Abstract
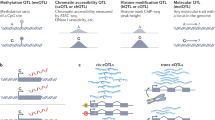
Association studies provide genome-wide information about the genetic basis of complex disease, but medical research has focused primarily on protein-coding variants, owing to the difficulty of interpreting noncoding mutations. This picture has changed with advances in the systematic annotation of functional noncoding elements. Evolutionary conservation, functional genomics, chromatin state, sequence motifs and molecular quantitative trait loci all provide complementary information about the function of noncoding sequences. These functional maps can help with prioritizing variants on risk haplotypes, filtering mutations encountered in the clinic and performing systems-level analyses to reveal processes underlying disease associations. Advances in predictive modeling can enable data-set integration to reveal pathways shared across loci and alleles, and richer regulatory models can guide the search for epistatic interactions. Lastly, new massively parallel reporter experiments can systematically validate regulatory predictions. Ultimately, advances in regulatory and systems genomics can help unleash the value of whole-genome sequencing for personalized genomic risk assessment, diagnosis and treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Collins, F. Has the revolution arrived? Nature 464, 674–675 (2010).
Lander, E.S. Initial impact of the sequencing of the human genome. Nature 470, 187–197 (2011).
Botstein, D. & Risch, N. Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease. Nat. Genet. 33, 228–237 (2003).
Hamosh, A. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 33, D514–D517 (2005).
Botstein, D., White, R.L., Skolnick, M. & Davis, R.W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32, 314–331 (1980).
Lander, E.S. & Botstein, D. Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121, 185–199 (1989).
Watson, J.D. The Human Genome Project: past, present, and future. Science 248, 44–49 (1990).
Lander, E. & Kruglyak, L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat. Genet. 11, 241–247 (1995).
International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003).
McCarthy, M.I. et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat. Rev. Genet. 9, 356–369 (2008).
Hindorff, L.A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009). The NHGRI GWAS Catalog reported here laid the groundwork for systematic intersection of functional annotations with disease-associated regions, and highlighted the preponderance of noncoding disease associations.
Manolio, T.A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009). This paper reports the deliberations of the NHGRI's expert working group on the sources of unexplained heritability, and their suggestions for future research strategies.
Cirulli, E.T. & Goldstein, D.B. Uncovering the roles of rare variants in common disease through whole-genome sequencing. Nat. Rev. Genet. 11, 415–425 (2010).
Fisher, R. The correlation between relatives on the supposition of Mendelian inheritance. Trans. R. Soc. Edinb. 52, 399–433 (1918).
Visscher, P.M. McEvoy, B. & Yang, J. From Galton to GWAS: quantitative genetics of human height. Genet. Res. 92, 371–379 (2010).
MacArthur, D.G. et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science 335, 823–828 (2012).
Nelson, M.R. et al. An abundance of rare functional variants in 202 drug target genes sequenced in 14,002 people. Science 337, 100–104 (2012).
Park, P.J. ChIP–seq: advantages and challenges of a maturing technology. Nat. Rev. Genet. 10, 669–680 (2009).
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 33, 5868–5877 (2005).
Boyle, A.P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012). The ENCODE consortium scale-up datasets represent the most comprehensive annotation of the noncoding genome at the time of this review.
Bernstein, B.E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Adams, D. et al. BLUEPRINT to decode the epigenetic signature written in blood. Nat. Biotechnol. 30, 224–226 (2012).
Bussemaker, H.J., Foat, B.C. & Ward, L.D. Predictive modeling of genome-wide mRNA expression: from modules to molecules. Annu. Rev. Biophys. Biomol. Struct. 36, 329–347 (2007).
Tompa, M. et al. Assessing computational tools for the discovery of transcription factor binding sites. Nat. Biotechnol. 23, 137–144 (2005).
Barash, Y. et al. Deciphering the splicing code. Nature 465, 53–59 (2010).
