Abstract
The nonhuman primates most commonly used in medical research are from the genus Macaca1. To better understand the genetic differences between these animal models, we present high-quality draft genome sequences from two macaque species, the cynomolgus/crab-eating macaque and the Chinese rhesus macaque. Comparison with the previously sequenced Indian rhesus macaque reveals that all three macaques maintain abundant genetic heterogeneity, including millions of single-nucleotide substitutions and many insertions, deletions and gross chromosomal rearrangements. By assessing genetic regions with reduced variability, we identify genes in each macaque species that may have experienced positive selection. Genetic divergence patterns suggest that the cynomolgus macaque genome has been shaped by introgression after hybridization with the Chinese rhesus macaque. Macaque genes display a high degree of sequence similarity with human disease gene orthologs and drug targets. However, we identify several putatively dysfunctional genetic differences between the three macaque species, which may explain functional differences between them previously observed in clinical studies.
Similar content being viewed by others
Main
The macaques are the most widespread of nonhuman primates, comprising more than 20 species that diverged from each other up to 5–6 million years ago2. The Macaca genus is closely related to humans, sharing a last common ancestor ∼25 million years ago3. The close relationship between humans and macaques has made several species attractive as animal models for different biomedical analyses. Although the Indian subspecies of the rhesus macaque (Macaca mulatta mulatta) was originally the research model of choice, a ban on the export of this macaque has greatly reduced the availability of these animals, leading to increased use of other macaque species and/or subspecies, in particular the Chinese rhesus (CR) macaque (Macaca mulatta lasiota) and the cynomolgus or crab-eating (CE) macaque (Macaca fascicularis).
We sequenced the genomes of a female CR macaque and a female CE macaque using a whole-genome shotgun strategy on a next-generation sequencing platform. Briefly mitochondrial genome sequence analysis verified the predicted origin of both individuals (Supplementary Section 1). We then constructed 19 and 18 multiple paired-end genomic DNA libraries with gradually increasing insert sizes for the CR macaque and CE macaque, respectively. The total size of the assembled CR macaque and CE macaque genomes was, respectively, ∼2.84 Gb and 2.85 Gb, providing 47-fold and 54-fold coverage, respectively, on average (Table 1 and Supplementary Section 1). The scaffolds were assigned onto the chromosomes according to the synteny displayed with the Indian rhesus (IR) macaque4 and human genome sequences. About 97% of CR macaque scaffolds and 92% of CE macaque scaffolds could be placed onto chromosomes. We also applied RNA-seq to profile transcripts in various tissues from one IR macaque and two CE macaques (Online Methods). An integrated analysis combining genomic and transcriptome data was then used to define transcript structure and ascertain the expression profile of each gene (Supplementary Section 2).
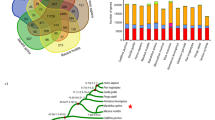
Macaque genetic diversity was evaluated by whole genome comparison and short read alignment using the IR macaque genome as a reference. In total, we detected >20 million single-nucleotide differences and 740,827 indel events in the three macaque species or subspecies (Supplementary Section 3), which will provide abundant genetic heterogeneity for use in future biomedical applications and analyses. We classified all of the single-nucleotide variable sites into three classes (shared, fixed and unique variants) based upon their presence or absence in the three individuals (Fig. 1a). Unique variants comprised >71.7% of the total variants, which is unsurprising given that even within a panmictic population, 44% of alleles are expected to be singletons in a sample of three individuals. It is noteworthy that a large number of genetic differences were shared between at least two macaques. Using only the fixed and unique variations, we estimated that the highest divergence rate, 0.40%, was between the CE macaque and the IR macaque (Fig. 1b). However, the sequence divergence between the CE macaque and CR macaque (0.34%), although nominally different species, was close to that observed between the subspecies, the CR macaque and the IR macaque (0.31%).
(a) Classification of single nucleotide divergence between macaque species. The ∼20 million single nucleotide differences among macaques were classified into three subclasses. The overlapping regions represent heterozygous variants shared between two individuals or all individuals. U, unique heterozygous variations evident in each species; F, the number of fixed homozygous variations in each species. (b) Single nucleotide divergence between macaque species in 100-kb windows across the genome. Heterozygous variants were ignored in this calculation. The divergence of X chromosomes between the two rhesus macaque subspecies was a significant outlier (P < 0.05, Grubbs' test). CE, crab-eating macaque; CR, Chinese rhesus macaque; IR, Indian rhesus macaque.
