Abstract
The poor correlation of mutational landscapes with phenotypes limits our understanding of the pathogenesis and metastasis of pancreatic ductal adenocarcinoma (PDAC). Here we show that oncogenic dosage-variation has a critical role in PDAC biology and phenotypic diversification. We find an increase in gene dosage of mutant KRAS in human PDAC precursors, which drives both early tumorigenesis and metastasis and thus rationalizes early PDAC dissemination. To overcome the limitations posed to gene dosage studies by the stromal richness of PDAC, we have developed large cell culture resources of metastatic mouse PDAC. Integration of cell culture genomes, transcriptomes and tumour phenotypes with functional studies and human data reveals additional widespread effects of oncogenic dosage variation on cell morphology and plasticity, histopathology and clinical outcome, with the highest KrasMUT levels underlying aggressive undifferentiated phenotypes. We also identify alternative oncogenic gains (Myc, Yap1 or Nfkb2), which collaborate with heterozygous KrasMUT in driving tumorigenesis, but have lower metastatic potential. Mechanistically, different oncogenic gains and dosages evolve along distinct evolutionary routes, licensed by defined allelic states and/or combinations of hallmark tumour suppressor alterations (Cdkn2a, Trp53, Tgfβ-pathway). Thus, evolutionary constraints and contingencies direct oncogenic dosage gain and variation along defined routes to drive the early progression of PDAC and shape its downstream biology. Our study uncovers universal principles of Ras-driven oncogenesis that have potential relevance beyond pancreatic cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Rahib, L. et al. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 74, 2913–2921 (2014)
Jones, S. et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008)
Campbell, P. J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010)
Biankin, A. V. et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491, 399–405 (2012)
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015)
Witkiewicz, A. K. et al. Whole-exome sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nat. Commun. 6, 6744 (2015)
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016)
Makohon-Moore, A. P. et al. Limited heterogeneity of known driver gene mutations among the metastases of individual patients with pancreatic cancer. Nat. Genet. 49, 358–366 (2017)
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001)
Schönhuber, N. et al. A next-generation dual-recombinase system for time- and host-specific targeting of pancreatic cancer. Nat. Med. 20, 1340–1347 (2014)
Pérez-Mancera, P. A. et al. The deubiquitinase USP9X suppresses pancreatic ductal adenocarcinoma. Nature 486, 266–270 (2012)
Mann, K. M. et al. Sleeping Beauty mutagenesis reveals cooperating mutations and pathways in pancreatic adenocarcinoma. Proc. Natl Acad. Sci. USA 109, 5934–5941 (2012)
Rad, R. et al. A conditional piggyBac transposition system for genetic screening in mice identifies oncogenic networks in pancreatic cancer. Nat. Genet. 47, 47–56 (2015)
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013)
Notta, F. et al. A renewed model of pancreatic cancer evolution based on genomic rearrangement patterns. Nature 538, 378–382 (2016)
Yamada, H. et al. Amplifications of both c-Ki-ras with a point mutation and c-myc in a primary pancreatic cancer and its metastatic tumors in lymph nodes. Jpn. J. Cancer Res. 77, 370–375 (1986)
Heidenblad, M. et al. Detailed genomic mapping and expression analyses of 12p amplifications in pancreatic carcinomas reveal a 3.5-Mb target region for amplification. Genes Chromosom. Cancer 34, 211–223 (2002)
Sarkisian, C. J. et al. Dose-dependent oncogene-induced senescence in vivo and its evasion during mammary tumorigenesis. Nat. Cell Biol. 9, 493–505 (2007)
Junttila, M. R. et al. Selective activation of p53-mediated tumour suppression in high-grade tumours. Nature 468, 567–571 (2010)
Feldser, D. M . et al. Stage-specific sensitivity to p53 restoration during lung cancer progression. Nature 468, 572–575 (2010)
Rad, R. et al. A genetic progression model of Braf(V600E)-induced intestinal tumorigenesis reveals targets for therapeutic intervention. Cancer Cell 24, 15–29 (2013)
Rhim, A. D. et al. EMT and dissemination precede pancreatic tumor formation. Cell 148, 349–361 (2012)
Stathis, A. & Moore, M. J. Advanced pancreatic carcinoma: current treatment and future challenges. Nat. Rev. Clin. Oncol. 7, 163–172 (2010)
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010)
Schneider, G. et al. IKKα controls p52/RelB at the skp2 gene promoter to regulate G1- to S-phase progression. EMBO J. 25, 3801–3812 (2006)
Hamidi, T. et al. Nuclear protein 1 promotes pancreatic cancer development and protects cells from stress by inhibiting apoptosis. J. Clin. Invest. 122, 2092–2103 (2012)
Redston, M. S . et al. p53 mutations in pancreatic carcinoma and evidence of common involvement of homocopolymer tracts in DNA microdeletions. Cancer Res. 54, 3025–3033 (1994)
Collisson, E. A. et al. Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nat. Med. 17, 500–503 (2011)
Moffitt, R. A. et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat. Genet. 47, 1168–1178 (2015)
Maresch, R. et al. Multiplexed pancreatic genome engineering and cancer induction by transfection-based CRISPR/Cas9 delivery in mice. Nat. Commun. 7, 10770 (2016)
Morohoshi, T., Held, G. & Klöppel, G. Exocrine pancreatic tumours and their histological classification. A study based on 167 autopsy and 97 surgical cases. Histopathology 7, 645–661 (1983)
Iacobuzio-Donahue, C. A. et al. DPC4 gene status of the primary carcinoma correlates with patterns of failure in patients with pancreatic cancer. J. Clin. Oncol. 27, 1806–1813 (2009)
Winter, J. M. et al. Absence of E-cadherin expression distinguishes noncohesive from cohesive pancreatic cancer. Clin. Cancer Res. 14, 412–418 (2008)
Hruban, R. H. et al. Pathology of genetically engineered mouse models of pancreatic exocrine cancer: consensus report and recommendations. Cancer Res. 66, 95–106 (2006)
Hingorani, S. R. et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 4, 437–450 (2003)
Nakhai, H. et al. Ptf1a is essential for the differentiation of GABAergic and glycinergic amacrine cells and horizontal cells in the mouse retina. Development 134, 1151–1160 (2007)
Aguirre, A. J. et al. Activated Kras and Ink4a/Arf deficiency cooperate to produce metastatic pancreatic ductal adenocarcinoma. Genes Dev. 17, 3112–3126 (2003)
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat. Genet. 29, 418–425 (2001)
