Abstract
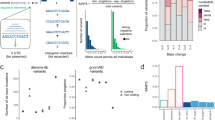
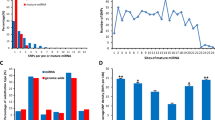
Genetic changes that help explain the differences between two individuals might create or disrupt sites complementary to microRNAs (miRNAs)1,2, but the extent to which such polymorphic sites influence miRNA-mediated repression is unknown. Here, we describe a method to measure mRNA allelic imbalances associated with a regulatory site found in mRNA transcribed from one allele but not found in that transcribed from the other. Applying this method, called allelic imbalance sequencing, to sites for three miRNAs (miR-1, miR-133 and miR-122) provided quantitative measurements of repression in vivo without altering either the miRNAs or their targets. A substantial fraction of polymorphic sites mediated repression in tissues that expressed the cognate miRNA, and downregulation was correlated with site type and site context. Extrapolating these results to the other broadly conserved miRNAs suggests that when comparing two mouse strains (or two human individuals), polymorphic miRNA sites cause expression of many genes (often hundreds) to differ.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Clop, A. et al. A mutation creating a potential illegitimate microRNA target site in the myostatin gene affects muscularity in sheep. Nat. Genet. 38, 813–818 (2006).
Sethupathy, P. & Collins, F.S. MicroRNA target site polymorphisms and human disease. Trends Genet. 24, 489–497 (2008).
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Brennecke, J., Stark, A., Russell, R.B. & Cohen, S.M. Principles of microRNA-target recognition. PLoS Biol. 3, e85 (2005).
Krek, A. et al. Combinatorial microRNA target predictions. Nat. Genet. 37, 495–500 (2005).
Lagos-Quintana, M. et al. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12, 735–739 (2002).
Frazer, K.A. et al. A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature 448, 1050–1053 (2007).
Sherry, S.T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Lim, L.P. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433, 769–773 (2005).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Chen, K. & Rajewsky, N. Natural selection on human microRNA binding sites inferred from SNP data. Nat. Genet. 38, 1452–1456 (2006).
Farh, K.K. et al. The widespread impact of mammalian MicroRNAs on mRNA repression and evolution. Science 310, 1817–1821 (2005).
Stark, A., Brennecke, J., Bushati, N., Russell, R.B. & Cohen, S.M. Animal MicroRNAs confer robustness to gene expression and have a significant impact on 3′UTR evolution. Cell 123, 1133–1146 (2005).
Nielsen, C.B. et al. Determinants of targeting by endogenous and exogenous microRNAs and siRNAs. RNA 13, 1894–1910 (2007).
Baek, D. et al. The impact of microRNAs on protein output. Nature 455, 64–71 (2008).
Selbach, M. et al. Widespread changes in protein synthesis induced by microRNAs. Nature 455, 58–63 (2008).
Friedman, R.C., Farh, K.K., Burge, C.B. & Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Cookson, W., Liang, L., Abecasis, G., Moffatt, M. & Lathrop, M. Mapping complex disease traits with global gene expression. Nat. Rev. Genet. 10, 184–194 (2009).
Schadt, E.E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Lo, H.S. et al. Allelic variation in gene expression is common in the human genome. Genome Res. 13, 1855–1862 (2003).
Gimelbrant, A., Hutchinson, J.N., Thompson, B.R. & Chess, A. Widespread monoallelic expression on human autosomes. Science 318, 1136–1140 (2007).
Tan, A.C. et al. Allele-specific expression in the germline of patients with familial pancreatic cancer: an unbiased approach to cancer gene discovery. Cancer Biol. Ther. 7, 135–144 (2008).
Serre, D. et al. Differential allelic expression in the human genome: a robust approach to identify genetic and epigenetic cis-acting mechanisms regulating gene expression. PLoS Genet. 4, e1000006 (2008).
International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Plagnol, V. et al. Extreme clonality in lymphoblastoid cell lines with implications for allele specific expression analyses. PLoS One 3, e2966 (2008).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007).
Su, A.I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
Acknowledgements
We thank Tim Harkins and the 454 Sequencing Facility for high-throughput sequencing, members of the laboratory for helpful comments on this manuscript, and T. DiCesare for illustration. Supported by a grant from the National Institutes of Health. D.P.B. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.K. performed all experiments and analyses. Both authors designed the experiments and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 and Supplementary Discussion (PDF 194 kb)
Supplementary Table 1
Data and information on the SNPs in 3′UTRs that create/disrupt miRNA target sites. (XLS 66 kb)
Supplementary Table 2
Data and information on the SNPs in ORFs. (XLS 23 kb)
Rights and permissions
About this article
Cite this article
Kim, J., Bartel, D. Allelic imbalance sequencing reveals that single-nucleotide polymorphisms frequently alter microRNA-directed repression. Nat Biotechnol 27, 472–477 (2009). https://doi.org/10.1038/nbt.1540
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1540
This article is cited by
-
Muscle allele-specific expression QTLs may affect meat quality traits in Bos indicus
Scientific Reports (2021)
-
Human IL12RB1 expression is allele-biased and produces a novel IL12 response regulator
Genes & Immunity (2019)
-
Vasodilator-stimulated phosphoprotein (VASP), a novel target of miR-4455, promotes gastric cancer cell proliferation, migration, and invasion, through activating the PI3K/AKT signaling pathway
Cancer Cell International (2018)
-
Heterogeneity and Individuality: microRNAs in Mental Disorders
Journal of Neural Transmission (2015)
-
RNA sequencing reveals small RNAs differentially expressed between incipient Japanese threespine sticklebacks
BMC Genomics (2013)