Abstract
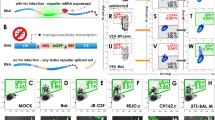
Hepatitis C virus (HCV), which infects 2–3% of the world population, is a causative agent of chronic hepatitis and the leading indication for liver transplantation1. The ability to propagate HCV in cell culture (HCVcc) is a relatively recent breakthrough and a key tool in the quest for specific antiviral therapeutics. Monitoring HCV infection in culture generally involves bulk population assays, use of genetically modified viruses and/or terminal processing of potentially precious samples. Here we develop a cell-based fluorescent reporter system that allows sensitive distinction of individual HCV-infected cells in live or fixed samples. We demonstrate use of this technology for several previously intractable applications, including live-cell imaging of viral propagation and host response, as well as visualizing infection of primary hepatocyte cultures. Integration of this reporter with modern image-based analysis methods could open new doors for HCV research.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Shepard, C.W., Finelli, L. & Alter, M.J. Global epidemiology of hepatitis C virus infection. Lancet Infect. Dis. 5, 558–567 (2005).
Bartenschlager, R. & Sparacio, S. Hepatitis C virus molecular clones and their replication capacity in vivo and in cell culture. Virus Res. 127, 195–207 (2007).
Bartenschlager, R. Hepatitis C virus molecular clones: from cDNA to infectious virus particles in cell culture. Curr. Opin. Microbiol. 9, 416–422 (2006).
Meylan, E. et al. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature 437, 1167–1172 (2005).
Foy, E. et al. Control of antiviral defenses through hepatitis C virus disruption of retinoic acid-inducible gene-I signaling. Proc. Natl. Acad. Sci. USA 102, 2986–2991 (2005).
Li, X.D., Sun, L., Seth, R.B., Pineda, G. & Chen, Z.J. Hepatitis C virus protease NS3/4A cleaves mitochondrial antiviral signaling protein off the mitochondria to evade innate immunity. Proc. Natl. Acad. Sci. USA 102, 17717–17722 (2005).
Kawai, T. et al. IPS-1, an adaptor triggering RIG-I- and Mda5-mediated type I interferon induction. Nat. Immunol. 6, 981–988 (2005).
Seth, R.B., Sun, L., Ea, C.K. & Chen, Z.J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-kappaB and IRF 3. Cell 122, 669–682 (2005).
Xu, L.G. et al. VISA is an adapter protein required for virus-triggered IFN-beta signaling. Mol. Cell 19, 727–740 (2005).
Tscherne, D.M. et al. Superinfection exclusion in cells infected with hepatitis C virus. J. Virol. 81, 3693–3703 (2007).
Loo, Y.M. et al. Viral and therapeutic control of IFN-beta promoter stimulator 1 during hepatitis C virus infection. Proc. Natl. Acad. Sci. USA 103, 6001–6006 (2006).
Simmonds, P. Genetic diversity and evolution of hepatitis C virus-15 years on. J. Gen. Virol. 85, 3173–3188 (2004).
Marukian, S. et al. Cell culture-produced hepatitis C virus does not infect peripheral blood mononuclear cells. Hepatology 48, 1843–1850 (2008).
Carroll, S.S. et al. Inhibition of hepatitis C virus RNA replication by 2′-modified nucleoside analogs. J. Biol. Chem. 278, 11979–11984 (2003).
Lin, K., Perni, R.B., Kwong, A.D. & Lin, C. VX-950, a novel hepatitis C virus (HCV) NS3–4A protease inhibitor, exhibits potent antiviral activities in HCV replicon cells. Antimicrob. Agents Chemother. 50, 1813–1822 (2006).
Pileri, P. et al. Binding of hepatitis C virus to CD81. Science 282, 938–941 (1998).
Scarselli, E. et al. The human scavenger receptor class B type I is a novel candidate receptor for the hepatitis C virus. EMBO J. 21, 5017–5025 (2002).
Evans, M.J. et al. Claudin-1 is a hepatitis C virus co-receptor required for a late step in entry. Nature 446, 801–805 (2007).
Ploss, A. et al. Human occludin is a hepatitis C virus entry factor required for infection of mouse cells. Nature 457, 882–886 (2009).
Timpe, J.M. et al. Hepatitis C virus cell-cell transmission in hepatoma cells in the presence of neutralizing antibodies. Hepatology 47, 17–24 (2008).
Witteveldt, J. et al. CD81 is dispensable for hepatitis C virus cell-to-cell transmission in hepatoma cells. J. Gen. Virol. 90, 48–58 (2009).
Diamond, D.L. et al. Temporal proteome and lipidome profiles reveal HCV-associated reprogramming of hepatocellular metabolism and bioenergetics. PLoS Pathog. 6, e1000719 (2010).
Anderson, P. & Kedersha, N. RNA granules. J. Cell Biol. 172, 803–808 (2006).
Schutz, S. & Sarnow, P. How viruses avoid stress. Cell Host Microbe 2, 284–285 (2007).
Tourriere, H. et al. The RasGAP-associated endoribonuclease G3BP assembles stress granules. J. Cell Biol. 160, 823–831 (2003).
Khetani, S.R. & Bhatia, S.N. Microscale culture of human liver cells for drug development. Nat. Biotechnol. 26, 120–126 (2008).
