Abstract
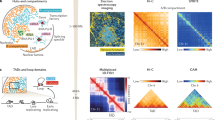
The spatial organization of chromosomes inside the cell nucleus is still poorly understood. This organization is guided by intra- and interchromosomal contacts and by interactions of specific chromosomal loci with relatively fixed nuclear 'landmarks' such as the nuclear envelope and the nucleolus. Researchers have begun to use new molecular genome-wide mapping techniques to uncover both types of molecular interactions, providing insights into the fundamental principles of interphase chromosome folding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pombo, A. & Branco, M.R. Functional organisation of the genome during interphase. Curr. Opin. Genet. Dev. 17, 451–455 (2007).
Misteli, T. Beyond the sequence: cellular organization of genome function. Cell 128, 787–800 (2007).
Zhao, R., Bodnar, M.S. & Spector, D.L. Nuclear neighborhoods and gene expression. Curr. Opin. Genet. Dev. 19, 172–179 (2009).
Hetzer, M.W. & Wente, S.R. Border control at the nucleus: biogenesis and organization of the nuclear membrane and pore complexes. Dev. Cell 17, 606–616 (2009).
Stuurman, N., Heins, S. & Aebi, U. Nuclear lamins: their structure, assembly, and interactions. J. Struct. Biol. 122, 42–66 (1998).
Herrmann, H. & Aebi, U. Intermediate filaments: molecular structure, assembly mechanism, and integration into functionally distinct intracellular Scaffolds. Annu. Rev. Biochem. 73, 749–789 (2004).
Prokocimer, M. et al. Nuclear lamins: key regulators of nuclear structure and activities. J. Cell Mol. Med. 13, 1059–1085 (2009).
Franke, W.W. Structure, biochemistry, and functions of the nuclear envelope. Int. Rev. Cytol. 4 (suppl.), 71–236 (1974).
Blobel, G. Gene gating: a hypothesis. Proc. Natl. Acad. Sci. USA 82, 8527–8529 (1985).
Takizawa, T., Meaburn, K.J. & Misteli, T. The meaning of gene positioning. Cell 135, 9–13 (2008).
Fedorova, E. & Zink, D. Nuclear genome organization: common themes and individual patterns. Curr. Opin. Genet. Dev. 19, 166–171 (2009).
Greil, F., Moorman, C. & van Steensel, B. DamID: mapping of in vivo protein-genome interactions using tethered DNA adenine methyltransferase. Methods Enzymol. 410, 342–359 (2006).
Vogel, M.J., Peric-Hupkes, D. & van Steensel, B. Detection of in vivo protein-DNA interactions using DamID in mammalian cells. Nat. Protoc. 2, 1467–1478 (2007).
Pickersgill, H. et al. Characterization of the Drosophila melanogaster genome at the nuclear lamina. Nat. Genet. 38, 1005–1014 (2006).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome— nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Shevelyov, Y.Y. et al. The B-type lamin is required for somatic repression of testis-specific gene clusters. Proc. Natl. Acad. Sci. USA 106, 3282–3287 (2009).
Reddy, K.L., Zullo, J.M., Bertolino, E. & Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452, 243–247 (2008).
Finlan, L.E. et al. Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet. 4, e1000039 (2008).
Kumaran, R.I. & Spector, D.L. A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J. Cell Biol. 180, 51–65 (2008).
Casolari, J.M. et al. Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization. Cell 117, 427–439 (2004).
Brown, C.R., Kennedy, C.J., Delmar, V.A., Forbes, D.J. & Silver, P.A. Global histone acetylation induces functional genomic reorganization at mammalian nuclear pore complexes. Genes Dev. 22, 627–639 (2008).
Kalverda, B., Pickersgill, H., Shloma, V.V. & Fornerod, M. Nucleoporins directly stimulate expression of developmental and cell-cycle genes inside the nucleoplasm. Cell 140, 360–371 (2010).
Capelson, M. et al. Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell 140, 372–383 (2010).
Vaquerizas, J.M. et al. Nuclear pore proteins nup153 and megator define transcriptionally active regions in the Drosophila genome. PLoS Genet. 6, e1000846 (2010).
Schermelleh, L. et al. Subdiffraction multicolor imaging of the nuclear periphery with 3D structured illumination microscopy. Science 320, 1332–1336 (2008).
Németh, A. et al. Initial genomics of the human nucleolus. PLoS Genet. 6, e1000889 (2010).
Stahl, A., Hartung, M., Vagner-Capodano, A.M. & Fouet, C. Chromosomal constitution of nucleolus-associated chromatin in man. Hum. Genet. 35, 27–34 (1976).