Recent research based on limited sequence data has suggested that an ancient introgression may have occurred from CR macaques to CE macaques living in an overlapping geographical distribution zone on the Indo-Chinese peninsula5,6,7. The two sequenced genomes allowed us to quantify the influence of this introgression at the whole-genome level. Specifically, we explored whether a DNA signal consistent with interspecies hybridization was apparent within the CR macaque and CE macaque genomes. We calculated the divergence ratio between the CE macaque and CR macaque and compared it with the divergence ratio between the CR macaque and IR macaque for 50-kb windows across the aligned genomes (Supplementary Section 4). For these calculations, we ignored variations at CpG sites because they are known to evolve particularly rapidly. Over 27% of the windows exhibited a divergence ratio less than zero, suggesting that CE and CR macaques are more closely related than the subspecies CR and IR macaques in these regions (Supplementary Section 4). In addition, >93% of 50-kb genomic windows displayed a lower divergence rate between the CE and CR macaques in comparison with the CE and IR macaques. Therefore, unsorted ancestral polymorphisms could not entirely explain the high proportion of inconsistent regions observed between the CE macaque and CR macaque. Furthermore, by combining previous single nucleotide polymorphism (SNP) data from IR and CR macaque populations with data from our own sequenced CR and CE macaque individuals8, we noted that our CE macaque individual clustered within the CR macaque population (Supplementary Section 4). This supports the occurrence of strong gene flow from the CR to the CE macaque genome. By screening the degree of asymmetry in the divergence between the CE and CR macaque and between the CE and IR macaque, we estimated that ∼30% of the CE macaque genome is of CR macaque origin (Supplementary Section 4).
We next sought to identify putative introgression regions (PIRs) in the CE macaque genome that might have been contributed by gene flow. We used simulated data (under a neutral no-migration model) as a control (Online Methods and Supplementary Section 4), and identified 8,942 PIRs spanning 778 Mb with a substantial lower-than-expected divergence rate between the CE and CR macaque (Fig. 2a,b). After merging overlapping PIRs, we found that most PIRs (>98%) were shorter than 500 kb. Because the length distribution of PIRs is a function of the time since gene flow occurred9, the prevalence of short PIRs suggests that gene flow occurred over an extended period of evolutionary time and was unlikely to have been simply a consequence of very recent human-mediated gene flow. We also observed a marked difference in variability between the X and autosomal chromosomes (Supplementary Section 4), which could have resulted from male-driven gene flow. One likely contributing factor to the restricted gene flow from CR macaque females to CE macaque males is that CR macaque females exhibit marked ovarian seasonality and only copulate during ovulation, whereas CE macaque females do not exhibit distinct reproductive seasonality and remain sexually receptive throughout the year10. Additionally, given that dispersal is primarily male-driven in macaques owing to female philopatry, this could also account for the gene flow from CR macaque males to CE macaque females and the absence of the reverse. These populations may therefore be of interest for studying physiological and behavioral aspects of reproduction between different species.
(a) The genetic distance between macaques (blue curve), human and macaques (red curve), and the distance between macaque species/subspecies (green curve for IR and CR; yellow curve for CR and CE) across chromosome 14. The dashed red line depicts the average genetic distance between human and macaque. The dotted blue line represents the average genetic distance between the macaques. The red bars at the bottom denote the candidate selective sweep regions, and the blue bars denote the putative introgression regions. The consecutive regions containing zero mutations in all species (such as the ∼20 Mb region) are sequencing gaps or alignment gap regions. (b) A potential introgression region (shaded blue), which contains fewer variations between CE macaque and CR macaque than between the two rhesus macaques (IR macaque and CR macaque). (c) A selective sweep region, encompassing 400 kb, which contains only one gene. The red bar denotes the coding region of the SBF2 gene; the red shaded box corresponds to the extent of the putative selective sweep.