Lee, C. L . et al. Generation of primary tumors with Flp recombinase in FRT-flanked p53 mice. Dis. Model. Mech. 5, 397–402 (2012)
Chytil, A., Magnuson, M. A., Wright, C. V. & Moses, H. L. Conditional inactivation of the TGF-β type II receptor using Cre:Lox. Genesis 32, 73–75 (2002)
Bronner, I. F., Quail, M. A., Turner, D. J. & Swerdlow, H. Improved protocols for Illumina sequencing. Curr. Protoc. Hum. Genet. 80, 18.2.1–18.2.42 (2014)
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011)
Jentsch, I., Adler, I. D., Carter, N. P. & Speicher, M. R. Karyotyping mouse chromosomes by multiplex-FISH (M-FISH). Chromosome Res. 9, 211–214 (2001)
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011)
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009)
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012)
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Parekh, S., Ziegenhain, C., Vieth, B., Enard, W. & Hellmann, I. The impact of amplification on differential expression analyses by RNA-seq. Sci. Rep. 6, 25533 (2016)
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protocols 4, 44–57 (2009)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Schaefer, C. F. et al. PID: the Pathway Interaction Database. Nucleic Acids Res. 37, D674–D679 (2009)
Du, P., Kibbe, W. A. & Lin, S. M. lumi: a pipeline for processing Illumina microarray. Bioinformatics 24, 1547–1548 (2008)
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015)
Friedrich, M. J. et al. Genome-wide transposon screening and quantitative insertion site sequencing for cancer gene discovery in mice. Nat. Protocols 12, 289–309 (2017)
Meerbrey, K. L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl Acad. Sci. USA 108, 3665–3670 (2011)
Kutner, R. H., Zhang, X. Y. & Reiser, J. Production, concentration and titration of pseudotyped HIV-1-based lentiviral vectors. Nat. Protocols 4, 495–505 (2009)
Weber, J. et al. CRISPR/Cas9 somatic multiplex-mutagenesis for high-throughput functional cancer genomics in mice. Proc. Natl Acad. Sci. USA 112, 13982–13987 (2015)
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012)
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013)
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015)
Hoadley, K. A. et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158, 929–944 (2014)
Acknowledgements
We thank the comparative experimental pathology team for discussions, and A. Selmeier, L. Dajka, O. Seelbach, P. Meyer, T. Schmidt, J. Eichinger and T. Stauber for technical assistance as well as M. Reichert for vector constructs. The work was supported by the German Cancer Consortium Joint Funding Program, the Helmholtz Gemeinschaft (PCCC Consortium), the German Research Foundation (SFB1243; A13/A14) and the European Research Council (ERC CoG number 648521).
Author information
Authors and Affiliations
Contributions
S.M., D.S., R.R. designed the study; S.M., T.E., R.M., D.S., R.R. interpreted and visualized data; T.E., S.La., M.Zw., M.B. conducted bioinformatic analyses. S.M., T.E., R.M., S.La., M.Zw., I.V. developed bioinformatic analysis strategies; S.M., R.M., M.Zu., T.K., A.S., B.S., J.G., K.Sc., Z.H., A.A., N.S., C.V., L.R. isolated mPDAC cell cultures; S.M., R.M., J.H., K.U. performed genomics with help from R.Ö.; R.Ö., C.Z. conducted RNA-seq; R.B., S.Lo., B.F., S.K., K.St., F.Y. performed cytogenetics; B.K. performed microdissection; B.K., H.-Y.Y., G.K., W.W., K.St. performed pathological assessment; C.Z., S.P., W.E., K.U., I.V. contributed analytical tools; M.Fri., O.M.D., S.E., F.C-C., J.R., M.I.S., M.Fra., J.M., G.K., R.M.S., J.C., P.L., G.V., W.W., K.St., W.E., G.S., A.B., D.S., R.R. provided resources and critical input; D.S., R.R. supervised the study; S.M., R.R. wrote the manuscript; T.E., R.M., D.S. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Mutational patterns, karyotype complexity and structural alterations in primary PDAC.
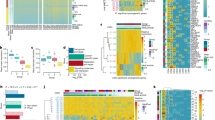
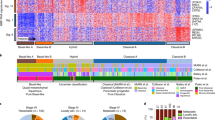
a, Single nucleotide variants (SNVs) and indels in primary PDAC cultures derived from 38 KrasG12D (PK) mice, as detected by whole-exome sequencing. Recurrently mutated genes that are frequently altered in human cancers and/or genome-wide pancreas-specific transposon screens are indicated. b, Frequency of somatic base substitutions based on trinucleotide context in mouse (n = 38 PK mice) and human PDAC (n = 51 patients, data used for analysis from ref. 6). b–f, Mutation spectra defined by trinucleotide contexts around base substitutions as detected by WES show similar patterns in PK mice (n = 38) and in relevant human pancreatic cancer cohorts. Base substitutions were extracted from BAM, VCF or MAF files from (b) ref. 6, (c) ref. 7, (d) the pancreatic adenocarcinoma cohort of The Cancer Genome Atlas, (e) ref. 46 and (f) ref. 47. Additional information about the analysis of each cohort is provided in Supplementary Table 2. g, Mutational signatures in mouse and human pancreatic cancer cohorts. Information on mutational signatures was used from ref. 47, which identified 21 mutational signatures operative in human cancer. The ‘deconstructSigs’ tool was used to determine the composition of the given set of 21 mutational signatures in each pancreatic cancer cohort. Extraction of mutational signatures strongly depends on SNV load per tumour. Owing to the low mutational burden of mPDACs from PK mice (median of 18 SNVs per tumour as detected by WES), the analyses of mutational signatures could not be performed at the level of individual tumours. We therefore investigated the contribution of each of the 21 mutational signatures to the SNV spectrum at the cohort level (see Methods). Signature 1, reflecting age-associated C>T transversions at NCG trinucleotides, was the only signature consistently identifiable in all cohorts of human and mouse pancreatic cancer. Compared with human cohorts, PK mice show C>G substitutions at GCC trinucleotides that cannot be attributed to one of 21 mutational signatures. Note that mutations at the GCC motif are not a general phenomenon of PDAC from PK mice, since only four samples predominantly contribute to this peak. h, i, Representative M-FISH karyotypes with no or few karyotypic changes are shown for a diploid (40 chromosomes) and tetraploid (81 chromosomes) mouse PDAC. Tumour 9591 shows gain of chr14. j, Representative karyotype of a complex diploid mPDAC genome with aneuploidy and translocations (46 chromosomes). Both copies of chr4 are involved in translocations: der(4)t(4;10) and der(4)t(4;16), probably affecting Cdkn2a. Further structural alterations and copy number changes are +5, der(5)t(4;5)*2, +6, +7, +8, del(9), +14, del(14), der(16)t(5;16), +17. k, Representative example of a complex tetraploid mPDAC