Abstract
The genus Citrus, comprising some of the most widely cultivated fruit crops worldwide, includes an uncertain number of species. Here we describe ten natural citrus species, using genomic, phylogenetic and biogeographic analyses of 60 accessions representing diverse citrus germ plasms, and propose that citrus diversified during the late Miocene epoch through a rapid southeast Asian radiation that correlates with a marked weakening of the monsoons. A second radiation enabled by migration across the Wallace line gave rise to the Australian limes in the early Pliocene epoch. Further identification and analyses of hybrids and admixed genomes provides insights into the genealogy of major commercial cultivars of citrus. Among mandarins and sweet orange, we find an extensive network of relatedness that illuminates the domestication of these groups. Widespread pummelo admixture among these mandarins and its correlation with fruit size and acidity suggests a plausible role of pummelo introgression in the selection of palatable mandarins. This work provides a new evolutionary framework for the genus Citrus.
Similar content being viewed by others
Main
The genus Citrus and related genera (Fortunella, Poncirus, Eremocitrus and Microcitrus) belong to the angiosperm subfamily Aurantioideae of the Rutaceae family, which is widely distributed across the monsoon region from west Pakistan to north-central China and south through the East Indian Archipelago to New Guinea and the Bismarck Archipelago, northeastern Australia, New Caledonia, Melanesia and the western Polynesian islands1. Native habitats of citrus and related genera roughly extend throughout this broad area (Extended Data Fig. 1a and Supplementary Table 1), although the geographical origin, timing and dispersal of citrus species across southeast Asia remain unclear. A major obstacle to resolving these uncertainties is our poor understanding of the genealogy of complex admixture in cultivated citrus, as has recently been shown2. Some citrus are clonally propagated apomictically3 through nucellar embryony, that is, the development of non-sexual embryos originating in the maternal nucellar tissue of the ovule, and this natural process may have been co-opted during domestication; grafting is a relatively recent phenomenon4. Both modes of clonal propagation have led to the domestication of fixed (desirable) genotypes, including interspecific hybrids, such as oranges, limes, lemons, grapefruits and other types.
Under this scenario, it is not surprising that the current chaotic citrus taxonomy—based on long-standing, conflicting proposals5,6—requires a solid reformulation consistent with a full understanding of the hybrid and/or admixture nature of cultivated citrus species. Here we analyse genome sequences of diverse citrus to characterize the diversity and evolution of citrus at the species level and identify citrus admixtures and interspecific hybrids. We further examine the network of relatedness among mandarins and sweet orange, as well as the pattern of the introgression of pummelos among mandarins for clues to the early stages of citrus domestication.
Diversity and evolution of the genus Citrus
To investigate the genetic diversity and evolutionary history of citrus, we analysed the genomes of 58 citrus accessions and two outgroup genera (Poncirus and Severinia) that were sequenced to high coverage, including recently published sequences2,3,7 as well as 30 new genome sequences described here. For our purpose, we do not include accessions related by somatic mutations. These sequences represent a diverse sampling of citrus species, their admixtures and hybrids (Supplementary Tables 2, 3 and Supplementary Notes 1, 2). Our collection includes accessions from eight previously unsequenced and/or unexamined citrus species, such as pure mandarins (Citrus reticulata), citron (Citrus medica), Citrus micrantha (a wild species from within the subgenus Papeda), Nagami kumquat (Fortunella margarita, also known as Citrus japonica var. margarita), and Citrus ichangensis (also known as Citrus cavaleriei; this species is also considered a Papeda), as well as three Australian citrus species (Supplementary Notes 3, 4). For each species, we have sequenced one or more pure accessions without interspecific admixture.
Local segmental ancestry of each accession can be delineated for both admixed and hybrid genotypes, based on genome-wide ancestry-informative single-nucleotide polymorphisms (Supplementary Note 5). Comparative genome analysis further identified shared haplotypes among the accessions (Supplementary Notes 6, 7). In particular, we demonstrate the F1 interspecific hybrid nature of Rangpur lime and red rough lemon (two different mandarin–citron hybrids), Mexican lime (a micrantha–citron hybrid) and calamondin (a kumquat–mandarin hybrid), and confirm, using whole-genome sequence data, the origins of grapefruit (a pummelo–sweet orange hybrid), lemon (a sour orange–citron hybrid) and eremorange (a sweet orange and Eremocitrus glauca (also known as Citrus glauca) hybrid). We also verified the parentage of Cocktail grapefruit, with low-acid pummelo as the seed parent and King and Dancy mandarins as the two grandparents on the paternal side. The origin of the Ambersweet orange is similarly confirmed to be a mandarin–sweet orange hybrid with Clementine as a grandparent. We have previously shown that sour orange (cv. Seville) (Citrus aurantium) is a pummelo–mandarin hybrid, and have analysed the more complex origin of sweet orange (Citrus sinensis)2. Re-analysing sequences from ten cultivars of sweet orange3 shows that they are all derived from the same genome by somatic mutations, and were thus not included in our study.