Thompson, M., Haeusler, R.A., Good, P.D. & Engelke, D.R. Nucleolar clustering of dispersed tRNA genes. Science 302, 1399–1401 (2003).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Tolhuis, B., Palstra, R.J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active β-globin locus. Mol. Cell 10, 1453–1465 (2002).
Murrell, A., Heeson, S. & Reik, W. Interaction between differentially methylated regions partitions the imprinted genes Igf2 and H19 into parent-specific chromatin loops. Nat. Genet. 36, 889–893 (2004).
Spilianakis, C.G. & Flavell, R.A. Long-range intrachromosomal interactions in the T helper type 2 cytokine locus. Nat. Immunol. 5, 1017–1027 (2004).
Vernimmen, D., De Gobbi, M., Sloane-Stanley, J.A., Wood, W.G. & Higgs, D.R. Long-range chromosomal interactions regulate the timing of the transition between poised and active gene expression. EMBO J. 26, 2041–2051 (2007).
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
Zhao, Z. et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat. Genet. 38, 1341–1347 (2006).
Dostie, J. et al. Chromosome Conformation Capture Carbon Copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Horike, S., Cai, S., Miyano, M., Cheng, J.F. & Kohwi-Shigematsu, T. Loss of silent-chromatin looping and impaired imprinting of DLX5 in Rett syndrome. Nat. Genet. 37, 31–40 (2005).
Tiwari, V.K., Cope, L., McGarvey, K.M., Ohm, J.E. & Baylin, S.B. A novel 6C assay uncovers Polycomb-mediated higher order chromatin conformations. Genome Res. 18, 1171–1179 (2008).
Fullwood, M.J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009).
Simonis, M., Kooren, J. & de Laat, W. An evaluation of 3C-based methods to capture DNA interactions. Nat. Methods 4, 895–901 (2007).
Dekker, J. The three 'C' s of chromosome conformation capture: controls, controls, controls. Nat. Methods 3, 17–21 (2006).
Xu, N., Tsai, C.L. & Lee, J.T. Transient homologous chromosome pairing marks the onset of X inactivation. Science 311, 1149–1152 (2006).
Bacher, C.P. et al. Transient colocalization of X-inactivation centres accompanies the initiation of X inactivation. Nat. Cell Biol. 8, 293–299 (2006).
Xu, N., Donohoe, M.E., Silva, S.S. & Lee, J.T. Evidence that homologous X-chromosome pairing requires transcription and Ctcf protein. Nat. Genet. 39, 1390–1396 (2007).
Sandhu, K.S. et al. Nonallelic transvection of multiple imprinted loci is organized by the H19 imprinting control region during germline development. Genes Dev. 23, 2598–2603 (2009).
Rodley, C.D., Bertels, F., Jones, B. & O'Sullivan, J.M. Global identification of yeast chromosome interactions using genome conformation capture. Fungal Genet. Biol. 46, 879–886 (2009).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Duan, Z. et al. A three-dimensional model of the yeast genome. Nature 465, 363–367 (2010).
Shopland, L.S. et al. Folding and organization of a contiguous chromosome region according to the gene distribution pattern in primary genomic sequence. J. Cell Biol. 174, 27–38 (2006).
Dekker, J. Mapping in vivo chromatin interactions in yeast suggests an extended chromatin fiber with regional variation in compaction. J. Biol. Chem. 283, 34532–34540 (2008).
Taddei, A., Schober, H. & Gasser, S.M. The budding yeast nucleus. Cold Spring Harb. Perspect. Biol. 2, a000612 (2010).
Hiratani, I. et al. Global reorganization of replication domains during embryonic stem cell differentiation. PLoS Biol. 6, e245 (2008).
Schwaiger, M. et al. Chromatin state marks cell-type- and gender-specific replication of the Drosophila genome. Genes Dev. 23, 589–601 (2009).
O'Keefe, R.T., Henderson, S.C. & Spector, D.L. Dynamic organization of DNA replication in mammalian cell nuclei: spatially and temporally defined replication of chromosome-specific α-satellite DNA sequences. J. Cell Biol. 116, 1095–1110 (1992).
Ryba, T. et al. Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res. 20, 761–770 (2010).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R.A. & Feinberg, A.P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat. Genet. 41, 246–250 (2009).
Yokochi, T. et al. G9a selectively represses a class of late-replicating genes at the nuclear periphery. Proc. Natl. Acad. Sci. USA 106, 19363–19368 (2009).
Gilbert, N. et al. Chromatin architecture of the human genome: gene-rich domains are enriched in open chromatin fibers. Cell 118, 555–566 (2004).
