Abstract
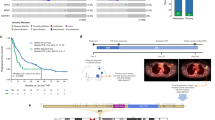
Somatic mutations of ERBB2 and ERBB3 (which encode HER2 and HER3, respectively) are found in a wide range of cancers. Preclinical modelling suggests that a subset of these mutations lead to constitutive HER2 activation, but most remain biologically uncharacterized. Here we define the biological and therapeutic importance of known oncogenic HER2 and HER3 mutations and variants of unknown biological importance by conducting a multi-histology, genomically selected, ‘basket’ trial using the pan-HER kinase inhibitor neratinib (SUMMIT; clinicaltrials.gov identifier NCT01953926). Efficacy in HER2-mutant cancers varied as a function of both tumour type and mutant allele to a degree not predicted by preclinical models, with the greatest activity seen in breast, cervical and biliary cancers and with tumours that contain kinase domain missense mutations. This study demonstrates how a molecularly driven clinical trial can be used to refine our biological understanding of both characterized and new genomic alterations with potential broad applicability for advancing the paradigm of genome-driven oncology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chmielecki, J. et al. Oncogenic alterations in ERBB2/HER2 represent potential therapeutic targets across tumors from diverse anatomic sites of origin. Oncologist 20, 7–12 (2015)
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017)
Schram, A. et al. Landscape of somatic ERBB2 mutations: Findings from AACR GENIE and comparison to ongoing ERBB2 mutant basket study. Cancer Res. 77, Abstract LB-103 (2017)
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016)
Bose, R. et al. Activating HER2 mutations in HER2 gene amplification negative breast cancer. Cancer Discov. 3, 224–237 (2013)
Kavuri, S. M. et al. HER2 activating mutations are targets for colorectal cancer treatment. Cancer Discov. 5, 832–841 (2015)
Jaiswal, B. S. et al. Oncogenic ERBB3 mutations in human cancers. Cancer Cell 23, 603–617 (2013)
Chumsri, S. et al. Prolonged response to trastuzumab in a patient with HER2-nonamplified breast cancer with elevated HER2 dimerization harboring an ERBB2 S310F mutation. J. Natl. Compr. Canc. Netw. 13, 1066–1070 (2015)
Zabransky, D. J. et al. HER2 missense mutations have distinct effects on oncogenic signaling and migration. Proc. Natl Acad. Sci. USA 112, E6205–E6214 (2015)
Hyman, D. M. et al. Vemurafenib in multiple nonmelanoma cancers with BRAF V600 mutations. N. Engl. J. Med. 373, 726–736 (2015)
Hyman, D. M. et al. AKT inhibition in solid tumors with AKT1 mutations. J. Clin. Oncol. 35, 2251–2259 (2017)
Ross, J. S. et al. A high frequency of activating extracellular domain ERBB2 (HER2) mutation in micropapillary urothelial carcinoma. Clin. Cancer Res. 20, 68–75 (2014)
Ross, J. S. et al. Relapsed classic E-cadherin (CDH1)-mutated invasive lobular breast cancer shows a high frequency of HER2 (ERBB2) gene mutations. Clin. Cancer Res. 19, 2668–2676 (2013)
Wolff, A. C. et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J. Clin. Oncol. 31, 3997–4013 (2013)
Ma, C. X. et al. Neratinib efficacy and circulating tumor DNA detection of HER2 mutations in HER2 non-amplified metastatic breast cancer. Clin. Cancer Res. 23, 5687–5695 (2017)
Yasuda, H. et al. Structural, biochemical, and clinical characterization of epidermal growth factor receptor (EGFR) exon 20 insertion mutations in lung cancer. Sci. Transl. Med. 5, 216ra177 (2013)
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015)
Kosaka, T. et al. Response heterogeneity of EGFR and HER2 exon 20 insertions to covalent EGFR and HER2 inhibitors. Cancer Res. 77, 2712–2721 (2017)