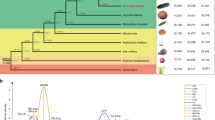
We identified ten progenitor citrus species (Supplementary Note 4.1) by combining diversity analysis (Extended Data Table 1), multidimensional scaling and chloroplast genome phylogeny (Extended Data Fig. 1b). The first two principal coordinates in the multidimensional scaling (Fig. 1a) separate three ancestral (sometimes called ‘fundamental’) Citrus species associated with commercially important types8,9—citrons (C. medica), mandarins (C. reticulata) and pummelos (Citrus maxima)—and display lemons, limes, oranges and grapefruits as hybrids involving these three species. The nucleotide diversity distributions (Fig. 1b) show distinct scales for interspecific divergence and intraspecific variation, and reflect the genetic origin of each accession. Hybrid accessions (sour orange, calamondin, lemon and non-Australian limes) with ancestry from two or more citrus species are readily identified on the basis of their higher segmental heterozygosity (1.5–2.4%) relative to intraspecific diversity (0.1–0.6%). Other citrus accessions show bimodal distributions in heterozygosity (sweet orange, grapefruits and some highly heterozygous mandarins) due to interspecific admixture, a process that generally involves complex backcrosses. Among the pure genotypes without interspecific admixture, citrons show significantly lower intraspecific diversity (around 0.1%) than the other species (0.3–0.6%). The reduced heterozygosity of citrons, a mono-embryonic species, is probably due to the cleistogamy of its flowers10, a mechanism that promotes pollination and self-fertilization in unopened flower buds, which in turn reduces heterozygosity.
a, Principal coordinate analysis of 58 citrus accessions based on pairwise nuclear genome distances and metric multidimensional scaling. The first two axes separate the three main citrus groups (citrons, pummelos and mandarins) with interspecific hybrids (oranges, grapefruit, lemon and limes) situated at intermediate positions relative to their parental genotypes. b, Violin plots of the heterozygosity distribution in 58 citrus accessions, representing 10 taxonomic groups as well as 2 related genera, Poncirus (Poncirus trifoliata, also known as Citrus trifoliata) and Chinese box orange (Severinia). White dot, median; bar limits, upper and lower quartiles; whiskers, 1.5× interquartile range. The bimodal separation of intraspecies (light blue) and interspecies (light pink) genetic diversity is manifested among the admixed mandarins and across different genotypes including interspecific hybrids. Three-letter codes are listed in parenthesis with additional descriptions in Supplementary Table 2. c, Chronogram of citrus speciation. Two distinct and temporally well-separated phases of species radiation are apparent, with the southeast Asian citrus radiation followed by the Australian citrus diversification. Age calibration is based on the citrus fossil C. linczangensis16 from the Late Miocene (denoted by a filled red circle). The 95% confidence intervals are derived from 200 bootstraps. Bayesian posterior probability is 1.0 for all nodes. d, Proposed origin of citrus and ancient dispersal routes. Arrows suggest plausible migration directions of the ancestral citrus species from the centre of origin—the triangle formed by northeastern India, northern Myanmar and northwestern Yunnan. The proposal is compatible with citrus biogeography, phylogenetic relationships, the inferred timing of diversification and the paleogeography of the region, especially the geological history of Wallacea and Japan. The red star marks the fossil location of C. linczangensis. Citrus fruit images in c and d are not drawn to scale.
The identification of a set of pure citrus species provides new insights into the phylogeny of citrus, their origins, evolution and dispersal. Citrus phylogeny is controversial1,5,6,11,12, in part owing to the difficulty of identifying pure or wild progenitor species, because of substantial interspecific hybridization that has resulted in several clonally propagated and cultivated accessions. Some authors assign separate binomial species designations to clonally propagated genotypes1,6. Our nuclear genome-based phylogeny, which is derived from 362,748 single-nucleotide polymorphisms in non-genic and non-pericentromeric genomic regions, reveals that citrus species are a monophyletic group and establishes well-defined relationships among its lineages (Fig. 1c and Supplementary Note 8). Notably, the nuclear genome-derived phylogeny differs in detail from the chloroplast-derived phylogeny (Extended Data Fig. 1). This is not unexpected, as chloroplast DNA is a single, non-recombining unit and is unlikely to show perfect lineage sorting during rapid radiation (Supplementary Note 8.3).