Phillips, J.E. & Corces, V.G. CTCF: master weaver of the genome. Cell 137, 1194–1211 (2009).
Splinter, E. et al. CTCF mediates long-range chromatin looping and local histone modification in the β-globin locus. Genes Dev. 20, 2349–2354 (2006).
Majumder, P., Gomez, J.A., Chadwick, B.P. & Boss, J.M. The insulator factor CTCF controls MHC class II gene expression and is required for the formation of long-distance chromatin interactions. J. Exp. Med. 205, 785–798 (2008).
Soutoglou, E. & Misteli, T. Mobility and immobility of chromatin in transcription and genome stability. Curr. Opin. Genet. Dev. 17, 435–442 (2007).
Chuang, C.H. & Belmont, A.S. Moving chromatin within the interphase nucleus-controlled transitions? Semin. Cell Dev. Biol. 18, 698–706 (2007).
Bolzer, A. et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. 3, e157 (2005).
Osborne, C.S. et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat. Genet. 36, 1065–1071 (2004).
Spilianakis, C.G., Lalioti, M.D., Town, T., Lee, G.R. & Flavell, R.A. Interchromosomal associations between alternatively expressed loci. Nature 435, 637–645 (2005).
Miele, A., Bystricky, K. & Dekker, J. Yeast silent mating type loci form heterochromatic clusters through silencer protein-dependent long-range interactions. PLoS Genet. 5, e1000478 (2009).
Shaw, C.J. & Lupski, J.R. Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum. Mol. Genet. 13 Spec No 1, R57–R64 (2004).
Mitelman, F., Johansson, B. & Mertens, F. The impact of translocations and gene fusions on cancer causation. Nat. Rev. Cancer 7, 233–245 (2007).
Harewood, L. et al. The effect of translocation-induced nuclear reorganization on gene expression. Genome Res. 20, 554–564 (2010).
Simonis, M. et al. High-resolution identification of balanced and complex chromosomal rearrangements by 4C technology. Nat. Methods 6, 837–842 (2009).
Roix, J.J., McQueen, P.G., Munson, P.J., Parada, L.A. & Misteli, T. Spatial proximity of translocation-prone gene loci in human lymphomas. Nat. Genet. 34, 287–291 (2003).
Lin, C. et al. Nuclear receptor-induced chromosomal proximity and DNA breaks underlie specific translocations in cancer. Cell 139, 1069–1083 (2009).
Mani, R.S. et al. Induced chromosomal proximity and gene fusions in prostate cancer. Science 326, 1230 (2009).
Worman, H.J., Fong, L.G., Muchir, A. & Young, S.G. Laminopathies and the long strange trip from basic cell biology to therapy. J. Clin. Invest. 119, 1825–1836 (2009).
Goldman, R.D. et al. Accumulation of mutant lamin A causes progressive changes in nuclear architecture in Hutchinson-Gilford progeria syndrome. Proc. Natl. Acad. Sci. USA 101, 8963–8968 (2004).
Taimen, P. et al. A progeria mutation reveals functions for lamin A in nuclear assembly, architecture, and chromosome organization. Proc. Natl. Acad. Sci. USA 106, 20788–20793 (2009).
Pegoraro, G. et al. Ageing-related chromatin defects through loss of the NURD complex. Nat. Cell Biol. 11, 1261–1267 (2009).
Acknowledgements
We thank members of the van Steensel and Dekker labs and M. Walhout for suggestions. This work was supported by the Netherlands Genomics Initiative and an Netherlands Organization for Scientific Research–Earth and Life Sciences (NWO-ALW) VICI grant to B.v.S., a grant from the US National Institutes of Health (HG003143) and a W.M. Keck Foundation Distinguished Young Scholar Award to J.D.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
van Steensel, B., Dekker, J. Genomics tools for unraveling chromosome architecture. Nat Biotechnol 28, 1089–1095 (2010). https://doi.org/10.1038/nbt.1680
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1680
This article is cited by
-
StackEPI: identification of cell line-specific enhancer–promoter interactions based on stacking ensemble learning
BMC Bioinformatics (2022)
-
Regulation and dysregulation of spatial chromatin structure in the central nervous system
Anatomical Science International (2021)
-
3DeFDR: statistical methods for identifying cell type-specific looping interactions in 5C and Hi-C data
Genome Biology (2020)
-
Modeling and analysis of Hi-C data by HiSIF identifies characteristic promoter-distal loops
Genome Medicine (2020)
-
Prediction of enhancer–promoter interactions using the cross-cell type information and domain adversarial neural network
BMC Bioinformatics (2020)