Abstract
The organellar two-pore channel (TPC) functions as a homodimer, in which each subunit contains two homologous Shaker-like six-transmembrane (6-TM)-domain repeats1. TPCs belong to the voltage-gated ion channel superfamily2 and are ubiquitously expressed in animals and plants3,4. Mammalian TPC1 and TPC2 are localized at the endolysosomal membrane, and have critical roles in regulating the physiological functions of these acidic organelles5,6,7. Here we present electron cryo-microscopy structures of mouse TPC1 (MmTPC1)—a voltage-dependent, phosphatidylinositol 3,5-bisphosphate (PtdIns(3,5)P2)-activated Na+-selective channel—in both the apo closed state and ligand-bound open state. Combined with functional analysis, these structures provide comprehensive structural insights into the selectivity and gating mechanisms of mammalian TPC channels. The channel has a coin-slot-shaped ion pathway in the filter that defines the selectivity of mammalian TPCs. Only the voltage-sensing domain from the second 6-TM domain confers voltage dependence on MmTPC1. Endolysosome-specific PtdIns(3,5)P2 binds to the first 6-TM domain and activates the channel under conditions of depolarizing membrane potential. Structural comparisons between the apo and PtdIns(3,5)P2-bound structures show the interplay between voltage and ligand in channel activation. These MmTPC1 structures reveal lipid binding and regulation in a 6-TM voltage-gated channel, which is of interest in light of the emerging recognition of the importance of phosphoinositide regulation of ion channels.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rahman, T. et al. Two-pore channels provide insight into the evolution of voltage-gated Ca2+ and Na+ channels. Sci. Signal. 7, ra109 (2014)
Yu, F. H. & Catterall, W. A. The VGL-chanome: a protein superfamily specialized for electrical signaling and ionic homeostasis. Sci. STKE 2004, re15 (2004). 10.1126/stke.2532004re15
Ishibashi, K., Suzuki, M. & Imai, M. Molecular cloning of a novel form (two-repeat) protein related to voltage-gated sodium and calcium channels. Biochem. Biophys. Res. Commun. 270, 370–376 (2000)
Furuichi, T., Cunningham, K. W. & Muto, S. A putative two pore channel AtTPC1 mediates Ca2+ flux in Arabidopsis leaf cells. Plant Cell Physiol. 42, 900–905 (2001)
Patel, S. Function and dysfunction of two-pore channels. Sci. Signal. 8, re7 (2015)
Grimm, C., Chen, C. C., Wahl-Schott, C. & Biel, M. Two-pore channels: catalyzers of endolysosomal transport and function. Front. Pharmacol. 8, 45 (2017)
Xu, H. & Ren, D. Lysosomal physiology. Annu. Rev. Physiol. 77, 57–80 (2015)
Ambrosio, A. L., Boyle, J. A., Aradi, A. E., Christian, K. A. & Di Pietro, S. M. TPC2 controls pigmentation by regulating melanosome pH and size. Proc. Natl Acad. Sci. USA 113, 5622–5627 (2016)
Bellono, N. W., Escobar, I. E. & Oancea, E. A melanosomal two-pore sodium channel regulates pigmentation. Sci. Rep. 6, 26570 (2016)
Sulem, P. et al. Two newly identified genetic determinants of pigmentation in Europeans. Nat. Genet. 40, 835–837 (2008)
Fernández, B. et al. Iron overload causes endolysosomal deficits modulated by NAADP-regulated 2-pore channels and RAB7A. Autophagy 12, 1487–1506 (2016)
Pereira, G. J. et al. Nicotinic acid adenine dinucleotide phosphate (NAADP) regulates autophagy in cultured astrocytes. J. Biol. Chem. 286, 27875–27881 (2011)
Favia, A. et al. VEGF-induced neoangiogenesis is mediated by NAADP and two-pore channel-2-dependent Ca2+ signaling. Proc. Natl Acad. Sci. USA 111, E4706–E4715 (2014)
Arndt, L. et al. NAADP and the two-pore channel protein 1 participate in the acrosome reaction in mammalian spermatozoa. Mol. Biol. Cell 25, 948–964 (2014)
Cang, C. et al. mTOR regulates lysosomal ATP-sensitive two-pore Na+ channels to adapt to metabolic state. Cell 152, 778–790 (2013)
Grimm, C. et al. High susceptibility to fatty liver disease in two-pore channel 2-deficient mice. Nat. Commun. 5, 4699 (2014)
Sakurai, Y. et al. Two-pore channels control Ebola virus host cell entry and are drug targets for disease treatment. Science 347, 995–998 (2015)
Brailoiu, E. et al. Essential requirement for two-pore channel 1 in NAADP-mediated calcium signaling. J. Cell Biol. 186, 201–209 (2009)
Calcraft, P. J. et al. NAADP mobilizes calcium from acidic organelles through two-pore channels. Nature 459, 596–600 (2009)