The origin of citrus has generally been considered to be in southeast Asia1, a biodiversity hotspot13 with a climate that has been influenced by both east and south Asian monsoons14 (Supplementary Note 9). Specific regions include the Yunnan province of southwest China15, Myanmar and northeastern India in the Himalayan foothills1. A fossil specimen from the late Miocene epoch of Lincang in Yunnan, Citrus linczangensis16, has traits that are characteristic of current major citrus groups, and provides definite evidence for the existence of a common Citrus ancestor within the Yunnan province approximately 8 million years ago (Ma).
Our analysis establishes a relatively rapid Asian radiation of citrus species in the late Miocene (6–8 Ma; Fig. 1c, d), a period coincident with an extensive weakening of monsoons and a pronounced climate transition from wet to drier conditions17. In southeast Asia, this marked climate alteration caused major changes in biota, including the migration of mammals18 and rapid radiation of various plant lineages19,20. Australian citrus species form a distinct clade that was proposed to be nested with citrons12, although distinct generic names (Eremocitrus and Microcitrus) were assigned in botanical classifications by Swingle1,5. Both molecular dating analysis21 and our whole-genome phylogenetic analysis do not support an Australian origin for citrus22. Rather, citrus species spread from southeast Asia to Australasia, probably via transoceanic dispersals. Our genomic analysis indicates that the Australian radiation occurred during the early Pliocene epoch, around 4 Ma. This is contemporaneous with other west-to-east angiosperm migrations from southeast Asia23,24, presumably taking advantage of the elevation of Malesia and Wallacea in the late Miocene and Pliocene25,26 (Supplementary Note 9).
The nuclear and chloroplast genome phylogenies indicate that there are three Australian species in our collection. One of the two Australian finger limes shows clear signs of admixture with round limes (Supplementary Note 5.4). The closest relative to Australian citrus is Fortunella, a species that has been reported to grow in the wild in southern China27. Australian citrus species are diverse, and found natively in both dry and rainforest environments in northeast Australia, depending on the species28. Our phylogeny shows that the progenitor citrus probably migrated across the Wallace line, a natural barrier for species dispersal from southeast Asia to Australasia, and later adapted to these diverse climates.
The results also show that the Tachibana mandarin, naturally found in Taiwan, the Ryukyu archipelago and Japan29, split from mainland Asian mandarins (Fig. 1c, d) during the early Pleistocene (around 2 Ma), a geological epoch with strong glacial maxima30. Tachibana, as did other flora and fauna in the region, very probably arrived in these islands from the adjacent mainland31 during the drop in the sea level of the South China Sea and the emergence of land bridges32,33, a process promoted by the expansion of ice sheets that repetitively occurred during glacial maxima (Supplementary Note 9).
Although Tachibana5,6 has been assigned its own species (Citrus tachibana), sequence analysis reveals that it has a close affinity to C. reticulata34,35 and does not support its taxonomic position as a separate species (Supplementary Note 4.1). However, both chloroplast genome phylogeny (Extended Data Fig. 1b) and nuclear genome clustering (Fig. 1a) clearly distinguish Tachibana from the mainland Asian mandarins. This suggests that Tachibana should be designated a subspecies of C. reticulata. By contrast, the wild Mangshan ‘mandarin’ (Citrus mangshanensis)7 represents a distinct species, with comparable distances to C. reticulata, pummelo and citron2 (Extended Data Table 1).
Pattern of pummelo admixture in the mandarins
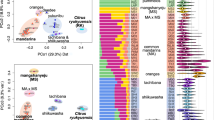
Using 588,583 ancestry-informative single-nucleotide polymorphisms derived from three species, C. medica, C. maxima and C. reticulata, we delineate the segmental ancestry of 46 citrus accessions (Extended Data Fig. 2 and Supplementary Note 5). Pummelo admixture is found in all but 5 of the 28 sequenced mandarins, and the amount and pattern of pummelo admixture, as identified by phased pummelo haplotypes (Fig. 2a and Supplementary Note 6), suggests the classification of the mandarins into three types.
