experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1, 3, 5, 7 and Supplementary Note (PDF 22832 kb)
Supplementary Table 2
Penotyping primers for first two PCR reactions (XLSX 57 kb)
Supplementary Table 4
TCR sequences from the TCR validation panel (XLSX 62 kb)
Supplementary Table 6
Reads counts per well of each phenotyping parameter illustrated in Fig. 2. (XLSX 119 kb)
Supplementary Table 8
Reads counts per well of each phenotyping parameter illustrated in Fig. 2. (XLSX 108 kb)
Supplementary Table 9
Paired TCR alpha/beta sequences for 309 CD4+ T cells from adjacent colon for which a TCR beta chain was obtained. (XLSX 71 kb)
Supplementary Table 10
Reads counts per well of each tumor CD4+ T cell analyzed. (XLSX 106 kb)
Supplementary Table 11
Reads counts per well of each adjacent colon CD4+ T cell analyzed. (XLSX 89 kb)
Rights and permissions
About this article
Cite this article
Han, A., Glanville, J., Hansmann, L. et al. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat Biotechnol 32, 684–692 (2014). https://doi.org/10.1038/nbt.2938
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2938
This article is cited by
-
An open protocol for modeling T Cell Clonotype repertoires using TCRβ CDR3 sequences
BMC Genomics (2023)
-
Bioengineering translational models of lymphoid tissues
Nature Reviews Bioengineering (2023)
-
Immune phenotypes and checkpoint molecule expression of clonally expanded lymph node-infiltrating T cells in classical Hodgkin lymphoma
Cancer Immunology, Immunotherapy (2023)
-
T cell receptor repertoires associated with control and disease progression following Mycobacterium tuberculosis infection
Nature Medicine (2023)
-
T-cell repertoire diversity: friend or foe for protective antitumor response?
Journal of Experimental & Clinical Cancer Research (2022)