Strong selection in favor of new advantageous alleles results in a 'selective sweep' which reduces genetic diversity relative to unselected regions. We developed an algorithm to identify putative sweep regions containing reduced variation between the three macaque species/subspecies, and generated simulated data under the assumption of a neutral model to evaluate its statistical significance (Online Methods and Supplementary Section 5). We identified 217 strong selective sweep regions that exhibited a reduced level of variation between macaques and that deviated substantially from neutral expectation (Fig. 2c, Supplementary Section 5 and Supplementary Table 17). Notably, one of the ten largest selective sweep regions, located on macaque chromosome 14, contains only one gene, the SET binding factor 2 (SBF2) (Fig. 2c). Thus, it is likely that this gene, which encodes a peripheral membrane protein from the protein-tyrosine phosphatase family, was the target of positive selection during the early evolution of macaques. Of potential biomedical interest, defects in the human SBF2 ortholog cause an autosomal recessive demyelinating form of Charcot-Marie-Tooth disease (CMT4B2).
To reveal the potential targets of positive selection in each macaque branch, we assigned 14,978 1:1 gene orthologs for human, chimpanzee and the three macaque species/subspecies by genome alignment (Supplementary Section 6). Comparison of the macaque ortholog trios revealed that they share an extremely high level of nucleotide sequence similarity within gene regions. It is noteworthy that 20.7% of the orthologs exhibit a higher degree of similarity between CR macaque and CE macaque than that between CR macaque and IR macaque, which may imply the influence of introgression. Gene Ontology–based gene category comparison between Macaca, Hominid and Murid lineages indicated that microtubule-based processes and the insulin receptor–signaling pathway evolved particularly rapidly in the Macaca lineage (Supplementary Section 6). Likelihood ratio tests based on a branch site model revealed 16 positively selected genes specifically in the IR macaque branch, 7 in the CR macaque branch and 13 in the CE macaque branch (Supplementary Section 6). It is intriguing that 31 of the 36 positively selected genes in macaques encode binding proteins that play major roles in regulating gene expression. It is also worth noting that two genes encoding dendrite proteins, CLCN2 in the IR macaque lineage, and activity-regulated, cytoskeleton-associated protein (ARC) in the CE macaque lineage, experienced positive selection. These two genes, together with another five positively selected genes, are already known to be relevant to human genetic disease, indicating the likely importance of their biological functions.
The availability of the CR macaque and CE macaque genome sequences allowed us to evaluate their genetic diversity, as well as the genetic differences between macaques and humans, which is important given the prominent use of macaques in biomedical research. Comparisons between the macaque genomes revealed the absence of 25 human single-copy genes (Online Methods and Supplementary Section 6), including a chemokine receptor gene, IL32, which may play a role in both innate and adaptive immune responses, and is consequently important to consider when these macaques are used in infectious disease studies. In addition, a total of 170 genes related to disease or immunity in one or another macaque species either contain frameshift mutations or premature stop codons, which would be predicted to have pseudogenized these genes (Supplementary Section 6). The authenticity of these truncating mutations is supported both by transcriptome data and high-depth sequencing reads, as well as independent PCR validation. Thirty-two of these genes function in immunity pathways and appear to have been lost in macaques. For example, an important innate-immunity gene, DEFA4, which encodes one of the microbicidal and cytotoxic peptides made by neutrophils11, has been pseudogenized in all three macaques because of loss of its first exon. Furthermore, the Toll-like receptor 4 (TLR4) gene contained a 1-bp deletion, which generates a premature stop codon in its third exon in all three macaques (Supplementary Section 6). TLR4 has been reported to have been under positive selection in Old World primates12. Notably, some human disease–related genes also contain frameshifts in their macaque homologs. For instance, we found that all three macaques had a premature stop codon in the second exon of the opioid receptor mu1 (OPRM1) gene, which encodes a protein distributed throughout the neuraxis and peripheral nervous system, and which is the primary target of opioids13 (Supplementary Section 6).