Freedman, R. A. et al. Translational Breast Cancer Research Consortium (TBCRC) 022: a phase II trial of neratinib for patients with human epidermal growth factor receptor 2-positive breast cancer and brain metastases. J. Clin. Oncol. 34, 945–952 (2016)
Shi, W. et al. Pathway level alterations rather than mutations in single genes predict response to HER2-targeted therapies in the neo-ALTTO trial. Ann. Oncol. 28, 128–135 (2017)
Loibl, S. et al. PIK3CA mutations are associated with reduced pathological complete response rates in primary HER2-positive breast cancer: pooled analysis of 967 patients from five prospective trials investigating lapatinib and trastuzumab. Ann. Oncol. 27, 1519–1525 (2016)
Baselga, J. et al. Biomarker analyses in CLEOPATRA: a phase III, placebo-controlled study of pertuzumab in human epidermal growth factor receptor 2-positive, first-line metastatic breast cancer. J. Clin. Oncol. 32, 3753–3761 (2014)
Schram, A. M., Berger, M. F. & Hyman, D. M. Precision oncology: charting a path forward to broader deployment of genomic profiling. PLoS Med. 14, e1002242 (2017)
Hyman, D. M., Taylor, B. S. & Baselga, J. Implementing genome-driven oncology. Cell 168, 584–599 (2017)
Jordan, E. J. et al. Prospective comprehensive molecular characterization of lung adenocarcinomas for efficient patient matching to approved and emerging therapies. Cancer Discov. 7, 596–609 (2017)
Baselga, J. et al. Phase II study of weekly intravenous recombinant humanized anti-p185HER2 monoclonal antibody in patients with HER2/neu-overexpressing metastatic breast cancer. J. Clin. Oncol. 14, 737–744 (1996)
Vogel, C. L. et al. Efficacy and safety of trastuzumab as a single agent in first-line treatment of HER2-overexpressing metastatic breast cancer. J. Clin. Oncol. 20, 719–726 (2002)
Slamon, D. J. et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N. Engl. J. Med. 344, 783–792 (2001)
Bang, Y. J. et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): a phase 3, open-label, randomised controlled trial. Lancet 376, 687–697 (2010)
Swain, S. M. et al. Pertuzumab, trastuzumab, and docetaxel in HER2-positive metastatic breast cancer. N. Engl. J. Med. 372, 724–734 (2015)
Blackwell, K. L. et al. Randomized study of lapatinib alone or in combination with trastuzumab in women with ErbB2-positive, trastuzumab-refractory metastatic breast cancer. J. Clin. Oncol. 28, 1124–1130 (2010)
Baselga, J. et al. Lapatinib with trastuzumab for HER2-positive early breast cancer (NeoALTTO): a randomised, open-label, multicentre, phase 3 trial. Lancet 379, 633–640 (2012)
Bertotti, A. et al. A molecularly annotated platform of patient-derived xenografts (‘xenopatients’) identifies HER2 as an effective therapeutic target in cetuximab-resistant colorectal cancer. Cancer Discov. 1, 508–523 (2011)
Sartore-Bianchi, A. et al. Dual-targeted therapy with trastuzumab and lapatinib in treatment-refractory, KRAS codon 12/13 wild-type, HER2-positive metastatic colorectal cancer (HERACLES): a proof-of-concept, multicentre, open-label, phase 2 trial. Lancet Oncol. 17, 738–746 (2016)
Kaufman, B. et al. Olaparib monotherapy in patients with advanced cancer and a germline BRCA1/2 mutation. J. Clin. Oncol. 33, 244–250 (2015)
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017)
Hyman, D. M. et al. The efficacy of larotrectinib (LOXO-101), a selective tropomyosin receptor kinase (TRK) inhibitor, in adult and pediatric TRK fusion cancers. J. Clin. Oncol. 35, abstract LBA2501 (2017)