Wang, Z. & Burge, C.B. Splicing regulation: from a parts list of regulatory elements to an integrated splicing code. RNA 14, 802–813 (2008).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Moses, A.M., Chiang, D., Pollard, D., Iyer, V. & Eisen, M. MONKEY: identifying conserved transcription-factor binding sites in multiple alignments using a binding site-specific evolutionary model. Genome Biol. 5, R98 (2004).
Hesselberth, J.R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nat. Methods 6, 283–289 (2009).
Henikoff, J.G., Belsky, J.A., Krassovsky, K., MacAlpine, D.M. & Henikoff, S. Epigenome characterization at single base-pair resolution. Proc. Natl. Acad. Sci. USA 108, 18318–18323 (2011).
Rhee, H.S. & Pugh, B.F. Comprehensive genome-wide protein–DNA interactions detected at single-nucleotide resolution. Cell 147, 1408–1419 (2011).
Beer, M.A. & Tavazoie, S. Predicting gene expression from sequence. Cell 117, 185–198 (2004).
Roy, S. et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Gerstein, M.B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Davidson, E.H. et al. A genomic regulatory network for development. Science 295, 1669–1678 (2002).
Patwardhan, R.P. et al. Massively parallel functional dissection of mammalian enhancers in vivo. Nat. Biotechnol. 30, 265–270 (2012).
Sharon, E. et al. Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters. Nat. Biotechnol. 30, 521–530 (2012).
Melnikov, A. et al. Systematic dissection and optimization of inducible enhancers in human cells using a massively parallel reporter assay. Nat. Biotechnol. 30, 271–277 (2012).
Davydov, E.V. et al. Identifying a high fraction of the human to be under selective constraint using GERP++. PLOS Comput. Biol. 6, e1001025 (2010).
Lindblad-Toh, K. et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–482 (2011). Conserved elements were shown to be enriched among disease-associated variants, motivating the use of conservation to guide candidate causal SNP selection.
Kellis, M., Patterson, N., Endrizzi, M., Birren, B. & Lander, E.S. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423, 241–254 (2003).
Stark, A. et al. Discovery of functional elements in 12 Drosophila genomes using evolutionary signatures. Nature 450, 219–232 (2007).
Papatsenko, D., Kislyuk, A., Levine, M. & Dubchak, I. Conservation patterns in different functional sequence categories of divergent Drosophila species. Genomics 88, 431–442 (2006).
Dermitzakis, E.T. & Clark, A.G. Evolution of transcription factor binding sites in mammalian gene regulatory regions: conservation and turnover. Mol. Biol. Evol. 19, 1114–1121 (2002).
Meader, S., Ponting, C.P. & Lunter, G. Massive turnover of functional sequence in human and other mammalian genomes. Genome Res. 20, 1335–1343 (2010).
Schmidt, D. et al. Five-vertebrate ChIP-seq reveals the evolutionary dynamics of transcription factor binding. Science 328, 1036–1040 (2010).
Brawand, D. et al. The evolution of gene expression levels in mammalian organs. Nature 478, 343–348 (2011).
Ng, P.C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Yue, P., Melamud, E. & Moult, J. SNPs3D: candidate gene and SNP selection for association studies. BMC Bioinformatics 7, 166 (2006).
Ramensky, V., Bork, P. & Sunyaev, S. Human non-synonymous SNPs: server and survey. Nucleic Acids Res. 30, 3894–3900 (2002).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Baker, M. Functional genomics: the changes that count. Nature 482, 257–262 (2012).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Boyle, A.P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP effect predictor. Bioinformatics 26, 2069–2070 (2010).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Yandell, M. et al. A probabilistic disease-gene finder for personal genomes. Genome Res. 10.1101/gr.123158.111 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Wang, K., Li, M. & Hakonarson, H. Analysing biological pathways in genome-wide association studies. Nat. Rev. Genet. 11, 843–854 (2010).
McKinney, B.A. & Pajewski, N.M. Six degrees of epistasis: statistical network models for GWAS. Front. Genet 2, 109 (2012).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011). This was the first demonstration that cross-tissue enhancer maps can link noncoding variants from GWAS to relevant cell types and candidate regulatory mechanisms.