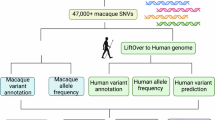
We also investigated genetic differences in orthologs that are specifically important in biomedical studies. The cytoplasmic tripartite-motif protein 5α (encoded by the TRIM5 gene), which can restrict replication of a broad range of retroviruses, is a key biomarker used to select animal models of HIV infection14. To survey the population-wide genetic diversity of TRIM5, we PCR amplified and sequenced TRIM5 from 33 unrelated CE macaque individuals of Vietnamese origin and 28 CR macaque individuals (Online Methods). We did not detect a previously reported15 Trim5-cyclophilin A chimera (TRIM-CypA2) in any individual, suggesting that this genotype is rare in these populations. However, 19 nonsynonymous polymorphisms and one microdeletion were identified in the TRIM5 gene relative to the IR macaque reference; nearly all of these polymorphisms displayed different frequencies between the two populations (Fig. 3 and Supplementary Section 7). We also identified a 6-bp deletion in the TRIM5 gene in the CE macaque that results in the loss of two amino acids (Thr339 and Phe340). Recent research has indicated that deletion of these residues could lead to increased HIV or SIV pathogenicity16. A high frequency (97.5%) of this mutation was detected in the CE macaque population, indicating that this deletion has become virtually fixed in the CE macaque. By contrast, in the CR macaque population, the frequency of this mutation is about 50%, only marginally higher than in the IR macaque population (36%)17. The variation in frequency of this 6-bp deletion and of other polymorphisms between macaques of different geographic origins may well be responsible for the observed differences in HIV resistance between these macaque species/subspecies16. We also surveyed genetic variation in other disease-related genes in the same population of CE or CR macaques, observing that mutations often occur at different frequencies in the two species (Supplementary Section 7).
(a) Schematic of protein encoded by TRIM5 in macaque. Annotated functional domains are marked with the names of domains in the colored boxes. The positions of nonsynonymous polymorphisms and the two-amino-acid deletion (in red) are marked.(b) The frequencies of all the nonsynonymous polymorphisms and the two-amino-acid deletion in the CR macaque and CE macaque populations. The frequency is counted for the genotype that appears in the IR macaque reference.
To study the orthologs of human druggable protein domains in macaques and to create a resource for the therapeutic exploitation of the 'druggable genome,' we screened the macaque orthologs for currently known drug domains. Almost all of the druggable orthologs can be detected in the three macaque species/subspecies, indicating that these animal models are likely to be functionally equivalent. However, in a very few cases, the ortholog found in macaque is different from its human counterpart. For instance, a mitochondrial acyltransferase (GLYATL2), which transfers an acyl group to glycine, has been completely lost in all three macaques. In addition, we identified 19 human genes with druggable domains, which have become pseudogenes in macaques (Supplementary Section 7). For example, the parathyroid hormone 1 receptor (PTH1R) gene, the target of the anti-osteoporosis drug teriparatide (Forteo)18,19, contains a premature stop codon in macaques. One of the targets of recombinant human keratinocyte growth factor (Palifermin20), fibroblast growth factor receptor 3, encoded by FGFR3, has also been pseudogenized in macaques owing to the presence of a premature stop codon.
Of additional biomedical interest are compensated pathogenic deviations. These represent human putatively pathological missense alleles where the substituting amino acids are identical to the wild-type amino acid residues at orthologous positions in other organisms. We identified 931 compensated pathogenic deviations in four closely related primate species (chimpanzee and the three macaques), of which 220 varied between the nonhuman primates, including 65 that varied between the three macaque species (Supplementary Section 8 and Supplementary Table 26). For example, one mutation (R40→H40) in the ornithine transcarbamylase (OTC) gene was evident in the two rhesus macaque subspecies but not in the CE macaque. Based on the examples of identified genetic differences outlined above, it is clear that the potential existence of such interspecies differences should be considered when selecting macaques for use as disease models.
Comparison of gene expression profiles (Supplementary Section 9) between the CE macaque and the IR macaque revealed that their orthologs displayed conserved expression profiles in the same tissue. However, we noted that the testis exhibited expression levels that were more divergent between those orthologs that had lower Pearson correlation coefficients (Supplementary Section 9). The observation that more genes display inconsistent expression levels in testis as compared with the other tissues might be related to the rapid evolutionary rate manifested by primate sperm-expressed genes21. Transcriptome data also served to identify several novel genes in CE macaques with respect to rhesus macaques.