a, Allelic proportion of five progenitor citrus species in 50 accessions. CI, C. medica; FO, Fortunella; MA, C. reticulata; MC, C. micrantha; PU, C. maxima; UNK, unknown. The pummelos and citrons represent pure citrus species, whereas in the heterogeneous set of mandarins, the degree of pummelo introgression subdivides the group into pure (type-1) and admixed (type-2 and -3) mandarins. Three-letter code as in Fig. 1, see Supplementary Table 2 for details. b, Genealogy of major citrus genotypes. The five progenitor species are shown at the top. Blue lines represent simple crosses between two parental genotypes, whereas red lines represent more complex processes involving multiple individuals, generations and/or backcrosses. Whereas type-1 mandarins are pure species, type-2 (early-admixture) mandarins contain a small amount of pummelo admixture that can be traced back to a common pummelo ancestor (with P1 or P2 haplotypes). Later, additional pummelo introgressions into type-2 mandarins gave rise to both type-3 (late-admixture) mandarins and sweet orange. Further breeding between sweet orange and mandarins or within late-admixture mandarins produced additional modern mandarins. Fruit images are not to scale and represent the most popular citrus types. See Supplementary Note 1.1 for nomenclature usage.
Type-1 mandarins represent pure C. reticulata with no evidence of interspecific admixture and include Tachibana, three unnamed Chinese mandarins (M01, M02, M04)3 and the ancient Chinese cultivar Sun Chu Sha Kat reported here, a small tart mandarin commonly grown in China and Japan, and also found in Assam. This cultivar is likely described in Han Yen-Chih’s ad 1178 monograph ‘Chü Lu’36, which includes references to citrus cultivated during the reign of Emperor Ta Yu (2205–2197 bc). Sixteen of the twenty-eight mandarins belong to type-2 mandarins, which have a small amount of pummelo admixture (1–10% of the length of the genetic map; Fig. 2a), usually in the form of a few short segments distributed across the genome. Although the lengths and locations of these admixed segments may be distinct in different mandarins, they share one or two common pummelo haplotypes (designated as P1 and P2) (Extended Data Fig. 3). By contrast, the seven remaining mandarins (type-3) contain higher proportions of pummelo alleles (12–38%; Fig. 2a) in longer segments. Although the P1 and P2 pummelo haplotypes are also detectable among type-3 mandarins, other more extensive pummelo haplotypes dominate the pummelo admixture in type-3 mandarins (Fig. 2b and Extended Data Table 2).
These observations suggest that the initial pummelo introgression into the mandarin gene pool may have involved as few as one pummelo tree (carrying both P1 and P2 haplotypes), the contribution of which was diluted by repeated backcrosses with mandarins (Supplementary Note 6.3). The introgressed pummelo haplotypes became widespread and gave rise to type-2 (early-admixture) mandarins (Fig. 2b). We propose that later, additional pummelo introgressions gave rise to type-3 (late-admixture) mandarins and sweet orange, and that some modern type-3 mandarins were derived from hybridizations among existing mandarins and sweet orange. This late-admixture model for type-3 mandarins is consistent with the historical records for Clementine and Kiyomi (both mandarin–sweet orange hybrids), and for W. Murcott, Wilking and Fallglo (hybrids involving other type-3 mandarins), whereas definitive records for the remaining two late-admixture mandarins (King and Satsuma) are not available.
Domestication of mandarins and sweet orange
Citrus domestication probably began with the identification and asexual propagation of selected, possibly hybrid or admixed individuals, rather than recurrent selection from a breeding population as for annual crops37,38. Additional diversity was obtained by capturing somatic mutations that occur within a relatively few basic genotypes. Therefore, conventional approaches to identifying selective pressures under recurrent breeding39 cannot be applied. We can, however, use genome sequences to infer some features of the early stages of citrus domestication. Here we focus on mandarins, a class of citrus comprising small and easily peeled fruits that are of high commercial value.