Maurano, M.T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Nica, A.C. et al. Candidate causal regulatory effects by integration of expression QTLs with complex trait genetic associations. PLoS Genet. 6, e1000895 (2010). This study uses eQTLs to investigate the tissue specificity of gene regulatory mechanisms, and suggests that assaying many tissues will be critical to developing a cis-regulatory map of the human genome.
Nicolae, D.L. et al. Trait-associated SNPs are more likely to be eQTLs: annotation to enhance discovery from GWAS. PLoS Genet. 6, e1000888 (2010).
Cantor, R.M., Lange, K. & Sinsheimer, J.S. Prioritizing GWAS results: a review of statistical methods and recommendations for their application. Am. J. Hum. Genet. 86, 6–22 (2010). The authors present an extensive review of how biological annotations are being used in association studies and to interpret their results. They show how knowledge of molecular pathways can be used to enhance discovery, test for epistasis and aggregate results.
Knight, J., Barnes, M.R., Breen, G. & Weale, M.E. Using functional annotation for the empirical determination of Bayes factors for genome-wide association study analysis. PLoS ONE 6, e14808 (2011).
Lewinger, J.P., Conti, D.V., Baurley, J.W., Triche, T.J. & Thomas, D.C. Hierarchical Bayes prioritization of marker associations from a genome-wide association scan for further investigation. Genet. Epidemiol. 31, 871–882 (2007).
Chen, G.K. & Witte, J.S. Enriching the analysis of genomewide association studies with hierarchical modeling. Am. J. Hum. Genet. 81, 397–404 (2007).
Lee, I., Blom, U.M., Wang, P.I., Shim, J.E. & Marcotte, E.M. Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21, 1109–1121 (2011).
Dering, C., Hemmelmann, C., Pugh, E. & Ziegler, A. Statistical analysis of rare sequence variants: an overview of collapsing methods. Genet. Epidemiol. 35, S12–S17 (2011).
Bansal, V., Libiger, O., Torkamani, A. & Schork, N.J. Statistical analysis strategies for association studies involving rare variants. Nat. Rev. Genet. 11, 773–785 (2010).
Pai, A.A., Bell, J.T., Marioni, J.C., Pritchard, J.K. & Gilad, Y. A genome-wide study of DNA methylation patterns and gene expression levels in multiple human and chimpanzee tissues. PLoS Genet. 7, e1001316 (2011).
Degner, J.F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010).
Majewski, J. & Pastinen, T. The study of eQTL variations by RNA-seq: from SNPs to phenotypes. Trends Genet. 27, 72–79 (2011).
Pickrell, J.K. et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464, 768–772 (2010).
Lappalainen, T., Montgomery, S.B., Nica, A.C. & Dermitzakis, E.T. Epistatic selection between coding and regulatory variation in human evolution and disease. Am. J. Hum. Genet. 89, 459–463 (2011).
Kerkel, K. et al. Genomic surveys by methylation-sensitive SNP analysis identify sequence-dependent allele-specific DNA methylation. Nat. Genet. 40, 904–908 (2008).
Prendergast, J.G., Tong, P., Hay, D.C., Farrington, S.M. & Semple, C.A. A genome-wide screen in human embryonic stem cells reveals novel sites of allele-specific histone modification associated with known disease loci. Epigenetics Chromatin 5, 6 (2012).
McDaniell, R. et al. Heritable individual-specific and allele-specific chromatin signatures in humans. Science 328, 235–239 (2010). In this study, the authors demonstrated that both genomic protein binding and DNase I hypersensitivity were heritable, and therefore under genetic control.
Maynard, N.D., Chen, J., Stuart, R.K., Fan, J.-B. & Ren, B. Genome-wide mapping of allele-specific protein-DNA interactions in human cells. Nat. Methods 5, 307–309 (2008).