karyotype (77 chromosomes). Structural alterations are der(1)t(1;11), dic(9;9), der(11)t(1;11) and der(14)t(14;19). Single chromosomal copy number changes are +2, −3, −9, −10, −11, −13, −14, +15 and +19. Del, deletion; der, derivative chromosome; dic, dicentric chromosome; t, translocation; ‘−’, chromosome loss; ‘+’, chromosome gain. l, (Extension to Fig. 1c.) Circos plot shows CNAs assessed by aCGH as well as translocations and ploidy states detected by M-FISH in 38 primary PDACs derived from PK mice (n = 38). CNAs for each mPDAC are displayed as log2(difference from tail control). Frequencies of translocations per chromosome are indicated in green in the inner circle of the graph. Connecting lines indicate individual translocations and involved chromosomes. On chr4, genomic alterations frequently involve Cdkn2a or Ncruc, a non-coding regulatory region upstream of Cdkn2a (27 out of 38 cancers with homozygous and 10 out of 38 with heterozygous inactivation of Cdkn2a and/or Ncruc). Only one cancer remained Cdkn2aWT. The target of copy number changes on chr6 is KrasG12D, either through arm level gain or focal amplification. In addition, primary mPDAC of PK mice exhibited recurrent genetic amplifications affecting other known oncogenes, such as Myc or Yap1, or Nfkb2, a novel oncogenic PDAC driver identified in this study (see also Fig. 2e, f and Extended Data Figure 4).
Extended Data Figure 2 Characterization of complex rearrangements in PDAC from PK mice and statistical inference of chromothripsis on the basis of WGS.
a–n, Copy-number profiles of chromosomes with complex rearrangements (defined as n ≥ 10 CNAs per chromosome) from primary mPDAC cell cultures as detected by aCGH. A total of 14 mPDACs had chromosomes with complex rearrangements. a–i, Nine primary mPDACs show copy-number patterns characterized by heterozygous deletions and oscillation of copy number around few states, indicating chromothripsis as the underlying mechanism. g, mPDAC-S821 was subjected to WGS for the inference of chromothripsis using previously established criteria14(see Fig. 1d and Extended Data Fig. 2p–w). j–m, Four primary mPDACs showed complex rearrangements with multiple copy number states on chr4, probably acquired through progressive or sequential rearrangement cycles. n, Cancer 5671 carries a complex rearrangement on chr15 characterized by oscillating copy number states and three prominent focal amplifications, of which one contained the Myc oncogene. Myc amplification is most probably the result of double minute chromosome formation during chromothriptic rearrangement of chr15. o, Comparison of age at tumour diagnosis in Cdkn2a∆HOM-deleted cancers with (n = 10) or without (n = 15) complex clustered chromosomal rearrangements (n ≥ 10 CNAs per chromosome). Complex clustered rearrangements are associated with significantly shortened time to tumour diagnosis, indicating accelerated tumour evolution through genetic crisis. Two-sided log-rank test. p, Criteria proposed in ref. 14 were tested for the inference of chromothripsis. Circos plot displays SNP ratio (inner circle, red dashed line indicating heterozygosity), CNA (outer circle, blue area indicating deletion, red amplification) and structural variations (colours as in v) as detected by WGS. Chr4 shows a complex deletion pattern and massive rearrangements associated with loss of one copy of Cdkn2a. The second copy of Cdkn2a is focally deleted. In addition, a balanced translocation of an approximately 200 kb segment from trisomic chr6 to chr4 and a far smaller segment of chr4 into chr6 was detected. The Kras locus is not directly affected by this inter-chromosomal translocation. LOH, CNAs and rearrangements are not detected on other chromosomes. q, In a chromothriptic model, DNA breakpoints tend to cluster on a chromosome. Testing against an exponential distribution (parameter λ derived from mean of observed distance between adjacent breakpoints) revealed significantly shorter distances than expected in a progressive model (n = 146 breakpoints). P < 10−12; χ2 goodness-of-fit test. r, In a progressive model of acquisition of massive rearrangements or structural variations, copy-number states tend to be more complex than in the chromothriptic model. Monte Carlo simulations were used to generate a progressive evolution model with sequential accumulation of observed rearrangements (n = 100 simulations per number of structural variations). mPDAC S821 showed fewer copy-number states on chr4 than expected in the progressive model. Mean is indicated as a black point and lines represent the 95% confidence interval. s, Chromothriptic tumours typically feature interspersed loss and retention of heterozygosity. Accordingly, there was a high overlap between deleted regions and LOH segments on chr4 (Jaccard index (J) = 0.99). t, In a chromothriptic model, DNA shattering typically occurs on a single haplotype. M-FISH showed that significant loss of chromosomal content occurred on only one copy of chr4. u, To show random chromothriptic DNA shattering and re-joining, observed segments (n = 73) were re-ordered by running Monte Carlo simulations (n = 103) generating a background probability distribution. S821 segment order lies within the chromothriptic null model. Two-sided P = 0.78. v, All four structural variation types are uniformly distributed in a chromothriptic tumour model. P = 0.43; χ2 goodness-of-fit test. w, In a chromothriptic model, paired-end connection types (as given by the structural variation type) induce an alternating sequence of DNA segment ends when ordered according to the genomic position on the original chromosome. Tendency towards this alternating 3′-to-5′ pattern of rearranged DNA segment ends (n = 146) was tested by using right-sided Wald–Wolfowitz runs test. P < 10−12. x, Mutation clusters in relation to breakpoint junctions involved in chromothripsis are shown as a rainfall plot for primary PDAC from PK mouse S821. Each dot represents a single SNV and is ordered on the x axis according to its position in the mouse genome. The distance of each SNV to the previous SNV in the genome is shown on the y axis. The colouring of individual SNV dots indicates the type of nucleotide substitution. y, Chr4 ‘zoom-in’ from x. Breakpoint junctions are shown according to their genomic position on chr4. No mutation clusters—either in absence or in combination with breakpoint junctions—were detected, consistent with chromothripsis involving end joining DNA repair mechanisms. This is in contrast to other complex rearrangement types, such as chromoanasynthesis, which arise through replication-based mechanisms with breakpoint-associated high mutation rates (for example, kataegis).