All 28 mandarin accessions, except for Tachibana, exhibit an extensive network of relatedness (with a coefficient of relatedness, r > 1/8), and all but four mandarins (three of the four are pure or type-1 mandarins) show second degree or higher relatedness (r > 1/4) to at least one (mean = 7) other mandarin (Fig. 3a and Supplementary Note 7). By contrast, sequenced pummelos and citrons appear to be independent selections from relatively large populations. In the absence of historical records for most mandarins, the actual kinships are difficult to infer, owing to extensive haplotype sharing among the ancestors, although some parent–child pairs can be identified. In addition to confirming, using the whole-genome sequence, the parentage of Wilking (King–Willowleaf), Kiyomi (Satsuma–sweet orange) and Fallglo (one grandparent is Clementine), we find parent–child relationships between two pairs of mandarins (Ponkan is a parent of Dancy; Huanglingmiao (a somatic mutant of Kishu) is a parent of Satsuma)34, in addition to the previously established parent–child pair of Willowleaf and Clementine mandarins2. Additional parent–child pairs involving the recently sequenced Chinese mandarins3 are also identified (Supplementary Note 7.3). A few cultivar types in this network (Satsuma, Dancy, Clementine, Kiyomi, Fallgo and the Chinese cultivar BTJ mandarins) have marked signs of inbreeding, indicated by runs of homozygosity (Extended Data Fig. 4a) as a result of shared haplotypes between their parents. The high degree of relatedness among mandarins implies extensive sharing of C. reticulata haplotypes.
a, Genetic relatedness among 48 citrus accessions derived from four progenitor species including citrons, pummelos, pure mandarins and micrantha. Solid lines connect pairs with coefficient of relatedness r > 0.45, with parent–child pairs denoted by arrows pointing from parent to child. Dashed and dotted lines correspond to 0.35 <r <0.45 and 0.25 <r <0.35, respectively. Mandarins are distinguished from other taxonomic groups by an extensive relatedness network, indicating shared haplotypes in the ancestral gene pool. Three-letter code as in Fig. 1, see Supplementary Table 2 for details. b, Shown in decreasing order are the values of coefficient of relatedness between sweet orange and other accessions, with C. maxima (rP) and C. reticulata (rM) components in green and light salmon, respectively.There is significant haplotype sharing between sweet orange and all mandarins, except for three of the type-1 mandarins. Five accessions (Clementine and Kiyomi mandarins, eremorange, Marsh grapefruit, and Ambersweet orange) have sweet orange as the male parent.
Sweet orange also shows extensive haplotype sharing at the level of r > 0.1 with 25 of the 28 sequenced mandarins (except for three pure or type-1 mandarins; Fig. 3b and Extended Data Fig. 4b). Two late-admixture mandarins (Clementine and Kiyomi) are direct offspring of sweet orange. Among the early-admixture (type-2) mandarins, Ponkan shows the highest affinity to sweet orange2 with r ≈ 0.36. Even the pure mandarin, Sun Chu Sha Kat has r ≈ 0.23, equivalent to second degree relatedness to sweet orange. We can rule out the scenario that sweet orange is the common ancestor of the mandarins, because of a lack of pummelo haplotypes (derived from sweet orange) among the mandarins. Rather, the extensive C. reticulata haplotype sharing between sweet orange and mandarins suggests that the mandarin parent of sweet orange was part of an expansive network of relatedness among mandarins.
Because our collection of mandarins represents a diverse set of both ancient and modern varieties, including economically important accessions with mostly unknown parentage, the presence of an extensive relatedness network was not anticipated a priori. The shared C. reticulata haplotypes are suggestive of and consistent with signatures of the human selection process, during which mandarins with desirable traits were necessarily maintained through clonal propagation (nucellar polyembryony or grafting). Although one cannot preclude the possibility that the relatedness network was initiated before domestication from a small number of founder trees, human selection of accessions resulting from natural hybridization probably had a key role in the process of domestication that eventually led to the extensive relatedness network observed today. For example, modern mandarins, such as Clementine and W. Murcott, are known to be selections from chance seedlings found in Algeria40 and Morocco2, at the onset and middle of the last century, respectively.
Pummelo admixture is correlated with fruit size and acidity, suggesting a role for pummelo introgression in citrus domestication. As both fruit size and acidity profile for the most recently sequenced accessions3 are not described, we used 37 citrus accessions in this analysis. We find that the fruit sizes of mandarins, oranges, grapefruit and pummelos show a strong positive correlation (Pearson correlation coefficient r = 0.88) with pummelo admixture proportion (Extended Data Fig. 5a, b and Supplementary Note 10.1). In addition to fruit size, a pivotal driver of fruit domestication is palatability, a characteristic that in citrus requires low to moderate levels of acidity. In mandarins, palatability appears to be linked to pummelo introgression at a major locus at the start of chromosome 8 (0.3–2.2 Mb), where all nine known palatable mandarins, but none of the four known acidic mandarins, show pummelo admixture in at least part of the genomic region (Extended Data Fig. 3). This locus is also found to be significant in a genome scan for palatability association (Extended Data Fig. 5c, d and Extended Data Table 3) and contains several potentially relevant genes (Supplementary Note 10.2). Among these genes is a gene encoding the mitochondrial NAD+-dependent isocitrate dehydrogenase (IDH) which regulates citric-acid synthesis41 (Extended Data Table 4).