Zong, X. et al. The two-pore channel TPCN2 mediates NAADP-dependent Ca2+-release from lysosomal stores. Pflugers Arch. 458, 891–899 (2009)
Wang, X. et al. TPC proteins are phosphoinositide-activated sodium-selective ion channels in endosomes and lysosomes. Cell 151, 372–383 (2012)
Jha, A., Ahuja, M., Patel, S., Brailoiu, E. & Muallem, S. Convergent regulation of the lysosomal two-pore channel-2 by Mg2+, NAADP, PI(3,5)P2 and multiple protein kinases. EMBO J. 33, 501–511 (2014)
Cang, C., Bekele, B. & Ren, D. The voltage-gated sodium channel TPC1 confers endolysosomal excitability. Nat. Chem. Biol. 10, 463–469 (2014)
Rybalchenko, V. et al. Membrane potential regulates nicotinic acid adenine dinucleotide phosphate (NAADP) dependence of the pH- and Ca2+-sensitive organellar two-pore channel TPC1. J. Biol. Chem. 287, 20407–20416 (2012)
Guo, J. et al. Structure of the voltage-gated two-pore channel TPC1 from Arabidopsis thaliana. Nature 531, 196–201 (2016)
Kintzer, A. F. & Stroud, R. M. Structure, inhibition and regulation of two-pore channel TPC1 from Arabidopsis thaliana. Nature 531, 258–264 (2016)
Hedrich, R. & Marten, I. TPC1-SV channels gain shape. Mol. Plant 4, 428–441 (2011)
Guo, J., Zeng, W. & Jiang, Y. Tuning the ion selectivity of two-pore channels. Proc. Natl Acad. Sci. USA 114, 1009–1014 (2017)
Hooper, R., Churamani, D., Brailoiu, E., Taylor, C. W. & Patel, S. Membrane topology of NAADP-sensitive two-pore channels and their regulation by N-linked glycosylation. J. Biol. Chem. 286, 9141–9149 (2011)
Patel, S., Churamani, D. & Brailoiu, E. NAADP-evoked Ca2+signals through two-pore channel-1 require arginine residues in the first S4–S5 linker. Cell Calcium 68, 1–4 (2017)
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. Manipulation of subunit stoichiometry in heteromeric membrane proteins. Structure 24, 797–805 (2016)
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017)
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016)
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014)
DiMaio, F., Zhang, J., Chiu, W. & Baker, D. Cryo-EM model validation using independent map reconstructions. Protein Sci. 22, 865–868 (2013)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Schrödinger, L. The PyMOL Molecular Graphics System, Version 1.8. (Schrödinger, 2015)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013)
Brailoiu, E. et al. An NAADP-gated two-pore channel targeted to the plasma membrane uncouples triggering from amplifying Ca2+ signals. J. Biol. Chem. 285, 38511–38516 (2010)
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006)
Acknowledgements
We thank N. Nguyen for manuscript preparation and M. X. Zhu for providing clones of animal TPC genes. Single particle cryo-EM data were collected at the University of Texas Southwestern Medical Center (UTSW) Cryo-Electron Microscopy Facility that is funded by the CPRIT Core Facility Support Award RP170644. We thank D. Nicastro and Z. Chen for facility access and data acquisition. Negatively stained sample screening was performed at UTSW Electron Microscopy core. This work was supported in part by the Howard Hughes Medical Institute (Y.J.) and by grants from the National Institute of Health (GM079179 to Y.J.) and the Welch Foundation (Grant I-1578 to Y.J.). X.B. is supported by the Cancer Prevention and Research Initiative of Texas and Virginia Murchison Linthicum Scholar in Medical Research fund.
Author information
Authors and Affiliations
Contributions
J.S., J.G. and Q.C. prepared the samples; J.S., J.G., Q.C. and X.B. performed data acquisition, image processing and structure determination; W.Z. performed electrophysiology; Y.J. supervised the project and revised the manuscript; all authors participated in research design, data analysis and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of MmTPC1, HsTPC1, AtTPC1, MmTPC2 and HsTPC2.
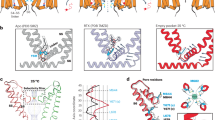
Secondary structure assignments are based on the structure of PtdIns(3,5)P2-bound MmTPC1. Red dots mark the ligand-binding residues; black dots mark the S4 arginine residues and residues at the gating-charge transfer centre; cyan dots mark the key S6 gating residues; green dots mark the residues predicted to participate in Ca2+ coordination in EF-hand domains of AtTPC1. MmTPC1 and AtTPC1 share about 25% sequence identity.