In conclusion, our sequencing and analyses of two macaque genomes confirmed that introgressive hybridization probably played an important role in the formation of the genome of the extant mainland-origin CE macaque. Thus, the CE macaque could be a useful model for exploring gene interchange between primate species, and the consequent role of this process in primate evolution and speciation. The two new macaque genomes presented here also highlight the degree of variation existing between these widely used nonhuman primate animal models. The abundant genetic diversity evident in individual macaques from distinct geographic populations is of direct interest to primatology, preclinical medicine, population genetics and phylogeographic studies.
Methods
Source of samples.
A 5-year-old female CR macaque and a 4-year-old female CE macaque of Vietnamese origin were used in this study. The CR macaque individual was descended from an individual captured from the wild in Yunnan Province. The origins of these two individuals were confirmed by mitochondrial DNA sequencing. Genomic DNA was collected from the peripheral blood cells of these two individuals. Two CE macaques of Indonesian origin were euthanized for tissue collection in transcriptome sequencing. Samples from brain, kidney, liver and white adipose tissue were collected from a 2-year-old male whereas tissue from testis and ileum were collected from a 6-year-old male. One male Rhesus macaque of Indian origin was euthanized for tissue collection in transcriptome sequencing. Samples from brain, heart, kidney, liver, quadriceps and testis were collected. We declare that the experiments on animals involved in this study have been approved by the institutional committee.
Sample preparation and sequencing.
We constructed 19 and 18 paired-end libraries, with spanning size ranges of 200 bp to 10 kb (Supplementary Section 1), from the CR macaque and CE macaque, respectively. The libraries were prepared following the manufacturer's standard instructions and sequenced on Illumina HiSeq (2000) platform. Whole genome sequencing was done as described previously22. A total of 178.98 Gb data and 198.39 Gb data were generated from these libraries for the CR macaque and CE macaque, respectively.
Genome assembly.
Before assembly, a series of filtering steps were undertaken to filter the low-quality sequencing reads. A total of 142 G (or 47.4×) and 162 G (or 54.0×) data for the CR macaque and CE macaque, respectively, were retained for assembly. The two macaque genomes were assembled de novo by the de Bruijn graph–based assembler SOAPdenovo23 (http://soap.genomics.org.cn/). The reads from the short insert size libraries (<2 kb) were first used to build the contigs, then all the paired-end reads were realigned onto the contig sequences to construct the scaffolds. We then determined the extent of the shared paired-end relationships between each pair of contigs, weighted the rate of consistent and conflicting paired ends and then constructed the scaffolds step by step, in increasing order of insert size. Finally, we used the paired-end information to retrieve the read pairs (that had one end mapped to the unique contig and the other located in the gap region) and performed a local assembly for these collected reads to fill the gaps. The statistics on genome assembly are shown in Table 1. The scaffolds were then mapped and assembled onto the chromosomes based upon their syntenies with the IR macaque genome and the human genome.
Transcriptome sequencing.
Tissue RNA extraction was performed using the QiagenRNeasy Kit. RNA sequencing libraries were constructed using an Illumina standard mRNA-Seq Prep Kit. The paired-end libraries were sequenced on the IlluminaHiSeq for 100 bp at each end.
Single nucleotide variation (SNV) detection.
To identify the SNVs among macaques, we aligned all the high-quality reads onto the rheMac2 assembly using SOAPaligner24 with gap-free mode, allowing two mismatches for 44-bp reads or 5 mismatches for 75-bp reads. The SOAPsnp25 was used in SNV calling. After quality control and filtering, 9.4M SNVs (37.34% homozygous, 62.66% heterozygous) and 12.0M SNVs (44.29% homozygous, 55.71% heterozygous) were identified from the CR macaque and CE macaque, respectively, in relation to the IR macaque reference.
Detection of putative introgression regions.