Our study finds that domesticated citrus fruit crops, such as mandarins and sweet orange, experienced a complex history of admixture, conceptually similar to those well-recognized in annual crops, such as rice42 and maize43, and in other fruit trees, such as apple44 and grape45, for which the current genomic diversity is linked to widespread ancient introgression. Other cultivated citrus groups, the interspecific F1 hybrids in particular, originated from hybridizations of two pure parental species. Several of these involve C. medica (citron), including limes and lemons10. A unique and critical characteristic of the three pivotal species (C. maxima, C. reticulata and C. medica) that gave rise to most cultivated citrus fruits is the occurrence of a complex floral anatomy (Extended Data Fig. 6), thus leading to the development of more complex fruit. Other species were also involved in hybridizations, including Fortunella and C. micrantha. Distinct from the mandarin lineages, these hybrids are characterized by their acidic fruit, and their selection must have been made on the basis of other characteristics, such as a sweet edible peel and aroma2, respectively.
Conclusion
On the basis of genomic, phylogenetic and biogeographic analyses of 60 diverse citrus and related accessions, we propose that the centre of origin of citrus species was the southeast foothills of the Himalayas, in a region that includes the eastern area of Assam, northern Myanmar and western Yunnan. Our analyses suggest that the ancestral citrus species underwent a sudden speciation event during the late Miocene. This radiation coincided with a pronounced transition from wet monsoon conditions to a drier climate, as observed in nearby areas in many other plant and animal lineages. The Australian citrus species and Tachibana, a native Japanese mandarin, split later from mainland citrus during the early Pliocene and Pleistocene, respectively. By distinguishing between pure species, hybrids and admixtures, we could trace the genealogy and genetic origin of the major citrus commercial cultivars. Both the extensive relatedness network among mandarins and sweet orange, and the association of pummelo admixture with desirable fruit traits suggest a complex domestication process.
Our work challenges previous proposals for citrus taxonomy. For example, we find that several named genera (Fortunella, Eremocitrus and Microcitrus) are in fact nested within the citrus clade. These and other distinct clades that we have identified are therefore more appropriately considered species within the genus Citrus, on a par with those that formerly were referred to as the three ‘true’ or ‘biological’ species (C. reticulata, C. maxima and C. medica). Additionally, the related genus, Poncirus, a subject of continuous controversy since it was originally proposed to be within the genus Citrus12,46, is clearly a distinct clade that is separate from Citrus based on sequence divergence and whole-genome phylogeny.
In summary, this work presents insights into the origin, evolution and domestication of citrus, and the genealogy of the most important wild and cultivated varieties. Taken together, these findings draw a new evolutionary framework for these fruit crops, a scenario that challenges current taxonomic and phylogenetic thoughts, and points towards a reformulation of the genus Citrus.
Methods
Sample collection and sequencing
Whole-genome sequences from a total of 60 accessions were analysed: 58 citrus accessions with different geographical origins and two representative outgroup genera. Twelve of these genomes, including five mandarins, four pummelos, two oranges and a wild Mangshan mandarin (C. mangshanensis) were reanalysed from previous works2,7. We also reanalysed 19 genomes from Chinese collections, including 15 unnamed mandarins, 2 Chinese sour oranges, Ambersweet orange and Cocktail grapefruit (a hybrid resembling grapefruit) that have been previously reported3.
The 30 accessions that were newly sequenced came from citrus germ-plasm banks and collections at IVIA, Valencia, Spain; SRA, Corse, France; UCR, Riverside and FDACS/DPI, Florida and included nine mandarins, two limes, one rough lemon, one grapefruit, one lemon, four citrons, one Australian desert lime, one eremorange, two Australian finger limes, two Australian round limes, one kumquat, one calamondin, one micrantha, one Ichang papeda, one trifoliate orange and one Chinese box orange (Supplementary Note 1).
DNA libraries were constructed using standard protocols with some modifications. Library insert sizes range from 325 to 500 bp. Sequencing was performed on HiSeq2000/2500 instruments using 100-bp paired-end reads. Primary analysis of the data included quality control on the Illumina RTA sequence analysis pipeline (Supplementary Note 2).
Variant calls and Citrus species diversity
Illumina paired-end reads were aligned to the haploid Clementine reference sequence2 and the sweet orange chloroplast genome assembly47 using bwa-mem48. PCR duplicates were removed using Picard. Raw variants were called using GATK HaplotypeCaller49 with subsequent filtering based on read map quality score, base quality score, read depth and so on (Supplementary Note 3.1).
