Abstract
Huntingtin (HTT) is a large (348 kDa) protein that is essential for embryonic development and is involved in diverse cellular activities such as vesicular transport, endocytosis, autophagy and the regulation of transcription1,2. Although an integrative understanding of the biological functions of HTT is lacking, the large number of identified HTT interactors suggests that it serves as a protein–protein interaction hub1,3,4. Furthermore, Huntington’s disease is caused by a mutation in the HTT gene, resulting in a pathogenic expansion of a polyglutamine repeat at the amino terminus of HTT5,6. However, only limited structural information regarding HTT is currently available. Here we use cryo-electron microscopy to determine the structure of full-length human HTT in a complex with HTT-associated protein 40 (HAP40; encoded by three F8A genes in humans)7 to an overall resolution of 4 Å. HTT is largely α-helical and consists of three major domains. The amino- and carboxy-terminal domains contain multiple HEAT (huntingtin, elongation factor 3, protein phosphatase 2A and lipid kinase TOR) repeats arranged in a solenoid fashion. These domains are connected by a smaller bridge domain containing different types of tandem repeats. HAP40 is also largely α-helical and has a tetratricopeptide repeat-like organization. HAP40 binds in a cleft and contacts the three HTT domains by hydrophobic and electrostatic interactions, thereby stabilizing the conformation of HTT. These data rationalize previous biochemical results and pave the way for improved understanding of the diverse cellular functions of HTT.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Saudou, F. & Humbert, S. The biology of huntingtin. Neuron 89, 910–926 (2016)
Zuccato, C. & Cattaneo, E. in Huntington’s Disease 4th edn (eds Bates, G. et al.) Ch. 11 (Oxford Univ. Press, 2014)
Kaltenbach, L. S. et al. Huntingtin interacting proteins are genetic modifiers of neurodegeneration. PLoS Genet. 3, e82 (2007)
Shirasaki, D. I. et al. Network organization of the huntingtin proteomic interactome in mammalian brain. Neuron 75, 41–57 (2012)
MacDonald, M. E. et al. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983 (1993)
Finkbeiner, S. Huntington’s disease. Cold Spring Harb. Perspect. Biol. 3, a007476 (2011)
Peters, M. F. & Ross, C. A. Isolation of a 40-kDa Huntingtin-associated protein. J. Biol. Chem. 276, 3188–3194 (2001)
Palidwor, G. A. et al. Detection of alpha-rod protein repeats using a neural network and application to huntingtin. PLoS Comput. Biol. 5, e1000304 (2009)
Vijayvargia, R. et al. Huntingtin’s spherical solenoid structure enables polyglutamine tract-dependent modulation of its structure and function. eLife 5, e11184 (2016)
Andrade, M. A. & Bork, P. HEAT repeats in the Huntington’s disease protein. Nat. Genet. 11, 115–116 (1995)
Takano, H. & Gusella, J. F. The predominantly HEAT-like motif structure of huntingtin and its association and coincident nuclear entry with dorsal, an NF-kB/Rel/dorsal family transcription factor. BMC Neurosci. 3, 15 (2002)
Tartari, M. et al. Phylogenetic comparison of huntingtin homologues reveals the appearance of a primitive polyQ in sea urchin. Mol. Biol. Evol. 25, 330–338 (2008)
Seong, I. S. et al. Huntingtin facilitates polycomb repressive complex 2. Hum. Mol. Genet. 19, 573–583 (2010)
Wetzel, R. & Mishra, R. in Huntington’s Disease 4th edn (eds Bates, G. et al.) Ch. 12 (Oxford Univ. Press, 2014)
Li, W., Serpell, L. C., Carter, W. J., Rubinsztein, D. C. & Huntington, J. A. Expression and characterization of full-length human huntingtin, an elongated HEAT repeat protein. J. Biol. Chem. 281, 15916–15922 (2006)
Pal, A., Severin, F., Lommer, B., Shevchenko, A. & Zerial, M. Huntingtin–HAP40 complex is a novel Rab5 effector that regulates early endosome motility and is up-regulated in Huntington’s disease. J. Cell Biol. 172, 605–618 (2006)
Huang, B. et al. Scalable production in human cells and biochemical characterization of full-length normal and mutant huntingtin. PLoS ONE 10, e0121055 (2015)
Chari, A. et al. ProteoPlex: stability optimization of macromolecular complexes by sparse-matrix screening of chemical space. Nat. Methods 12, 859–865 (2015)
Buchan, D. W., Minneci, F., Nugent, T. C., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED protein analysis workbench. Nucleic Acids Res. 41, W349–W357 (2013)
Arndt, J. R., Chaibva, M. & Legleiter, J. The emerging role of the first 17 amino acids of huntingtin in Huntington’s disease. Biomol. Concepts 6, 33–46 (2015)