Extended Data Figure 2 Gating and selectivity of MmTPC1.
a, Sample traces and current density (current/capacitance) of wild-type MmTPC1 and the L11A/I12A mutant of MmTPC1, recorded in the whole-cell configuration with 100 μM PtdIns(3,5)P2 in the pipette (cytosolic). The experiments were repeated five times independently with similar results. Data points for current density are mean ± s.e.m. (n = 5 independent experiments). The L11A/I12A mutant elicited much larger whole-cell currents and was therefore used as the wild-type channel in all recordings. The extracellular side of MmTPC1 in plasma membrane is equivalent to the luminal side of MmTPC1 in lysosomes. b, Sample traces of PtdIns(3,5)P2-dependent voltage activation of MmTPC1. Whole-cell currents were recorded with varying PtdIns(3,5)P2 concentrations in the pipette (cytosolic) at pH 7.4. The experiments were repeated five times independently with similar results. c, G/Gmax–V curves of MmTPC1 at various PtdIns(3,5)P2 concentrations. Boltzmann fit yields V1/2 (mV) = 21.6 ± 1.2, 15.2 ± 1.0, 16.1 ± 0.9 and −2.0 ± 1.0, and Z = 0.78 ± 0.04, 0.82 ± 0.03, 0.89 ± 0.02 and 0.84 ± 0.05 for voltage activation in 0.05, 0.2, 2.0 and 10 μM cytosolic PtdIns(3,5)P2, respectively, in which V1/2 is the membrane potential for half maximum activation and Z is apparent valence. All data points are mean ± s.e.m. (n = 5 independent experiments). d, Luminal pH modulates the voltage activation of MmTPC1. Whole-cell currents of MmTPC1 recorded in the presence of 2 μM cytosolic PtdIns(3,5)P2 with a varying luminal (bath) pH of 7.4, 6.0 or 4.6. Sample traces were obtained from the same patch. The experiments were repeated five times independently with similar results. e, G/Gmax–V curves of MmTPC1 at various luminal pH values. Boltzmann fit yields V1/2 = 16.2 ± 0.8 mV, Z = 0.91 ± 0.02 at pH 7.4, V1/2 = 38.2 ± 1.2 mV, Z = 0.95 ± 0.02 at pH 6.0. All data points were normalized against Gmax obtained at 100 mV activation voltage and pH 7.4. All data points are mean ± s.e.m. (n = 5 independent experiments). f, Sample traces of whole-cell currents with 150 mM Na+ in the pipette solution, and either 150 mM Na+ or 145 mM K+ and 5 mM Na+ in the bath solution, and the I–V curves generated from the tail currents of the sample traces. g, Sample traces of whole-cell currents with 150 mM Na+ in the pipette solution and 150 mM Na+ or 100 mM Ca2+ in the bath solution, and the I–V curves generated from the tail currents of the sample traces. Data in f and g were recorded with 10 μM PtdIns(3,5)P2 in the pipette at pH 7.4 and both experiments were repeated five times independently with similar results.
Extended Data Figure 3 Structure determination of PtdIns(3,5)P2-bound MmTPC1.
a, Representative electron micrograph of PtdIns(3,5)P2-bound MmTPC1; 2,348 micrographs were used for structure determination. b, 2D class averages. c, Euler angle distribution of particles used in the final 3D reconstruction, with the heights of the cylinders corresponding to the number of particles. d, Final density maps coloured by local resolution. e, Gold-standard FSC curves of the final 3D reconstructions. f, FSC curves for cross-validation between the models and the maps. Curves for model versus summed map in black (sum), for model versus half map in blue (work) and for model versus half map not used for refinement in red (free). g, Flowchart of electron microscopy data processing for PtdIns(3,5)P2-bound MmTPC1 particles.
Extended Data Figure 4 Structure determination of apo MmTPC1.
a, Representative electron micrograph of apo MmTPC1; 2,937 micrographs were used for structure determination. b, 2D class averages. c, Euler angle distribution of particles used in the final 3D reconstruction, with the heights of the cylinders corresponding to the number of particles. d, Final density maps coloured by local resolution. e, Gold-standard FSC curves of the final 3D reconstructions. f, FSC curves for cross-validation between the models and the maps. Curves for model versus summed map in black (sum), for model versus half map in blue (work) and for model versus half map not used for refinement in red (free). g, Flowchart of electron microscopy data processing for apo MmTPC1 particles.