Ge, B. et al. Global patterns of cis variation in human cells revealed by high-density allelic expression analysis. Nat. Genet. 41, 1216–1222 (2009).
Ng, P.C., Murray, S.S., Levy, S. & Venter, J.C. An agenda for personalized medicine. Nature 461, 724–726 (2009).
Patterson, N. et al. Methods for high-density admixture mapping of disease genes. Am. J. Hum. Genet. 74, 979–1000 (2004).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Coop, G. et al. The role of geography in human adaptation. PLoS Genet. 5, e1000500 (2009).
Hernandez, R.D. et al. Classic selective sweeps were rare in recent human evolution. Science 331, 920–924 (2011).
Sabeti, P.C. et al. Positive natural selection in the human lineage. Science 312, 1614–1620 (2006).
Grossman, S.R. et al. A composite of multiple signals distinguishes causal variants in regions of positive selection. Science 327, 883–886 (2010).
Ott, J., Kamatani, Y. & Lathrop, M. Family-based designs for genome-wide association studies. Nat. Rev. Genet. 12, 465–474 (2011).
Minichiello, M.J. & Durbin, R. Mapping trait loci by use of inferred ancestral recombination graphs. Am. J. Hum. Genet. 79, 910–922 (2006).
Wu, Y. Association mapping of complex diseases with ancestral recombination graphs: models and efficient algorithms. J. Comput. Biol. 15, 667–684 (2008).
Asthana, S. et al. Widely distributed noncoding purifying selection in the human genome. Proc. Natl. Acad. Sci. USA 104, 12410–12415 (2007).
Ward, L.D. & Kellis, M. Evidence of abundant purifying selection in humans for recently acquired regulatory functions. Science 10.1126/science.1225057 (2012).
Hill, W.G., Goddard, M.E. & Visscher, P.M. Data and theory point to mainly additive genetic variance for complex traits. PLoS Genet. 4, e1000008 (2008).
Shao, H. et al. Genetic architecture of complex traits: large phenotypic effects and pervasive epistasis. Proc. Natl. Acad. Sci. USA 105, 19910–19914 (2008).
Zuk, O., Hechter, E., Sunyaev, S.R. & Lander, E.S. The mystery of missing heritability: Genetic interactions create phantom heritability. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1119675109 (2012).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010).
Cordell, H.J. Detecting gene–gene interactions that underlie human diseases. Nat. Rev. Genet. 10, 392–404 (2009).
Musani, S.K. et al. Detection of gene x gene interactions in genome-wide association studies of human population data. Hum. Hered. 63, 67–84 (2007).
Lou, X.-Y. et al. A combinatorial approach to detecting gene–gene and gene–environment interactions in family studies. Am. J. Hum. Genet. 83, 457–467 (2008).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Emily, M., Mailund, T., Hein, J., Schauser, L. & Schierup, M.H. Using biological networks to search for interacting loci in genome-wide association studies. Eur. J. Hum. Genet. 17, 1231–1240 (2009).
Mechanic, L.E., Luke, B.T., Goodman, J.E., Chanock, S.J. & Harris, C.C. Polymorphism Interaction Analysis (PIA): a method for investigating complex gene-gene interactions. BMC Bioinformatics 9, 146 (2008).
Pattin, K.A. & Moore, J.H. Exploiting the proteome to improve the genome-wide genetic analysis of epistasis in common human diseases. Hum. Genet. 124, 19–29 (2008).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Fullwood, M.J. et al. An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 462, 58–64 (2009).
Cheng, C. et al. Construction and analysis of an integrated regulatory network derived from high-throughput sequencing data. PLOS Comput. Biol. 7, e1002190 (2011).
Zhu, J. et al. Integrating large-scale functional genomic data to dissect the complexity of yeast regulatory networks. Nat. Genet. 40, 854–861 (2008).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Burke, M.K. et al. Genome-wide analysis of a long-term evolution experiment with Drosophila. Nature 467, 587–590 (2010).