We first used the degree of asymmetry in the divergence between CE macaque/CR macaque and CE macaque/IR macaque to estimate the proportion of the CE macaque genome that is of CR macaque origin. In particular, the proportion of the genome that is introgressed (m) should adhere to the following equation: m = (D13 − D12)/(D13 − D22), where Dij is the proportion of average pair-wise differences from individuals sampled from populations i and j, and the population indices are CE macaque: 1, CR: 2, IR macaque: 3. Using this equation, we found that ∼30% of the CE macaque genome is of CR macaque origin. We next sought to identify putative introgression regions (PIRs) in the macaque. We noted that if a given chromosomal region had originated as a consequence of hybridization between CE macaques and CR macaques, the sequence diversity between CE macaques and CR macaques (denoted as DIVCE–CR) should be lower than that between CR macaques and IR macaques (denoted as DIVIR–CR). The diversity between two species/subspecies was scaled in terms of genetic distance using the matrix given in Supplementary Section 4. Then, we calculated DIVCE–CR and DIVCR–IR for nonoverlapping windows of fixed size (denoted  and
and  for the window i) using the method described below:
for the window i) using the method described below:

where S1 and S2 are two different species/subspecies. Further, we introduced a statistic to quantify the difference between  and
and  :
:

Under this definition, a negative  indicates that CR macaque is closer to CE macaque than to IR macaque. To filter out the regions where the CR macaque sequence was closer to CE macaque by chance alone, we generated simulation data assuming a neutral model using parameters estimated by demographic analysis. The demographic model used for simulation assumed no migration between CE macaque and CR macaque. The program ms26 was used to generate segregating sites for the three macaques. Then, we calculated Rdiff for each window in the simulated data. The 1% quartile of Rdiff in the simulated data was used as a cut-off (denoted as Rcutoff), that is P(Rdiff <= Rcutoff)=0.01, for all windows in simulated data.
indicates that CR macaque is closer to CE macaque than to IR macaque. To filter out the regions where the CR macaque sequence was closer to CE macaque by chance alone, we generated simulation data assuming a neutral model using parameters estimated by demographic analysis. The demographic model used for simulation assumed no migration between CE macaque and CR macaque. The program ms26 was used to generate segregating sites for the three macaques. Then, we calculated Rdiff for each window in the simulated data. The 1% quartile of Rdiff in the simulated data was used as a cut-off (denoted as Rcutoff), that is P(Rdiff <= Rcutoff)=0.01, for all windows in simulated data.
Then, the cut-off from the simulated data was applied to our actual data, with  to predict PIRs. We performed this analysis using a series of window sizes from 10 kb to 1 Mb. The total size of PIRs is shown in Supplementary Figure 14. We found that fewer PIRs could be detected using small windows (10 kb and 20 kb) or large windows (>100 kb). This could be due to the smaller number of SNVs existing within a small window, which can reduce the power to detect PIRs. Although the large window may have exceeded the average size of real PIRs, the total PIR sizes do not change much when 40 kb to 100 kb windows are used, yielding a peak at 50 kb. To maximize the precision of PIR detection, we used 50 kb in our final version.
to predict PIRs. We performed this analysis using a series of window sizes from 10 kb to 1 Mb. The total size of PIRs is shown in Supplementary Figure 14. We found that fewer PIRs could be detected using small windows (10 kb and 20 kb) or large windows (>100 kb). This could be due to the smaller number of SNVs existing within a small window, which can reduce the power to detect PIRs. Although the large window may have exceeded the average size of real PIRs, the total PIR sizes do not change much when 40 kb to 100 kb windows are used, yielding a peak at 50 kb. To maximize the precision of PIR detection, we used 50 kb in our final version.
Detection of selective sweeps.
We used the HKA test26,27 to detect regions that contain potential selective sweeps and which have a low degree of divergence among macaques but normal levels of divergence between macaques and the outgroup (here we used human as the outgroup). The χ2 statistic, used for measuring the goodness-of-fit, was obtained after application of the HKA test for each window, and was used to infer the putative selective sweeps. We performed the simulations based on the demographic model of three macaque populations to calculate the significance of the deviations.
Pseudogene identification.
To detect homozygous pseudogenes in the three macaques, we first aligned all the human cDNAs (Ensembl release-56) onto the three macaque genomes using BLAT and used Exonerate software to predict the precise exon-intron structure for each gene in macaques. All homologous genes containing frameshifts or premature stop codons were considered as candidate pseudogenes. A series of filtering steps, including filtering artificial results resulting from mis-annotation, validation with high-depth sequencing reads or transcriptome reads, were done to identify pseudogenes with high confidence. For PCR validation, 1 kb genome sequences flanking each frameshift or premature stop codon site were cut down from the three macaque genomes and the human genome for PCR primer design. PCR amplification products from different macaque species were sequenced using an ABI-3730.