Wahl, R. L., Jacene, H., Kasamon, Y. & Lodge, M. A. From RECIST to PERCIST: Evolving considerations for PET response criteria in solid tumors. J. Nucl. Med. 50, 122S–150S (2009)
Cheng, D. T. et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015)
Chang, M. T. et al. Accelerating discovery of functional mutant alleles in cancer. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-17-0321 (2017)
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000)
Chakravarty D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. https://doi.org/10.1200/PO.17.00011 (2017)
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016)
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012)
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015)
Niu, B. et al. MSIsensor: microsatellite instability detection using paired tumor-normal sequence data. Bioinformatics 30, 1015–1016 (2014)
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Middha et al. Reliable pan-cancer microsatellite instability assessment by using targeted next-generation sequencing data. JCO Precis. Oncol. https://doi.org/10.1200/PO.17.00084 (2017)
Zhou, X. et al. Exploring genomic alteration in pediatric cancer using ProteinPaint. Nat. Genet. 48, 4–6 (2016)
Acknowledgements
We thank patients and their families for participating in this study. Editorial support, not including writing, was provided by L. Miller. This work was funded by Puma Biotechnology, and supported by grants from the National Institutes of Health (grants P30 CA008748, P30 CA016672, P30 CA014089, R01 CA204749, R01 CA80195, T32 CA009207, 1U01 CA180964 and UL1 TR000371), the National Institutes of Health/National Cancer Institute (Breast SPORE grant P50 CA098131), Cycle for Survival, Marie-Josée and Henry R. Kravis Center for Molecular Oncology, The Cancer Prevention and Research Institute of Texas (RP1100584), the Sheikh Khalifa Bin Zayed Al Nahyan Institute for Personalized Cancer Therapy, Nellie B. Connally Breast Cancer Research Endowment, and the Breast Cancer Research Foundation.
Author information
Authors and Affiliations
Contributions
D.M.H., H.W., M.F.B., R.E.C, F.X., A.B., L.D.E., G.M., C.F., A.S.L., R.P.B., J.B. and D.B.S. designed the study and supervised the analyses. R.E.C., F.X., L.D.E., G.M., C.F., A.S.L. and R.P.B. helped to collect and monitor the clinical outcome data. D.M.H., S.A.P., J.R., C.S., G.I.S., D.J., D.I.Q., V.M., B.D., I.A.M., V.B., E.C., S.L., A.C.L., J.P.E., B.T.L., A.J.H., R.M., A.M.S., A.D., L.M.S., K.J., G.I., J.J.H., C.L.A., F.M.B., J.B. and D.B.D. enrolled patients and provided patient samples. G.U. developed the PET response criteria and performed radiographic response assessments. B.S.T., J.P., J.T., S.D.S., N.B., M.M., M.F.B., J.B. and D.B.S. performed the tumour and plasma sequencing, provided computational infrastructure, and made final variant calls. D.M.H., H.W., M.S., B.S.T., J.P., J.T., H.B., M.F.B. and D.B.S. analysed clinical and genomic data and performed the integrated efficacy analyses. F.X. performed biostatistical analyses of the clinical efficacy data. D.M.H., H.W., B.S.T., C.L.A., F.M.B. and D.B.S. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
R.E.C., F.X., L.D.E., G.M., C.F., A.S.L. and R.P.B. are employees of Puma Biotechnology. D.M.H., M.S. and J.B. receive research support from Puma Biotechnology, B.T.L. and M.S. receive research funding from Diachi, A.D. receives personal fees from Roche, and D.S. received personal fees from Loxo Oncology and Pfizer.
Additional information
Reviewer Information Nature thanks E. Mardis and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Design of SUMMIT study.