Gresham, D. et al. The repertoire and dynamics of evolutionary adaptations to controlled nutrient-limited environments in yeast. PLoS Genet. 4, e1000303 (2008).
Perlstein, E.O., Ruderfer, D.M., Roberts, D.C., Schreiber, S.L. & Kruglyak, L. Genetic basis of individual differences in the response to small-molecule drugs in yeast. Nat. Genet. 39, 496–502 (2007).
Quackenbush, J. Microarray analysis and tumor classification. N. Engl. J. Med. 354, 2463–2472 (2006).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Rakyan, V.K., Down, T.A., Balding, D.J. & Beck, S. Epigenome-wide association studies for common human diseases. Nat. Rev. Genet. 12, 529–541 (2011). The authors review the challenges and promise of EWAS, and how their results can be used in conjunction with GWAS.
Petronis, A. Epigenetics as a unifying principle in the aetiology of complex traits and diseases. Nature 465, 721–727 (2010).
Chen, L.S., Emmert-Streib, F. & Storey, J.D. Harnessing naturally randomized transcription to infer regulatory relationships among genes. Genome Biol. 8, R219 (2007).
Lawlor, D.A., Harbord, R.M., Sterne, J.A.C., Timpson, N. & Davey Smith, G. Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat. Med. 27, 1133–1163 (2008).
Voight, B.F. et al. Plasma HDL cholesterol and risk of myocardial infarction: a mendelian randomisation study. Lancet 380, 572–580.
Chen, R. et al. Personal omics profiling reveals dynamic molecular and medical phenotypes. Cell 148, 1293–1307 (2012).
Anonymous. Asking for more. Nat. Genet. 44, 733 (2012).
Homer, N. et al. Resolving individuals contributing trace amounts of DNA to highly complex mixtures using high-density SNP genotyping microarrays. PLoS Genet. 4, e1000167 (2008).
Salathé, M. et al. Digital epidemiology. PLOS Comput. Biol. 8, e1002616 (2012).
Brownstein, J.S., Sordo, M., Kohane, I.S. & Mandl, K.D. The tell-tale heart: population-based surveillance reveals an association of rofecoxib and celecoxib with myocardial Infarction. PLoS ONE 2, e840 (2007).
Roque, F.S. et al. Using electronic patient records to discover disease correlations and stratify patient cohorts. PLOS Comput. Biol. 7, e1002141 (2011).
Wilke, R.A. et al. The emerging role of electronic medical records in pharmacogenomics. Clin. Pharmacol. Ther. 89, 379–386 (2011).
Nebert, D.W., Zhang, G. & Vesell, E.S. From human genetics and genomics to pharmacogenetics and pharmacogenomics: past lessons, future directions. Drug Metab. Rev. 40, 187–224 (2008). A critical review of current challenges in human genetics and the application of pharmacogenetic discoveries to clinical practice.
Garrod, A. E. & Harris, H. Inborn Errors of Metabolism (Henry Frowde and Hodder & Stoughton, London, 1909).
Woo, S.L., Lidsky, A.S., Güttler, F., Chandra, T. & Robson, K.J. Cloned human phenylalanine hydroxylase gene allows prenatal diagnosis and carrier detection of classical phenylketonuria. Nature 306, 151–155 (1983).
Riordan, J.R. et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science 245, 1066–1073 (1989).
Audrézet, M.P. et al. Genomic rearrangements in the CFTR gene: extensive allelic heterogeneity and diverse mutational mechanisms. Hum. Mutat. 23, 343–357 (2004).
Zschocke, J. Phenylketonuria mutations in Europe. Hum. Mutat. 21, 345–356 (2003).
Amiel, J. et al. Hirschsprung disease, associated syndromes and genetics: a review. J. Med. Genet. 45, 1–14 (2008).
Yang, J. et al. Common SNPs explain a large proportion of heritability for human height. Nat. Genet. 42, 565–569 (2010).
Purcell, S.M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Nica, A.C. et al. The architecture of gene regulatory variation across multiple human tissues: the MuTHER study. PLoS Genet. 7, e1002003 (2011).