Extended Data Figure 3 Specificity, timing, mechanisms and impact of KrasG12D gene dosage alterations on gene expression in pancreatic tumorigenesis.
a, Overlay of copy number profiles of primary mPDAC cell cultures from PK mice (n = 38) as determined by aCGH. The y axis shows the frequency of a genomic region to be amplified (up) or deleted (down) in the cohort, with Cdkn2a and Kras loci being most frequently affected by CNAs. b, Prevalence of LOH in primary mPDAC cell cultures from PK mice (n = 38) on the basis of WES data. A chromosome was considered to be affected by LOH if the SNP frequency was shifted to no more than 0.1 or at least 0.9 in a segment with a size of at least 200 kb. LOH on chr4 is frequently the consequence of heterozygous deletions involving the Cdkn2a locus. By contrast, LOH on chr6 is predominantly copy number neutral and linked to increased KrasG12D gene dosage. Chr4 (home of Cdkn2a) and chr6 (home of Kras) show markedly increased rates of LOH compared with all other chromosomes, reflecting their functional importance during tumorigenesis. c–h, Genetic mechanisms of KrasG12D gene dosage alterations as identified by aCGH, M-FISH and WES in pancreatic cancers from PK mice. The observed types of increased KrasG12D gene dosage acquisition were (1) focal gain (affecting no more than 50% of the chromosome length), arising either through replication-based mechanisms (two cases, one with high-level KrasG12D amplification (shown in c) and one with low level amplification) or translocation and subsequent amplification of the translocated chromosome (one case (shown in d)), (2) arm-level gain (affecting at least 50% of the chromosome length) arising through mitotic errors (seven cases of whole-chromosome gain (example shown in e), occasionally (two cases) with concomitant intra-chromosomal deletions or translocations not affecting Kras (example shown in f)) and (3) copy-number-neutral LOH (CN-LOH, KrasG12D homozygosity, acquired uniparental disomy), arising either through mitotic recombination (affecting parts of chr6 (shown in h)) or chromosomal missegregation (duplication of KrasG12D-mutant chr6 and loss of wild-type chr6 (shown in g)). c, mPDAC S134 shows a high-order focal amplification of KrasG12D. The sharp borders, small size of the amplification (600 kb) and strong increase in copy number (4×) indicate that KrasG12D was amplified through multiple cycles of repeated template-switching by a replication-based DNA repair mechanism. The KrasG12D mutant allele frequency is 89.1%. d, Tumour 4706 carries a focal amplification of KrasG12D. M-FISH analysis revealed that the mutant KrasG12D allele (chr6) was probably first affected by a reciprocal translocation of chr4 and chr6, resulting in two rearranged chromosomes: der(4)T(4;6) and der(6)T(4;6). Subsequently, der(4)T(4;6) was missegregated through mitotic error, resulting in focal gain of the KrasG12D locus. The KrasG12D mutant allele frequency is 72.2%. e, mPDAC R1035 shows ‘classic’ whole-chromosome gain (trisomy) of chr6, which was probably generated through mitotic error or missegregation. The KrasG12D mutant allele frequency is 69.8%. f, In tumour 8442, arm-level gain of KrasG12D was probably generated through mitotic missegregation of chr6. Intra-chromosomal deletion on one of three chromosomes (19.6 Mb) does not affect Kras. The KrasG12D mutant allele frequency is 66.4%. Asterisk, chr6 with reduced length resulting from intra-chromosomal deletion. g, h, mPDAC 16992 and B590 display CN-LOH, leading to increased KrasG12D gene dosage. The KrasG12D mutant allele frequencies are 99.2% and 96.3%, respectively. The SNP pattern of chr6 in mPDAC 16992 reveals that the whole chromosome is affected by CN-LOH, indicating chromosome missegregation (duplication of the KrasG12D-mutant chr6 and loss of wild-type chr6) as the underlying mechanism. By contrast, in mPDAC B590 only a partial region of chr6 is affected by CN-LOH, therefore probably resulting from mitotic recombination. i, Allele-specific KrasG12D mRNA expression in KrasG12D-HET (n = 12) versus KrasG12D-iGD (n = 26) primary PDAC cell cultures from PK mice as detected by combined analysis of amplicon-based RNA-seq (proportion of mutant/wild-type Kras mRNA) and 3′ polyadenylation RNA-seq (amount of total Kras mRNA, but not the proportion of mutant/wild-type Kras mRNA, owing to sequencing of 3′ transcript ends; see Methods). This figure is related to Fig. 2b. ***P ≤ 0.001, two-tailed Mann–Whitney test; bars, median. j, Mutant KrasG12D mRNA levels in Cdkn2a/Ncruc∆HET/WT (n = 11) versus Ckdn2a/Ncruc∆HOM (n = 27) primary PDAC cell cultures from PK mice as detected by combined amplicon-based RNA-seq and 3′ polyadenylation RNA-seq. This figure is related to Extended Data Fig. 5f. ***P ≤ 0.001, two-tailed Mann–Whitney test; bars, median. k, Mutant KrasG12D mRNA levels in transcriptional clusters of mPDAC from PK mice (C2a/b/c/C1, n = 5/7/6/15) as detected by combined amplicon-based RNA-seq and 3′ polyadenylation RNA-seq. This figure is related to Fig. 5d. P = 1.6 × 10−5, two-sided Pearson correlation; bars, median. l–n, Interphase FISH for the analysis of copy-number and ploidy states at the KRAS locus on chr12 in hPanIN with KRASG12 variant allele frequencies (VAFs) of approximately 100%. KRASG12 VAFs are indicated above each FISH profile as detected by amplicon-based deep sequencing. A VAF of approximately 100% can be caused either by loss of the wild-type KRAS locus (hemizygosity of KRASG12-MUT: one KRASG12-MUT allele per cell) or by CN-LOH (acquired uniparental disomy; homozygosity of KRASG12-MUT: two KRASG12-MUT alleles per cell). All samples show a diploid genome as suggested by CEN12 (centromere probe chr12; two red signals per nucleus). Neither loss of one KRAS allele nor monosomy of chr12 was observed, providing evidence for CN-LOH and increased KRASG12-MUT gene dosage in hPanIN. Scale bars, 2.5 μm.