Five tumour-specific HER2 (ERBB2)-mutant cohorts were pre-specified (endometrial, gastroesophageal, ovarian, colorectal and bladder/urinary tract). In addition, a sixth ‘solid tumour (not otherwise specified, NOS)’ HER2-mutant cohort allowed for the enrolment of patients with any other cancer types. A sufficient number of patients with breast, cervical, biliary and lung cancer were enrolled in the solid tumours (NOS) cohort to permit independent efficacy analysis using the same design as the pre-specified cohorts. Patients with HER3 (ERBB3)-mutant tumours were enrolled in a HER3-specific cohort regardless of tumour type. CBR, clinical benefit rate; cfDNA, cell-free (tumour) DNA; CI, confidence interval.
Extended Data Figure 2 Distribution of HER2 and HER3 mutations positioned by their amino acid coordinates across the respective protein domains.
a, b, HER2 (a) and HER3 (b) mutations (125 and 16 mutations, respectively). Each unique mutation is represented by a circle, with the circle size and number representing the frequency, and coloured to show the mutation class as indicated in the legend. The corresponding amino acid change and common hotspot mutations (shown in pink) are labelled next to the circles.
Extended Data Figure 3 Spectrum of HER2 and HER3 mutations observed in the neratinib study versus TCGA, ICGC and other public datasets.
a, b, Distribution of HER2 (a) and HER3 (b) mutations observed across our cohort in comparison to the spectrum of HER2 and HER3 mutations (reflected lollipop) from publically available datasets (TCGA, ICGC and other published studies).
Extended Data Figure 4 Distribution and outcome of 28 HER2 exon 20 insertions.
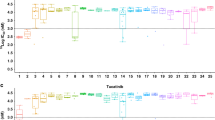
a, Percentage best change and PFS plots corresponding to each type of exon 20 insertion (colour coded by synonymous amino acid change). Three cases with no change are indicated in colour-coded circles above the x axis. b, Zoomed-in schematic of all exon 20 insertions positioned by their amino acid coordinates and frequencies. c, Five unique types of exon 20 insertions observed in the study with the resulting full amino acid sequences (insertion indicated in red).
Extended Data Figure 5 Genomic modifiers of response and outcome by treatment duration.
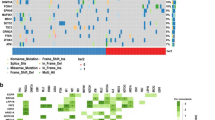
a, Cancer cell fractions with 95% confidence intervals and clonality status of all HER2 mutations in 74 patients with sufficient sequencing data ordered by increasing clinical benefit (weeks on therapy). b, Comparison of the percentage activation of known oncogenic alterations in the three pathways between the patients of clinical benefit (n = 20, biologically independent samples) and no benefit (n = 66, biologically independent samples). Nominal Fisher’s P values are shown.
Supplementary information
Supplementary Information
This file contains: 1 - list of genes covered in the MSK-IMPACT panel along with the HGNC ID, short gene description, chromosomal location, and panel version, 2 - list of all somatic mutations within the MSK-IMPACT genes for patient tumour samples with sequencing data and 3 - list of all somatic copy number alterations within the MSK-IMPACT genes for patient tumour samples with sequencing data. (PDF 745 kb)
Rights and permissions
About this article
Cite this article
Hyman, D., Piha-Paul, S., Won, H. et al. HER kinase inhibition in patients with HER2- and HER3-mutant cancers. Nature 554, 189–194 (2018). https://doi.org/10.1038/nature25475
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25475
This article is cited by
-
Biomarker discovery with quantum neural networks: a case-study in CTLA4-activation pathways
BMC Bioinformatics (2024)
-
Conformational diversity and protein–protein interfaces in drug repurposing in Ras signaling pathway
Scientific Reports (2024)
-
Management of patients with advanced-stage HER2-positive breast cancer: current evidence and future perspectives
Nature Reviews Clinical Oncology (2024)
-
Reporting on invasive lobular breast cancer in clinical trials: a systematic review
npj Breast Cancer (2024)
-
Progress in systemic therapy for advanced-stage urothelial carcinoma
Nature Reviews Clinical Oncology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.