Population study.
To characterize the polymorphisms of several important variants between the CE and CR macaques, we carried out a population survey in 33 unrelated CE macaque individuals of Vietnamese origin and 28 CR macaque individuals using PCR amplification and sequencing.
Accession codes.
The CR macaque (M. mulatta) and the CE macaque (M. fascicularis) whole genome shotgun projects have been deposited at DDBJ/EMBL/GenBank under the accession numbers AEHK00000000 and AEHL00000000. The versions described in this Letter are AEHK01000000 and AEHL01000000. All short read data have been deposited into the Short Read Archive under accession numbers SRA023855 and SRA023856. Raw sequencing data of transcriptome have been deposited in Gene Expression Omnibus as GSE29629. Genome assemblies are also available using the following data DOIs at our CLiMB repository: doi:10.5524/100002 and doi:10.5524/100003 <http://dx.doi.org/10.5524/100002 and < http://dx.doi.org/10.5524/100003>.
Accession codes
Accessions
DDBJ/GenBank/EMBL
Gene Expression Omnibus
Sequence Read Archive
References
Carlsson, H.E., Schapiro, S.J., Farah, I. & Hau, J. Use of primates in research: a global overview. Am. J. Primatol. 63, 225–237 (2004).
Fooden, J. The Macaques: Studies in Ecology, Behavior, and Evolution (Van Nostrand-Reinhold, 1980).
Kumar, S. & Hedges, S.B. A molecular timescale for vertebrate evolution. Nature 392, 917–920 (1998).
Gibbs, R.A. et al. Evolutionary and biomedical insights from the rhesus macaque genome. Science 316, 222–234 (2007).
Bonhomme, M., Cuartero, S., Blancher, A. & Crouau-roy, B. Assessing natural introgression in 2 biomedical model species, the rhesus macaque (Macaca mulatta) and the long-tailed macaque (Macaca fascicularis). J. Hered. 100, 158–169 (2009).
Stevison, L.S. & Kohn, M.H. Divergence population genetic analysis of hybridization between rhesus and cynomolgus macaques. Mol. Ecol. 18, 2457–2475 (2009).
Tosi, A., Morales, J. & Melnick, D. Y-chromosome and mitochondrial markers in Macaca fascicularis indicate introgression with Indochinese M. mulatta and a biogeographic barrier in the Isthmus of Kra. Int. J. Primatol. 23, 161–178 (2002).
Hernandez, R.D. et al. Demographic histories and patterns of linkage disequilibrium in Chinese and Indian rhesus macaques. Science 316, 240–243 (2007).
Pool, J.E. & Nielsen, R. Inference of historical changes in migration rate from the lengths of migrant tracts. Genetics 181, 711–719 (2009).
Weinbauer, G. et al. Physiology and endocrinology of the ovarian cycle in macaques. Toxicol. Pathol. 36, 7S–23S (2008).
Palfree, R.G., Sadro, L.C. & Solomon, S. The gene encoding the human corticostatin HP-4 precursor contains a recent 86-base duplication and is located on chromosome 8. Mol. Endocrinol. 7, 199–205 (1993).
Wlasiuk, G. & Nachman, M.W. Adaptation and constraint at Toll-like receptors in primates. Mol. Biol. Evol. 27, 2172–2186 (2010).
Kosarac, B., Fox, A.A. & Collard, C.D. Effect of genetic factors on opioid action. Curr. Opin. Anaesthesiol. 22, 476–482 (2009).
Wolf, D. & Goff, S.P. Host restriction factors blocking retroviral replication. Annu. Rev. Genet. 42, 143–163 (2008).
Newman, R.M. et al. Evolution of a TRIM5-CypA splice isoform in old world monkeys. PLoS Pathogens 4, e1000003 (2008).
Yap, M.W., Nisole, S. & Stoye, J.P. A single amino acid change in the SPRY domain of human Trim5alpha leads to HIV-1 restriction. Curr. Biol. 15, 73–78 (2005).