Interspecific admixtures versus pure citrus species were distinguished based on sliding window analysis of heterozygosity and pairwise genetic distance D (Supplementary Note 4). Genome-wide ancestry informative markers for the progenitor species were derived using pure accessions. Admixture analysis was carried out in sliding windows using ancestry informative markers (Supplementary Notes 5).
Citrus relatedness and haplotype sharing
Interspecific phasing was used to extract admixed haplotypes. Identical-by-descent sharing was calculated for each of the non-overlapping sliding windows across the genome and used to estimate coefficient of relatedness among citrus accessions (Supplementary Notes 6, 7).
Phylogeny and speciation dating
We used Chinese box orange (genus Severinia) as an outgroup. Time calibration is based on the C. linczangensis16 fossil from Lincang, Yunnan, China. MrBayes50 was used for whole genome Bayesian phylogenetic inference, and corroborated with a PhyML51 reconstructed maximum likelihood tree. A penalized likelihood method52 as implemented in APE53 was used to construct the chronogram (Supplementary Note 8).
Genome scan of palatability association
We used a mixed linear model as implemented in gemma54 for a case–control study of citrus acidity and palatability with 37 citrus accessions. A conservative Bonferroni correction was used to select significant genomic loci, with subsequent manual examination of each candidate variant in all accessions to identify most discriminatory loci for fruit palatability (Supplementary Note 10).
Data availability
Whole-genome shotgun-sequencing data generated in this study have been deposited at NCBI under BioProject PRJNA414519. Prior resequencing data analysed here can be accessed under BioProject accession numbers PRJNA320985 (mandarins) and PRJNA321100 (oranges), and also under the NCBI Sequence Read Archive accession codes SRX372786 (sour orange), SRX372703 (sweet orange), SRX372702 (low-acid pummelo), SRX372688 (Chandler pummelo), SRX372685 (Willowleaf mandarin), SRX372687 (W. Murcott mandarin), SRX372665 (Ponkan mandarin) and SRX371962 (Clementine mandarin). The Clementine reference sequence used here is available at https://phytozome.jgi.doe.gov/.
Accession codes
References
Swingle, W. T. & Reece, P. C. in The Citrus Industry, revised 2nd edn, History, World Distribution, Botany, and Varieties Vol. 1 (eds Reuther, W. et al.) 190–430 (Univ. California, 1967)
Wu, G. A. et al. Sequencing of diverse mandarin, pummelo and orange genomes reveals complex history of admixture during citrus domestication. Nat. Biotechnol. 32, 656–662 (2014)
Wang, X. et al. Genomic analyses of primitive, wild and cultivated citrus provide insights into asexual reproduction. Nat. Genet. 49, 765–772 (2017)
Mudge, K., Janick, J., Scofield, S. & Goldschmidt, E. E. A history of grafting. Hortic. Rev. (Am. Soc. Hortic. Sci.) 35, 437–493 (2009)
Swingle, W. in The Citrus Industry, History Botany and Breeding Vol. 1 (eds Webber, H. J. & Batchelor, L. D. ) 129–474 (Univ. California Press, 1943)
Tanaka, T. Species Problem in Citrus (Japanese Society for Promotion of Science, 1954)
Xu, Q. et al. The draft genome of sweet orange (Citrus sinensis). Nat. Genet. 45, 59–66 (2013)
Barrett, H. & Rhodes, A. A numerical taxonomic study of affinity relationships in cultivated Citrus and its close relatives. Syst. Bot. 1, 105–136 (1976)
Scora, R. W. On the history and origin of Citrus. Bull. Torrey Bot. Club 102, 369–375 (1975)
Curk, F. et al. Phylogenetic origin of limes and lemons revealed by cytoplasmic and nuclear markers. Ann. Bot. 117, 565–583 (2016)
Nicolosi, E. et al. Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor. Appl. Genet. 100, 1155–1166 (2000)
Bayer, R. J. et al. A molecular phylogeny of the orange subfamily (Rutaceae: Aurantioideae) using nine cpDNA sequences. Am. J. Bot. 96, 668–685 (2009)