Extended Data Figure 4 Enrichment for amplification of alternative oncogenic drivers in mPDACs of PK mice with KrasG12D-HET status.
a, b, Two primary mPDACs with strong focal Myc amplification on chr15 are shown, as detected by aCGH. Red dashed line indicates no copy-number change. c, d, Focal copy number gains targeting the Yap1 locus on chr9 in primary mPDACs 4072 and 9203 as revealed by aCGH. e, Chr19 was also frequently subject to arm-level gain (see Fig. 1c and Extended Data Fig. 1l). Primary mPDAC of PK mouse 4072 harbours a focal gain on chr19 containing 20 genes: 9130011E15Rik, Gm6813, Hps6, Ldb1, Pprc1, Nolc1, Elovl3, Pitx3, Gbf1, Nfkb2, Psd, Fbxl15, Cuedc2, Tmem180, Actr1a, Sufu, Trim8, Arl3, Sfxn2 and D19Wsu162e. f, Cross-species analyses revealed that the orthologous region on human chr10 is also subject to recurrent amplifications in human PDAC (8 out of 109 hPDACs have focal amplifications; data from ref. 6). Of the 20 mouse genes, 16 could be assigned to orthologues in humans. Further analyses revealed that only two genes, NFKB2 and PSD, are within the minimal overlapping region of recurrent amplification (data from ref. 6 and oncoplot from cBioPortal60,61). g, NFKB2, but not PSD, shows medium protein expression in exocrine glandular cells of normal pancreatic tissue, as detected by immunohistochemistry (data from the Human Protein Atlas62). h, NFKB2 is highly expressed in 17% (2 out of 12) of stained hPDAC biopsies as shown by immunohistochemistry. In contrast, there was no PSD expression in any of the analysed pancreatic cancers (0 out of 12). Protein expression data were used from the Human Protein Atlas62.
Extended Data Figure 5 Characterization of Cdkn2a (chr4) alterations and correlation with KrasMUT gene dosage variation and mRNA expression in mouse and human PDAC.
a–d, Cdkn2a alteration on mouse chr4 can occur through arm-level, complex or focal loss as well as uniparental disomy (see Fig. 3). In addition, chr4 is frequently involved in inter-chromosomal translocations. Examples of representative karyotypes of primary pancreatic cancer cultures derived from PK mice with translocations involving chr4, probably affecting the Cdkn2a locus. In all four cases, chr4 translocations were found in all ten metaphase spreads of each cancer, indicating their early acquisition during tumour evolution. a, mPDAC 4706 with diploid karyotype: 42, XX, del(X), +2, der(2)t(2;4)is(2;4), der(4)t(4;6)*2, +der(4)t(2;4), der(6)t(4;6). b, mPDAC 4900 also features a diploid karyotype: 41, XX, der(X)is(X;4), der(4)is(4;8), del(4), +6, der(8)t(4;8). c, mPDAC 5123 underwent polyploidization, after translocation of chr4 with chr1 and a deletion on the other copy: 78, XXXX, −1, del(1)*2, −2, +4*2, der(4)t(1;4)*3, del(4)*3, −5, −7, −9, +15, −17, +18. d, mPDAC 8349 shows a diploid karyotype: 40, XX, der(4)t(3;4), der(4)t(4;13), +del(4), der(13)t(4;13). e, KrasG12D variant allele frequencies detected by amplicon-based deep sequencing of the Kras locus are higher in Cdkn2a/Ncruc∆HOM mPDAC (n = 27) compared with Cdkn2a/Ncruc∆HET/WT (n = 11) pancreatic cancers. All cancers are from PK mice. Blue dots indicate tumours with complete Ncruc deletion. ***P ≤ 0.001, two-tailed Mann–Whitney test; bars, median. f, Allele-specific expression of mutant KrasG12D mRNA is increased in primary tumours from PK mice with Cdkn2a/Ncruc∆HOM (n = 27) background compared with Cdkn2a/Ncruc∆HET/WT (n = 11) cancers. Primary mPDACs with homozygous loss of Ncruc are highlighted in blue. KrasG12D expression was analysed by combining amplicon-based RNA-seq and qRT–PCR (as described in the Methods section). **P = 0.003, two-tailed Mann–Whitney test; bars, median. g, KRASMUT variant allele frequencies based on WES in a published data set of microdissected human PDAC (ref. 6, reduced stromal content) was analysed with respect to CDKN2A and TP53 status. KRASMUT allele frequency was higher in mutated/homozygous deleted CDKN2A and/or TP53 (CDKN2AMUT/∆HOM/TP53MUT/∆HOM; hPDACs compared with cancers with CDKN2A∆HET/WT/TP53∆HET/WT status (from left: n = 28, n = 14, n = 28, n = 30). Two-sided rank-based ANOVA (P = 5.8 × 10−6); post hoc testing with two-sided Tukey honest significant difference test, *adjusted P ≤ 0.05, ***adjusted P ≤ 0.001; bars, median. h, Fraction of the genome altered by copy number changes detected by aCGH in primary mPDACs of PK (n = 38), PKC (n = 16) and PKP (n = 16) mice. PKP mice show a significantly increased CNA load compared with PKC mice. Two-sided rank-based ANOVA (P = 0.01); post hoc testing with two-sided Tukey honest significant difference test, **adjusted P = 0.009, adjusted P values for group-wise comparisons are shown; bars, median. Del, deletion; der, derivative chromosome; is, insertion; t, translocation; ‘−’, chromosome loss; ‘+’, chromosome gain.
Extended Data Figure 6 Complete Cdkn2a barrier loss precedes KrasG12D-iGD in primary mPDAC of PK mouse 53704.