King, J.L. & Jukes, T.H. Non-Darwinian evolution. Science 164, 788–798 (1969).
Kimura, M. Evolutionary rate at the molecular level. Nature 217, 624–626 (1968).
Ohno, S. So much 'junk' DNA in our genome. Brookhaven Symp. Biol. 23, 366–370 (1972).
Stratton, M.R., Campbell, P.J. & Futreal, P.A. The cancer genome. Nature 458, 719–724 (2009).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Servin, B. & Stephens, M. Imputation-based analysis of association studies: candidate regions and quantitative traits. PLoS Genet. 3, e114 (2007).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Veyrieras, J.-B. et al. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet. 4, e1000214 (2008).
Shabalin, A.A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Rozowsky, J. et al. AlleleSeq: analysis of allele-specific expression and binding in a network framework. Mol. Syst. Biol. 7, 522 (2011).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 3 (2004).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Korn, J.M. et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 40, 1253–1260 (2008).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Faustino, N.A. & Cooper, T.A. Pre-mRNA splicing and human disease. Genes Dev. 17, 419–437 (2003).
Cáceres, J.F. & Kornblihtt, A.R. Alternative splicing: multiple control mechanisms and involvement in human disease. Trends Genet. 18, 186–193 (2002).
López-Bigas, N., Audit, B., Ouzounis, C., Parra, G. & Guigó, R. Are splicing mutations the most frequent cause of hereditary disease? FEBS Lett. 579, 1900–1903 (2005).
Barbaux, S. et al. Donor splice-site mutations in WT1 are responsible for Frasier syndrome. Nat. Genet. 17, 467–470 (1997).
Lorson, C.L., Hahnen, E., Androphy, E.J. & Wirth, B. A single nucleotide in the SMN gene regulates splicing and is responsible for spinal muscular atrophy. Proc. Natl. Acad. Sci. USA 96, 6307–6311 (1999).
Cazzola, M. & Skoda, R.C. Translational pathophysiology: a novel molecular mechanism of human disease. Blood 95, 3280–3288 (2000).
Bisio, A. et al. Functional analysis of CDKN2A/p16INK4a 5′-UTR variants predisposing to melanoma. Hum. Mol. Genet. 19, 1479–1491 (2010).
Abelson, J.F. et al. Sequence variants in SLITRK1 are associated with Tourette's syndrome. Science 310, 317–320 (2005).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Ponting, C.P., Oliver, P.L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Bonafé, L. et al. Evolutionary comparison provides evidence for pathogenicity of RMRP mutations. PLoS Genet. 1, e47 (2005).
Cooper, T.A., Wan, L. & Dreyfuss, G. RNA and disease. Cell 136, 777–793 (2009).
Knight, J.C. Regulatory polymorphisms underlying complex disease traits. J. Mol. Med. 83, 97–109 (2005).
Martin, M.P. et al. Genetic acceleration of AIDS progression by a promoter variant of CCR5. Science 282, 1907–1911 (1998).
Bream, J.H. et al. CCR5 promoter alleles and specific DNA binding factors. Science 284, 223 (1999).
Bray, N.J. et al. Allelic expression of APOE in human brain: effects of epsilon status and promoter haplotypes. Hum. Mol. Genet. 13, 2885–2892 (2004).
St George-Hyslop, P.H. & Petit, A. Molecular biology and genetics of Alzheimer's disease. C. R. Biol. 328, 119–130 (2005).
Exner, M., Minar, E., Wagner, O. & Schillinger, M. The role of heme oxygenase-1 promoter polymorphisms in human disease. Free Radic. Biol. Med. 37, 1097–1104 (2004).
Kleinjan, D.A. & van Heyningen, V. Long-range control of gene expression: Emerging mechanisms and disruption in disease. Am. J. Hum. Genet. 76, 8–32 (2005).