Bhatia, S.N., Balis, U.J., Yarmush, M.L. & Toner, M. Effect of cell-cell interactions in preservation of cellular phenotype: cocultivation of hepatocytes and nonparenchymal cells. FASEB J. 13, 1883–1900 (1999).
Ploss, A. et al. Persistent hepatitis C virus infection in microscale primary human hepatocyte cultures. Proc. Natl. Acad. Sci. USA (in the press).
Wollman, R. & Stuurman, N. High throughput microscopy: from raw images to discoveries. J. Cell Sci. 120, 3715–3722 (2007).
Pietschmann, T. et al. Construction and characterization of infectious intragenotypic and intergenotypic hepatitis C virus chimeras. Proc. Natl. Acad. Sci. USA 103, 7408–7413 (2006).
Lindenbach, B.D. et al. Complete replication of hepatitis C virus in cell culture. Science 309, 623–626 (2005).
Lindenbach, B.D. & Rice, C.M. Trans-complementation of yellow fever virus NS1 reveals a role in early RNA replication. J. Virol. 71, 9608–9617 (1997).
Petrakova, O. et al. Noncytopathic replication of Venezuelan equine encephalitis virus and eastern equine encephalitis virus replicons in mammalian cells. J. Virol. 79, 7597–7608 (2005).
Zennou, V. et al. HIV-1 genome nuclear import is mediated by a central DNA flap. Cell 101, 173–185 (2000).
Acknowledgements
We acknowledge the expert support of The Rockefeller University Bioimaging Core Facility, with special thanks to A. North, S. Galdeen and S. Bhuvanendran. We thank The Rockefeller University Flow Cytometry Resource Center, supported by the Empire State Stem Cell Fund through NY State Department of Health (NYSDOH) contract no. C023046; opinions expressed here are solely those of the authors and do not necessarily reflect those of the Empire State Stem Cell Fund, the NYSDOH, or the State of NY. We are grateful to C. Stoyanov (The Rockefeller University) for YF17D(5′C25Venus2AUbi), J. Tazi for G3BP (Institut de Génétique Moléculaire de Montpellier) and I. Frolov (UTMB) for Venezuelan equine encephalitis virus-EGFP. We thank M. Holz, A. Forest, M. Panis and A. Webson for laboratory support and C. Murray for critical reading of the manuscript. This work was supported by Public Health Service grants R01 AI075099 (C.M.R.) and R01 DK56966 (S.N.B.). This work was also funded by the Office of the Director/National Institutes of Health (NIH) through the NIH Roadmap for Medical Research, Grant 1 R01 DK085713-01 (C.M.R. and S.N.B.). Information on this Roadmap Transformative R01 Program can be found at http://grants.nih.gov/grants/guide/rfa-files/RFA-RM-08-029.html. Additional funding was provided by the Greenberg Medical Research Institute and the Starr Foundation (C.M.R.). S.N.B. is an Howard Hughes Medical Investigator investigator. C.T.J. was supported by National Research Service Award DK081193; M.T.C. was supported by a Women & Science Fellowship; L.M.J.L. is supported by a Natural Sciences and Engineering Research Council of Canada fellowship. A.P. is a recipient of a Kimberly Lawrence-Netter cancer research discovery fund award.
Author information
Authors and Affiliations
Contributions
C.T.J. and C.M.R. designed the project, analyzed results and wrote the manuscript. C.T.J., M.T.C., L.M.J.L., A.J.S., S.R.K., T.S.O., A.P. and J.W.S. performed the experimental work. S.R.K., J.W.S., T.S.O., M.R.M. and S.N.B. contributed reagents and technical expertise.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following conflicts of interest, which are managed under University policy: C.M.R. has equity in Apath, LLC, which holds commercial licenses for the Huh-7.5 cell line and the HCVcc cell culture system. S.R.K and S.N.B have equity in Hepregen Corporation, which holds commercial licenses from the Massachusetts Institute of Technology for micropatterned co-cultures (MPCCs).
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–3 (PDF 730 kb)
Supplementary Video 1a
Supplementary Video 1. Time-lapse live cell imaging of HCVcc infection. (MOV 834 kb)
Supplementary Video 1b
Supplementary Video 1. Time-lapse live cell imaging of HCVcc infection. (MOV 1057 kb)
Supplementary Video 1c
Supplementary Video 1. Time-lapse live cell imaging of HCVcc infection. (MOV 510 kb)
Supplementary Video 1d
Supplementary Video 1. Time-lapse live cell imaging of HCVcc infection. (MOV 629 kb)
Supplementary Video 2a
Supplementary Video 2. Time-lapse live cell imaging of HCV-induced stress response. (MOV 1837 kb)
Supplementary Video 2b
Supplementary Video 2. Time-lapse live cell imaging of HCV-induced stress response. (MOV 3523 kb)
Supplementary Video 2c
Supplementary Video 2. Time-lapse live cell imaging of HCV-induced stress response. (MOV 4147 kb)
Rights and permissions
About this article
Cite this article
Jones, C., Catanese, M., Law, L. et al. Real-time imaging of hepatitis C virus infection using a fluorescent cell-based reporter system. Nat Biotechnol 28, 167–171 (2010). https://doi.org/10.1038/nbt.1604
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1604