Extended Data Figure 5 Sample electron microscopy density maps (blue mesh) for MmTPC1.
a-d, Sample electron microscopy density maps for various parts of PtdIns(3,5)P2-bound MmTPC1: IS1–IS6 and filter I (a), IIS1–IS6 and filter II (b), NAGs of Asn600 and Asn612 (c), and PtdIns(3,5)P2-binding site (d). The maps are low-pass filtered to 3.2 Å and sharpened with a temperature factor of −105 Å2. e, f, Sample electron microscopy density maps for the key parts of apo MmTPC1: ligand binding site (e) and S6 helices (f). The maps are low-pass filtered to 3.4 Å and sharpened with a temperature factor of −98.5 Å2. Residues discussed in main text are labelled in red.
Extended Data Figure 6 Structure comparison between MmTPC1 and AtTPC1.
a, Superposition of the overall structures of MmTPC1 (blue) and AtTPC1(salmon). b, Superposition of the pore regions. c, Superposition of VSD1 domains. The comparison of the VSD2 domains is shown in Fig. 3f. d, Superposition of cytosolic soluble domains.
Extended Data Figure 7 Sample traces of whole-cell currents for Asn648Ala and Asn649Ala filter mutants.
The pipette solution contained 150 mM Na+ and the bath solution contained 150 mM Na+, or 145 mM K+ and 5 mM Na+. The tail currents were used to generate the I–V curves shown in Fig. 2g. The experiments were repeated five times independently with similar results.
Extended Data Figure 8 Voltage-sensing domains.
a, Superimposition of MmTPC1 VSD1 structures in the PtdIns(3,5)P2-bound (green) and apo (pink) states with S1 helices removed for clarity. The MmTPC1 VSD1 lacks some key features of canonical voltage sensors: the conserved aromatic residue on S2 and acidic residue on S3 that form the gating-charge transfer centre become Val152 and Lys177, respectively, in MmTPC1; the conserved basic residue at the R5 position becomes Phe209 in MmTPC1; no arginine from IS4 is positioned in the gating-charge transfer centre. b, Superimposition of MmTPC1 VSD2 structures in the PtdIns(3,5)P2-bound (orange) and apo (cyan) states. c, Sample traces of voltage activation of MmTPC1 and its IS4 arginine mutations, recorded in whole-cell configuration with 2 μM PtdIns(3,5)P2 in the pipette. Peak tail currents were used to generate the G/Gmax–V curves shown in Fig. 3c. The experiments were repeated five times independently with similar results. d, Sample traces of voltage activation of Arg546Gln mutation, recorded in whole-cell configuration with 2 μM and 100 μM PtdIns(3,5)P2 in the pipette. The experiments were repeated five times independently with similar results.
Extended Data Figure 9 PtdIns(3,5)P2-binding in MmTPC1.
a, Model of bound PtdIns(3,5)P2 (left) and its electron microscopy density (right). Density of two other membrane lipid molecules (blue mesh in left panel) was also observed near PtdIns(3,5)P2 in the structure. b, Current density of mutations at the PtdIns(3,5)P2-binding site measured at −100 mV in whole-cell recordings. All mutants were generated on the background of Arg540Gln mutant, which was used as control. All data points are mean ± s.e.m. with the number of independent experiments for each mutant shown in parentheses. c, Sample I–V curves of Arg540Gln mutant recorded in excised patches with varying concentrations of PtdIns(3,5)P2 in the bath (cytosolic). The experiments were repeated five times independently with similar results. Currents at −100 mV were used to generate the concentration-dependent PtdIns(3,5)P2 activation curve shown in Fig. 4c. Imax is the current recorded at −100 mV with 10 μM PtdIns(3,5)P2 in the bath. d, Structural comparison at the ligand-binding site between the PtdIns(3,5)P2-bound (green) and apo (salmon) states.
Supplementary information
Rights and permissions
About this article
Cite this article
She, J., Guo, J., Chen, Q. et al. Structural insights into the voltage and phospholipid activation of the mammalian TPC1 channel. Nature 556, 130–134 (2018). https://doi.org/10.1038/nature26139
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature26139
This article is cited by
-
Unexpected inhibition of the lipid kinase PIKfyve reveals an epistatic role for p38 MAPKs in endolysosomal fission and volume control
Cell Death & Disease (2024)
-
Two-pore channel blockade by phosphoinositide kinase inhibitors YM201636 and PI-103 determined by a histidine residue near pore-entrance
Communications Biology (2022)
-
Lsm12 is an NAADP receptor and a two-pore channel regulatory protein required for calcium mobilization from acidic organelles
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.