Noonan, J.P. & McCallion, A.S. Genomics of long-range regulatory elements. Annu. Rev. Genomics Hum. Genet. 11, 1–23 (2010).
Visel, A., Rubin, E.M. & Pennacchio, L.A. Genomic views of distant-acting enhancers. Nature 461, 199–205 (2009).
Lettice, L.A. et al. A long-range Shh enhancer regulates expression in the developing limb and Fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 12, 1725–1735 (2003).
Sakabe, N.J., Savic, D. & Nobrega, M.A. Transcriptional enhancers in development and disease. Genome Biol. 13, 238 (2012).
Pomerantz, M.M. et al. The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nat. Genet. 41, 882–884 (2009).
Tuupanen, S. et al. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat. Genet. 41, 885–890 (2009).
Wasserman, N.F., Aneas, I. & Nobrega, M.A. An 8q24 gene desert variant associated with prostate cancer risk confers differential in vivo activity to a MYC enhancer. Genome Res. 20, 1191–1197 (2010).
Duan, J. et al. Synonymous mutations in the human dopamine receptor D2 (DRD2) affect mRNA stability and synthesis of the receptor. Hum. Mol. Genet. 12, 205–216 (2003).
Burgner, D. et al. A genome-wide association study identifies novel and functionally related susceptibility loci for Kawasaki disease. PLoS Genet. 5, e1000319 (2009).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008).
Segrè, A.V., Groop, L., Mootha, V.K., Daly, M.J. & Altshuler, D. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Raychaudhuri, S. et al. Identifying relationships among genomic disease regions: predicting genes at pathogenic SNP associations and rare deletions. PLoS Genet. 5, e1000534 (2009).
Fransen, K. et al. Analysis of SNPs with an effect on gene expression identifies UBE2L3 and BCL3 as potential new risk genes for Crohn's disease. Hum. Mol. Genet. 19, 3482–3488 (2010).
Ernst, J. & Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol. 28, 817–825 (2010).
Schaub, M.A., Boyle, A.P., Kundaje, A., Batzoglou, S. & Snyder, M. Linking disease associations with regulatory information in the human genome. Genome Res. 22, 1748–1759 (2012).
John, S. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nat. Genet. 43, 264–268 (2011).
Cowper-Sal·lari, R. et al. Breast cancer risk–associated SNPs modulate the affinity of chromatin for FOXA1 and alter gene expression. Nat. Genet. 44, 1191–1191 (2012).
Kraft, P. & Hunter, D.J. Genetic risk prediction—Are we there yet? N. Engl. J. Med. 360, 1701–1703 (2009).
Yngvadottir, B., MacArthur, D.G., Jin, H. & Tyler-Smith, C. The promise and reality of personal genomics. Genome Biol. 10, 237 (2009).
Roberts, N.J. et al. The predictive capacity of personal genome sequencing. Sci. Transl. Med. 4, 133ra58 (2012).
Jostins, L. & Barrett, J.C. Genetic risk prediction in complex disease. Hum. Mol. Genet. 20, R182–R188 (2011).
Stahl, E.A. et al. Bayesian inference analyses of the polygenic architecture of rheumatoid arthritis. Nat. Genet. 44, 483–489 (2012).
Cooper, G.M. & Shendure, J. Needles in stacks of needles: finding disease-causal variants in a wealth of genomic data. Nat. Rev. Genet. 12, 628–640 (2011).
Gibson, G. Rare and common variants: twenty arguments. Nat. Rev. Genet. 13, 135–145 (2011).
Goldstein, D.B. The importance of synthetic associations will only be resolved empirically. PLoS Biol. 9, e1001008 (2011).
Acknowledgements
L.D.W. and M.K. were funded by NIH grants R01HG004037 and RC1HG005334 and US National Science Foundation CAREER grant 0644282.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Ward, L., Kellis, M. Interpreting noncoding genetic variation in complex traits and human disease. Nat Biotechnol 30, 1095–1106 (2012). https://doi.org/10.1038/nbt.2422
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2422