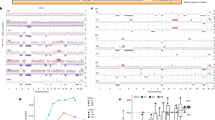
CNAs at chr4 (Cdkn2a) and chr6 (Kras) in mPDAC 53704 and corresponding metastases, as detected by aCGH (top) and WES-based SNP pattern analysis (bottom). The primary cancer and both liver metastases display identical focal deletions of Cdkn2a and similar SNP patterns on chr4, revealing that all lesions share the same ancestor cell with complete Cdkn2a loss. By contrast, SNP analysis on chr6 revealed discordant patterns in the primary mPDAC and both metastases. Li2 shows partial LOH of a distal region on chr6 involving the Kras locus, while LOH in Li3 involves the whole chr6. This explains the stepwise LOH pattern observed on chr6 in the primary mPDAC. The graphic on the right shows the combined interpretation of CNA and LOH profiles, which suggests the following sequence of genetic events during tumour evolution. The initial KrasG12D mutation was followed by focal deletion of one copy of Cdkn2a. In a subsequent genetic event, the second copy of Cdkn2a was lost by chr4 missegregation and copy-number-neutral LOH. Complete barrier loss allowed for convergent evolution of increased KrasG12D gene dosage through copy-number-neutral LOH and gave rise to independent metastases in the liver. Note that a major obstacle for equivalent human studies is the limited availability of human matched primary–metastasis samples, particularly of treatment-naive ones. We performed cross-species analyses using data from a recent study, which analysed human treatment-naive metastatic PDACs by WGS8 and provided CDKN2A and KRAS copy number data for matched primaries and metastases from three patients. In one patient the sequential order of CDKN2A deletion and KRAS amplification could be reconstructed: homozygous CDKN2A deletions were identical in all primaries and metastases, whereas there were five different KRAS gains in the six metastases. This suggests convergent evolution of mutant KRAS gene dosage gain upon homozygous CDKN2A loss in this patient, in line with similar data in large series of mouse cancers and their metastases (see Fig. 3e).
Extended Data Figure 7 Transcriptome-based subtyping of human primary pancreatic cancer and classification of human PDAC cell lines and primary PDAC cell cultures from PK mice.
a–c, Independent cross-comparison of transcriptional classification systems from Collisson et al.28, Moffitt et al.29 and Bailey et al.7. Collisson et al.28 performed PDAC microdissection and defined three transcriptional subtypes: classical, quasimesenchymal and exocrine-like. Moffitt et al.29 defined two subtypes (classical, basal-like) using (1) virtual separation of tumour and non-tumour gene expression patterns, (2) transplantation studies and (3) human PDAC cell lines; and they proposed that the exocrine-like signature stems from exocrine pancreatic cells, rather than from the cancer cells. Bailey et al.7 used bulk tumours and defined four subtypes (pancreatic progenitor, immunogenic, squamous, aberrantly differentiated endocrine exocrine (ADEX)). RNA-seq data from PDAC and adenosquamous pancreatic carcinoma from Bailey et al.7 were used for cross-comparison of classification systems. Other histological subentities of pancreatic cancer were excluded (for example, intraductal papillary mucinous neoplasm, mucinous cystic neoplasm, acinar cell carcinoma). The subtyping from Bailey et al.7 for this data set was available. a, Unbiased hierarchical clustering of primary pancreatic cancer samples (n = 71) from Bailey et al.7 using classifier genes of Collisson et al.28. b, Subtyping of primary pancreatic cancer samples (n = 71) from Bailey et al.7 using classifier genes defined by Moffitt et al.29. c, Consensus clustering based on analyses performed in a and b. There is considerable overlap between at least two subtypes, which are in large parts captured by the initially proposed classical and quasimesenchymal signatures from Collisson et al.28(which are also detected in mouse and human PDAC cell lines; see Extended Data Fig. 7e–h). The classification of Bailey et al.7(based on bulk tissue analyses) suggests that classical cancers of Collisson et al.28(microdissected cancer tissue) can be further sub-stratified into some with and some without a strong immune cell infiltration. The classification of Moffitt et al.29 suggests that the exocrine-like signature of Collisson et al.28(ADEX subtype in Bailey et al.7) stems from ‘contaminating’ healthy exocrine pancreatic cells, on the basis of the evidence described above. Given that the exocrine-like signature of Collisson et al.28 was derived from microdissected PDAC, such ‘contamination’ is only conceivable if exocrine-like signature genes were much more highly expressed in pancreatic acinar cells than in PDAC cells. d, Volcano plot showing strongly upregulated expression of exocrine-like genes in human wild-type pancreas (13- to 241-fold; median 183-fold upregulation). Note that 15 out of 19 exocrine-like signature genes (red dots) are among the top 50 genes upregulated in human wild-type pancreas (n = 3) compared with hPDAC cell lines (n = 30) (y axis is calculated on Benjamini–Hochberg adjusted P values derived from R package limma (see Methods)). Although these data do not exclude the existence of exocrine-like PDACs, they support the possibility that ‘contamination’ with few acinar cells can impose an exocrine-like signature on a cancer. This might explain why human or mouse PDAC cell lines do not cluster into the exocrine-like subtype (see also Extended Data Fig. 7e, f). e, Hierarchical clustering of microarray-based expression profiles using identifier genes from Collisson et al.28 on human PDAC cell lines (n = 19, Gene Expression Omnibus series GSE17891). As also described in Collisson et al.28, only two subtypes can be detected in human cell line collections: classical and quasimesenchymal. Of note, the most prominent change in the quasimesenchymal cell lines is downregulation (extinction) of the classical assigner genes, whereas expression of quasimesenchymal classifier genes is quite variable. We therefore also use here the terms ‘classical’ and ‘non-classical’. f, Projection of the Collisson et al.28 classifiers on to mouse PDAC cell culture transcriptomes (n = 33) also identified classical and non-classical subtypes. The non-classical subtype contained a subset of mPDAC cell cultures from cluster C2a/b/c (epithelial morphology; equivalent of human quasimesenchymal) and all cluster C1 mPDACs (mesenchymal morphology; ‘M’ cluster). g, Application of a human EMT hallmark gene set52 for hierarchical clustering of expression profiles from primary PDAC cultures (PK mice; n = 33) resulted in a separation of C1 (mesenchymal) and C2a/b/c (epithelial) cell lines. h, Projection of the EMT hallmark gene set on human PDAC cell line transcriptomes (n = 19, Gene Expression Omnibus series GSE17891) did not result in a clear separation of samples, indicating underrepresentation of the mesenchymal M subtype (equivalent to mouse C1/‘M’) in available human cell line collections. As shown in Extended Data Fig. 9b, however, the EMT signature is detectable in undifferentiated human pancreatic carcinoma, which is the human equivalent of the mesenchymal mouse PDACs in C1.