Lim, S.Y. et al. TRIM5alpha modulates immunodeficiency virus control in rhesus monkeys. PLoS Pathog. 6, e1000738 (2010).
Brixen, K.T., Christensen, P.M., Ejersted, C. & Langdahl, B.L. Teriparatide (biosynthetic human parathyroid hormone 1–34): a new paradigm in the treatment of osteoporosis. Basic Clin. Pharmacol. Toxicol. 94, 260–270 (2004).
Chen, X., Ji, Z.L. & Chen, Y.Z. TTD: therapeutic target database. Nucleic Acids Res. 30, 412–415 (2002).
Cancilla, B., Davies, A., Cauchi, J.A., Risbridger, G.P. & Bertram, J.F. Fibroblast growth factor receptors and their ligands in the adult rat kidney. Kidney Int. 60, 147–155 (2001).
Khaitovich, P., Enard, W., Lachmann, M. & Paabo, S. Evolution of primate gene expression. Nat. Rev. Genet. 7, 693–702 (2006).
Li, R. et al. The sequence and de novo assembly of the giant panda genome. Nature 463, 311–317 (2010).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
Hudson, R.R. Generating samples under a Wright-Fisher neutral model of genetic variation. Bioinformatics 18, 337–338 (2002).
Hudson, R.R., Kreitman, M. & Aguade, M. A test of neutral molecular evolution based on nucleotide data. Genetics 116, 153–159 (1987).
Acknowledgements
We thank the staff at the Beijing Genomics Institute in Shenzhen and in Merck Research Laboratories whose names were not included in the authors list but who nevertheless contributed to this project. We thank L. Goodman for providing valuable suggestions for this analysis and helping to edit the manuscript. This project was funded by the Southern China Center for Innovative Pharmaceuticals (SCCIP), the National Natural Science Foundation of China (30725008), the Shenzhen Municipal Government of China (grant no. CXB200903110066A), the Major State Basic Research Development Program of China (project no. 2006CB701500), the National Basic Research Program of China (2007CB512402) and the National Science and Technology Major Project of Key Drug Innovation and Development (2011ZX09307-303-03).
Author information
Authors and Affiliations
Contributions
G.Y., G.Z., X.F., Yanfeng Zhang, C.L. and F.L. contributed equally to this work. G.Y., Jun Wang, X.W. and Jian Wang managed the project. H.D., X.Z., L.L., F.L., Jufang Wang, T.W. and L.W. prepared the DNA and performed sequencing and PCR-based experiments. G.Z., X.F., Yanfeng Zhang, C.L., Q.L., Yong Zhang, P.D.S., Yan Li, X.L., P.Z., Z.H., J.L., Y.M., Y.B., M.Z., T.W., J.C., L.W., J.L., Y.H., Z.W., C.L., H.H., B.L., N.A., Q.H., W.F., Y.L., D.N.C., R.N., M.G.K. and E.V.B. performed the genome assembly, gene annotation, and human disease gene, druggable domain, gene evolution, introgression and selective sweep analyses. R.C., J.R.T., A.J.v.G. and P.Y. coordinated the RNA sequencing and analysis. G.Y., G.Z. and X.F. wrote the manuscript while D.N.C., W.W., B.S., R.B., H.Y., Jun Wang, X.W. and Jian Wang revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Sections 1–9 (PDF 1891 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar license.
About this article
Cite this article
Yan, G., Zhang, G., Fang, X. et al. Genome sequencing and comparison of two nonhuman primate animal models, the cynomolgus and Chinese rhesus macaques. Nat Biotechnol 29, 1019–1023 (2011). https://doi.org/10.1038/nbt.1992
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1992
This article is cited by
-
Design, methodology, and preliminary results of the non-human primates eye study
BMC Ophthalmology (2023)
-
Brain region-specific effects of nearly fixed sapiens-derived alleles
BMC Genomic Data (2022)
-
Temporal mapping of derived high-frequency gene variants supports the mosaic nature of the evolution of Homo sapiens
Scientific Reports (2022)
-
Evolutionary pattern of Macaca fascicularis in Southeast Asia inferred using Y-chromosomal gene
BMC Ecology and Evolution (2021)
-
Aggregation-induced emission luminogens for image-guided surgery in non-human primates
Nature Communications (2021)