Yanai, A. et al. Palmitoylation of huntingtin by HIP14 is essential for its trafficking and function. Nat. Neurosci. 9, 824–831 (2006)
Kegel, K. B. et al. Huntingtin associates with acidic phospholipids at the plasma membrane. J. Biol. Chem. 280, 36464–36473 (2005)
Wellington, C. L. et al. Caspase cleavage of gene products associated with triplet expansion disorders generates truncated fragments containing the polyglutamine tract. J. Biol. Chem. 273, 9158–9167 (1998)
El-Daher, M. T. et al. Huntingtin proteolysis releases non-polyQ fragments that cause toxicity through dynamin 1 dysregulation. EMBO J. 34, 2255–2271 (2015)
Luo, S., Vacher, C., Davies, J. E. & Rubinsztein, D. C. Cdk5 phosphorylation of huntingtin reduces its cleavage by caspases: implications for mutant huntingtin toxicity. J. Cell Biol. 169, 647–656 (2005)
Schilling, B. et al. Huntingtin phosphorylation sites mapped by mass spectrometry. Modulation of cleavage and toxicity. J. Biol. Chem. 281, 23686–23697 (2006)
Ratovitski, T. et al. Post-translational modifications (PTMs), identified on endogenous huntingtin, cluster within proteolytic domains between HEAT repeats. J. Proteome Res. 16, 2692–2708 (2017)
Ratovitski, T. et al. Mutant huntingtin N-terminal fragments of specific size mediate aggregation and toxicity in neuronal cells. J. Biol. Chem. 284, 10855–10867 (2009)
Gusella, J. F. & MacDonald, M. E. Huntingtin: a single bait hooks many species. Curr. Opin. Neurobiol. 8, 425–430 (1998)
Vedadi, M. et al. Chemical screening methods to identify ligands that promote protein stability, protein crystallization, and structure determination. Proc. Natl Acad. Sci. USA 103, 15835–15840 (2006)
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005)
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015)
Moriya, T. et al. High-resolution single particle analysis from electron cryo-microscopy images using SPHIRE. J. Vis. Exp. 123, e55448 (2017)
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
The PyMOL Molecular Graphics System v.1.8 (Schrödinger, 2015)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Acknowledgements
We thank J. Plitzko for electron microscopy support, F. Beck for help with image processing, E. Conti, H. Kiefer, B. Landwehrmeyer, K. Lindenberg, P. Mittl and L. Toledo-Sherman for discussions and A. Bracher, M. Hipp and E. Sakata for the critical reading of the manuscript. This work has been funded by the CHDI Foundation, the German Federal Ministry of Education and Research (FTLDc 01GI1007A), the German Research Foundation (SFB1279) and the European Commission (grant FP7 GA ERC-2012-SyG_318987–ToPAG). Q.G. is the recipient of postdoctoral fellowships from EMBO (EMBO ALTF 73-2015) and the Alexander von Humboldt Foundation.
Author information
Authors and Affiliations
Contributions
Q.G., B.H., P.O., A.P., W.B., R.F.-B. and S.K. designed experiments. F.M., M.M. and A.P. performed differential scanning fluorimetry studies. P.O. performed mass spectrometry analyses. B.H. prepared the HTT–HAP40 samples for cryo-EM and performed the majority of the biochemical work together with T.E. M.S. performed the experiments with truncated HAP40 constructs. Q.G. performed the majority of the cryo-EM work. Q.G. and G.P. optimized sample conditions for cryo-EM. J.C. built the atomic model. Q.G., J.C. and R.F.-B. analysed the structure. Q.G., B.H., J.C., M.S., P.O., M.O., A.P., R.F.-B. and S.K. analysed the data. Q.G. and B.H. prepared the figures. Q.G., B.H., W.B., R.F.-B. and S.K. wrote the manuscript. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. Scheres and R. Wetzel for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
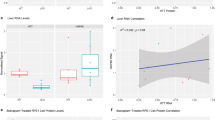
Extended Data Figure 1 Sedimentation analysis by rate-zonal ultracentrifugation.
a, Flag-tag purified HTT (top) and Strep-tag purified HTT–HAP40 complex (bottom) analysed by rate-zonal ultracentrifugation followed by SDS–PAGE and Coomassie staining. Twenty-five fractions from 5–20% sucrose gradients were collected from the bottom of the tube; fractions 1–18 are shown here. Whereas HTT alone was present in fractions 1–18, the HTT–HAP40 complex was found mainly in fractions 15–17, indicating lower conformational heterogeneity. b, Western blot analysis of fractions 10–18 of the HTT–HAP40 complex. Coomassie stainings and western blots are representative of three independent experiments with similar results. For gel source images, see Supplementary Fig. 1.