Extended Data Figure 8 Functional analyses to study the role of increased KrasG12D gene dosage in EMT.
a–d, Multiplexed somatic CRISPR–Cas9 mutagenesis for phylogenetic tracking of epithelial–mesenchymal mPDAC clones in vivo. a, Major steps of multiplexed gene editing by pooled delivery of CRISPR–Cas9 vectors, each targeting a different tumour suppressor gene in the pancreas of PK mice. Electroporation-based transfection enables low-frequency mosaic vector delivery (average of 120 cells per pancreas are transfected) to induce clonal tumours. Primary tumour cell cultures were screened for the simultaneous presence of epithelial and mesenchymal cells. Two such cancers were identified (mPDACs from mouse 021 and mouse 901) and subjected to differential trypsinization to enrich for each morphology. b, Amplicon-based deep sequencing of all single-guide RNA-targeted loci revealed identical indel patterns in both epithelial–mesenchymal culture pairs. This shows (1) that epithelial and mesenchymal cells originate from the same clone and (2) that the CRISPR-induced mutations do not contribute to the differential phenotype. c, KrasG12D VAFs in epithelial and mesenchymal cell cultures from mPDAC 021 and mPDAC 901, as detected by amplicon-based deep sequencing. Both cancers had increased KrasG12D expression in mesenchymal cells (see Fig. 5e). In mPDAC 021, this is due to selective amplification of the KrasG12D allele in mesenchymal cells. In mPDAC 901, genetic KrasG12D amplification was not observed, suggesting induction of increased Kras expression in mesenchymal cells by other mechanisms. d, Gene set enrichment analysis using MSigDB of differentially regulated genes in mesenchymal versus epithelial mPDACs on the basis of RNA-seq. Mesenchymal clones of mPDAC 021 and mPDAC 901 show an upregulation of genes involved in ‘MAPK signaling pathway’ and ‘EMT’ compared with the corresponding epithelial clones, in line with increased KrasG12D gene dosage (a full list of enriched gene sets is provided for comparison in Supplementary Table 15). False discovery rate-adjusted P values are shown on the y axis. Representative data from one experiment are shown. e–g, Induction of EMT-like transcriptional programs by KRASG12D overexpression in human PDAC cell lines. e, Graphic of experimental workflow. Two human PDAC cell lines (HUPT3 and PANC0327) with homozygous CKDN2A loss (CDKN2A∆HOM) and heterozygous KRASMUT (KRASMUT-HET) status were transduced with lentivirus carrying doxycycline-inducible KRASG12D or GFP-control expression constructs. KRASG12D or GFP expression was induced by adding doxycycline for 1, 3 or 5 days. f, Gene set enrichment analysis using MSigDB of differentially regulated genes in KRASG12D- versus GFP-induced hPDAC cell lines HUPT3 and PANC0327 on the basis of RNA-seq. Upon doxycycline treatment, both hPDAC cell lines showed consistent upregulation of genes involved in ‘KRAS signaling up’ and ‘EMT’ (a full list of enriched gene sets is provided for both cell lines in Supplementary Table 16). False discovery rate-adjusted P values are shown on the y axis. g, Expression of marker genes for epithelial (CDH1) or mesenchymal (VIM) cell differentiation and invasion or matrix disassembly (MMP1) was validated by qPCR (normalized to GAPDH and PPIA). In line with RNA-seq data, KRASG12D-induced cells show an increased expression of the mesenchymal marker gene VIM, increased expression of MMP1 and reduced levels of epithelial marker gene CDH1. *P ≤ 0.05, **P ≤ 0.005, NS, not significant, two-tailed t-test; bars, mean; error bars, s.e.m.
Extended Data Figure 9 Transcriptional profiles of human undifferentiated pancreatic carcinomas are enriched for signatures of oncogenic signalling intensification and EMT but not for activation of TP63∆N transcriptional network.
a, Primary pancreatic tumours from PK mice with a mesenchymal phenotype (C1 cluster, n = 15) are almost exclusively classified as undifferentiated or sarcomatoid by histopathological evaluation and tend to have a reduced age at diagnosis when compared with epithelial (C2a/b/c cluster, n = 18) tumours (histopathological grade 1–3 (G1–G3)). This aggressive behaviour of undifferentiated pancreatic carcinoma is also observed in human patients and is associated with worse clinical outcome33. P value calculated by two-sided log-rank test. b, Comparison of publicly available expression profiles of human undifferentiated pancreatic carcinoma (n = 4), PDAC (World Health Organization grades 1–3 (G1–G3), n = 64) and adenosquamous pancreatic carcinoma (n = 7). Human samples with the above histopathological characteristics for which expression-based subtype information from Bailey et al.7 was available were used and complemented with available undifferentiated pancreatic carcinomas from the ICGC PACA-AU cohort (Supplementary Table 18). Other histological subentities of pancreatic cancer were excluded (for example, intraductal papillary mucinous neoplasm, mucinous cystic neoplasm, acinar cell carcinoma). ANOVA was performed to select genes that were differentially expressed in at least one of the six defined subgroups of pancreatic cancer: (1) undifferentiated, (2) adenosquamous pancreatic carcinoma and (3–6) PDAC (G1–G3) sub-stratified in pancreatic progenitor, immunogenic, squamous and ADEX subtypes from Bailey et al.7. Differentially regulated genes were used for unbiased hierarchical clustering of these pancreatic cancer transcriptional profiles. Five sub-clusters of co-regulated gene expression could be identified according to the cluster tree on the y axis (separated by white horizontal bars in the heatmap). Gene set enrichment analysis using MSigDB was performed for individual sub-clusters and terms related to predominating gene sets or pathways are annotated for each cluster on the right (full list provided in Supplementary Table 17). Undifferentiated pancreatic carcinomas cluster together and are associated with (1) upregulation of genes in cluster 3 (containing MAPK signalling pathway and gene sets relevant during embryonic development or EMT) and (2) downregulation of genes in clusters 2 and 5, which contain gene sets related to epithelial cell differentiation, embryonic development or metabolic signatures. This reflects the pathway enrichment signature in the equivalent undifferentiated (mesenchymal) mouse PDACs (cluster C1/‘M’ in PK mice; see Extended Data Fig. 7g) and provides further support for the link between KRAS signalling intensification, EMT and the undifferentiated tumour phenotype. The immunogenic PDAC subtype showed high expression of cluster 4 genes, which was also strong (even elevated) in undifferentiated pancreatic carcinomas, suggesting increased immune cell infiltration in undifferentiated carcinomas. Cluster 1 contained gene sets related to cell proliferation/cell cycle, squamous differentiation and TP63∆N transcriptional targets, which were most highly overexpressed in pancreatic carcinomas with adenosquamous histology. Undifferentiated pancreatic carcinomas did not show activation of the TP63∆N transcriptional targets. This suggests that activation of TP63∆N transcriptional targets is not causally linked to KRAS signalling intensification and EMT (see also Extended Data Fig. 9c, d, showing a lack of association of undifferentiated carcinomas withTP63∆N transcriptional network activation). c, Unbiased hierarchical clustering of human pancreatic carcinomas with adenosquamous histology (n = 7) as well as PDACs (World Health Organization grades 1–3 (G1–G3), n = 64) and undifferentiated pancreatic carcinomas (n = 4) (sample set as in Extended Data Fig. 9b) using a list of validated TP63∆N transcriptional targets53. Pancreatic cancers with adenosquamous differentiation were significantly enriched in a cluster showing increased TP63∆N transcriptional network activity (P ≤ 0.001, two-sided Fisher’s exact test, odds ratio 130, 95% confidence interval 11.6–1,452). Undifferentiated pancreatic carcinomas did not contribute to this cluster. In line with these results, pancreatic cancers from PK mice did not show differential regulation of the TP63∆N network, reflecting the lack of adenosquamous tumours in this cohort (not shown). d, Unbiased hierarchical clustering across solid cancers (Cancer Cell Line Encyclopedia, n = 856) using the same gene list showed a strong enrichment of tumours with squamous differentiation in the sub-cluster with the highest TP63∆N transcriptional network expression (P ≤ 0.001, two-sided Fisher’s exact test, odds ratio 28.1, 95% confidence interval 16.4–48.1), in line with the observation in ref. 63 that TP63∆N is a signature for squamous differentiation across cancers.
Extended Data Figure 10 KrasG12D gene dosage is a critical determinant of PDAC biology in a mouse model with high mutational load.
a, The mutational burden in primary PDAC cultures of PK mice was significantly lower compared with human PDAC studies (see Fig. 1b). To account for this potential confounding factor and to test whether our discoveries in PK mice also applied in a setting of high mutational burden, we used a mouse model combining KrasG12D mutation and PiggyBac transposon-based insertional mutagenesis (PK–PB mice13). PK–PB mice show accelerated tumourigenesis compared with PK mice. PK–PB-derived tumours had an extensive mutational burden (median of 494 transposon insertions per tumour). Primary cultures of PDAC from PK–PB mice (n = 17) were subjected to comprehensive genetic characterization using aCGH, microarray-based gene expression profiling, quantitative transposon insertion-site sequencing and amplicon-based deep sequencing of the Kras locus. b, Transcriptome profiles of primary PDAC cultures from PK–PB mice (n = 17) were used for unbiased hierarchical clustering that resulted in two major clusters (C1 and C2), as in PK mice. KrasG12D gene dosage status (as determined by aCGH and amplicon-based deep sequencing of the Kras locus) and Cdkn2a status (as determined by aCGH and quantitative transposon insertion-site sequencing) are indicated below the cluster tree for each individual tumour. Similarly to PK mice, cluster C2a was characterized by KrasG12D-HET and Cdkn2a/Ncruc∆HET/WT status, whereas mPDACs in clusters C2b/c and C1 had increased KrasG12D gene dosage (KrasG12D-iGD) and were Cdkn2a/Ncruc∆HOM. The genetic KrasG12D status was significantly associated with expression clusters (P = 0.01, two-sided Fisher’s exact test), providing further evidence that expression clusters are associated with KrasG12D gene dosage. c, Prevalence of KrasG12D-iGD in cultures of primary mPDAC (from PK–PB mice) with homozygous (n = 12) or heterozygous/wild-type (n = 5) Cdkn2a/Ncruc status. *P = 0.03, two-sided Fisher’s exact test, odds ratio 20.0, 95% confidence interval 1.4–287.8. d, Gene set enrichment analysis using DAVID of upregulated genes in cluster C1 (n = 5) compared with cluster C2 (n = 12) of primary mPDAC cultures from PK–PB mice. As in PK mice, PK–PB tumours in C1 are characterized by upregulation of genes enriched in gene sets describing mesenchymal cell differentiation and revealed a strong enrichment for Ras downstream signalling pathways (full list in Supplementary Table 19). False discovery rate-adjusted P values are shown on the y axis. Overall, these analyses show that the biological principles discovered in the PK model also apply to pancreatic cancers from PK–PB mice with high mutational load.
Supplementary information
Supplementary Data
This zipped file contains Supplementary Tables 1-20 and a Supplementary Table Guide. (ZIP 2161 kb)
Rights and permissions
About this article
Cite this article
Mueller, S., Engleitner, T., Maresch, R. et al. Evolutionary routes and KRAS dosage define pancreatic cancer phenotypes. Nature 554, 62–68 (2018). https://doi.org/10.1038/nature25459
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25459
This article is cited by
-
The protein phosphatase-2A subunit PR130 is involved in the formation of cytotoxic protein aggregates in pancreatic ductal adenocarcinoma cells
Cell Communication and Signaling (2024)
-
KRAS allelic imbalance drives tumour initiation yet suppresses metastasis in colorectal cancer in vivo
Nature Communications (2024)
-
Inhibition of ADAM9 promotes the selective degradation of KRAS and sensitizes pancreatic cancers to chemotherapy
Nature Cancer (2024)
-
Clinical significance of serum-derived exosomal PD-L1 expression in patients with advanced pancreatic cancer
BMC Cancer (2023)
-
SOX2 dosage sustains tumor-promoting inflammation to drive disease aggressiveness by modulating the FOSL2/IL6 axis
Molecular Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.