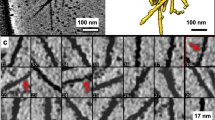
Extended Data Figure 2 Cryo-EM analysis of the HTT–HAP40 complex.
a, Representative micrograph of HTT–HAP40 complex. b, 2D class averages. c, FSC plots. Cyan, gold-standard FSC curve; orange, FSC curve calculated between the cryo-EM map and the refined atomic model. FSC cut-off values of 0.143 and 0.5 were used for the half versus half and model versus map comparisons, respectively. The initial and final numbers of micrographs and particles were 707 and 635 and 418,627 and 98,310, respectively. d, Final density map of the HTT–HAP40 complex, coloured according to local resolution. The map was low-pass filtered to 4 Å and sharpened with a B-factor of −174 Å2. e, Detail of the electron density maps (mesh) for parts of HTT (top) and HAP40 (bottom). Source Data for the FSC plots are available online.
Extended Data Figure 3 Atomic model of HTT within the HTT–HAP40 complex.
a–d, The atomic model is shown in ribbon representation with a rainbow colour code from the N terminus (blue arrowhead in d) to the C terminus (red arrowhead in a). a–d show different views of the complex, as indicated. Dashed lines mark unresolved regions.
Extended Data Figure 4 Amino acid sequences of 17QHTT and HAP40.
Structural elements of the atomic models are indicated as follows: elements not visible in the model (red boxes), unstructured regions (no box) and α-helices (yellow boxes). The sites of previously reported protease cleavage and post-translational modifications of HTT1,21,23,27 are indicated by text colour as follows: acetylation (dark blue), palmitoylation (red), phosphorylation (green) and proteolytic cleavage (cyan).
Extended Data Figure 5 PSIPRED secondary-structure predictions for HTT and HAP40.
Structural elements are indicated as follows: unstructured regions (no box), α-helices (yellow boxes) and β-sheet (grey box).
Extended Data Figure 6 Truncation analysis of HAP40 binding to HTT.
a, Schematic representation of the HAP40 constructs used in this study (all have C-terminal Strep-tags). b, HAP40 constructs were co-expressed with 17QHTT (bearing a C-terminal Flag-tag), immunoprecipitated using Strep-Tactin beads and analysed by western blot. Lanes are labelled as follows: 1, cell lysates; 2, cell lysates after incubation with Strep beads; 3, Strep-bead eluates. Note that full-length HAP40 and the construct lacking the central domain immunoprecipitate HTT, but constructs with deletion of the N- or C-terminal regions of HAP40 do not. Western blots are representative of two independent experiments with similar results. For gel source images, see Supplementary Fig. 1.
Extended Data Figure 7 Evolutionary analysis of HTT.
The human HTT model is shown in ribbon representation, coloured on the basis of sequence conservation across 16 metazoan species (Homo sapiens, Rattus norvegicus, Mus musculus, Sus scrofa, Bos taurus, Canis lupus familiaris, Monodelphis domestica, Gallus gallus, Danio rerio, Tetraodon nigroviridis, Fugu rubripes, Ciona savignyi, Ciona intestinalis, Strongylocentrotus purpuratus, Tribolium castaneum, Apis mellifera), using a previously reported sequence alignment12. The orientations of the HTT–HAP40 complex with respect to Fig. 2 are also indicated.
Extended Data Figure 8 Workflow for initial model validation for 3D reconstruction of the HTT–HAP40 complex.
A subset of particles with well-resolved 2D averages were used for initial model generation using RELION33 or SPHIRE35. The resulting models were used as reference for 3D classification of all the good particles. A featureless sphere was also used as a classification reference. Most of the particles were classified to identical structures with sufficient detail, indicating no reference bias in the reconstruction.
Supplementary information
Supplementary Figure 1
This file contains the uncropped scans with size marker indications. (PDF 419 kb)
Rights and permissions
About this article
Cite this article
Guo, Q., Bin Huang, Cheng, J. et al. The cryo-electron microscopy structure of huntingtin. Nature 555, 117–120 (2018). https://doi.org/10.1038/nature25502
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25502
This article is cited by
-
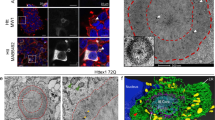
CryoET reveals organelle phenotypes in huntington disease patient iPSC-derived and mouse primary neurons
Nature Communications (2023)
-
Krankheitsmodifizierende Therapieansätze bei der Huntington-Krankheit
Der Nervenarzt (2022)
-
Proteinopathies: Deciphering Physiology and Mechanisms to Develop Effective Therapies for Neurodegenerative Diseases
Molecular Neurobiology (2022)
-
Huntingtin structure is orchestrated by HAP40 and shows a polyglutamine expansion-specific interaction with exon 1
Communications Biology (2021)
-
Mitochondrial Abnormalities and Synaptic Damage in Huntington’s Disease: a Focus on Defective Mitophagy and Mitochondria-Targeted Therapeutics
Molecular Neurobiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.