Jacques, F. M. et al. Late Miocene southwestern Chinese floristic diversity shaped by the southeastern uplift of the Tibetan Plateau. Palaeogeogr. Palaeoclimatol. Palaeoecol. 411, 208–215 (2014)
Jacques, F. M. et al. Quantitative reconstruction of the Late Miocene monsoon climates of southwest China: a case study of the Lincang flora from Yunnan Province. Palaeogeogr. Palaeoclimatol. Palaeoecol. 304, 318–327 (2011)
Gmitter, F. G. & Hu, X. L. The possible role of Yunnan, China, in the origin of contemporary citrus species (Rutaceae). Econ. Bot. 44, 267–277 (1990)
Xie, S., Manchester, S. R., Liu, K., Wang, Y. & Sun, B. Citrus linczangensis sp n., a leaf fossil of Rutaceae from the late Miocene of Yunnan, China. Int. J. Plant Sci. 174, 1201–1207 (2013)
Clift, P. D., Wan, S. & Blusztajn, J. Reconstructing chemical weathering, physical erosion and monsoon intensity since 25 Ma in the northern South China Sea: a review of competing proxies. Earth Sci. Rev. 130, 86–102 (2014)
Valdiya, K. S. Emergence and evolution of Himalaya: reconstructing history in the light of recent studies. Prog. Phys. Geogr. 26, 360–399 (2002)
Wen, J., Zhang, J. Q., Nie, Z. L., Zhong, Y. & Sun, H. Evolutionary diversifications of plants on the Qinghai–Tibetan Plateau. Front. Genet. 5, 4 (2014)
Favre, A. et al. The role of the uplift of the Qinghai–Tibetan Plateau for the evolution of Tibetan biotas. Biol. Rev. Camb. Philos. Soc. 90, 236–253 (2015)
Pfeil, B. E. & Crisp, M. D. The age and biogeography of Citrus and the orange subfamily (Rutaceae: Aurantioideae) in Australasia and New Caledonia. Am. J. Bot. 95, 1621–1631 (2008)
Beattie, G. A. C., Holford, P., Mabberley, D. J., Haigh, A. M. & Broadbent, P. in On the origins of Citrus, Huanglongbing, Diaphorina citri and Trioza erytreae. International Conference of Huanglongbing (eds Gottwald, T. R. & Graham, J. H. ) 25–57 (Plant Management Network, 2009)
Thomas, D. C. et al. West to east dispersal and subsequent rapid diversification of the mega-diverse genus Begonia (Begoniaceae) in the Malesian archipelago. J. Biogeogr. 39, 98–113 (2012)
Richardson, J. E., Costion, C. M. & Muellner, A. N. in Biotic Evolution and Environmental Change in Southeast Asia Ch. 6 (eds Gower, D. et al.) 138–163 (Cambridge Univ. Press, 2012)
van Welzen, P. C., Slik, J. W. F. & Alahuhta, J. Plant distribution patterns and plate tectonics in Malesia. Biol. Skr. 55, 199–217 (2005)
Hall, R. Southeast Asia’s changing palaeogeography. Blumea 54, 148–161 (2009)
Zhang, W. Thirty years achievements in citrus varietal improvement in China in Proc. International Citrus Congress (ed. Matsumoto, K. ) 51–53 (International Society of Citriculture, 1982–1983)
Brophy, J. J., Goldsack, R. J. & Forster, P. I. The leaf oils of the Australian species of Citrus (Rutaceae). J. Essent. Oil Res. 13, 264–268 (2001)
Tanaka, T. The discovery of Citrus tachibana in Formosa, and its scientific and industrial significance. Studia Citrologica 5, 1–20 (1931)
Gibbard, P. & Cohen, K. M. Global chronostratigraphical correlation table for the last 2.7 million years. Episodes 31, 243–247 (2008)
Chiang, T.-Y. & Schaal, B. A. Phylogeography of plants in Taiwan and the Ryukyu Archipelago. Taxon 55, 31–41 (2006)
Voris, H. K. Maps of Pleistocene sea levels in southeast Asia: shorelines, river systems and time durations. J. Biogeogr. 27, 1153–1167 (2000)
Huang, S.-F. Hypothesizing origin, migration routes and distribution patterns of gymnosperms in Taiwan. Taiwania 59, 139–163 (2014)
Shimizu, T. et al. Hybrid origins of citrus varieties inferred from DNA marker analysis of nuclear and organelle genomes. PLoS ONE 11, e0166969 (2016)
Hirai, M., Mitsue, S., Kita, K. & Kajiura, I. A survey and isozyme analysis of wild mandarin, Tachibana (Citrus tachibana (Mak.) Tanaka) growing in Japan. J. Jpn. Soc. Hortic. Sci. 59, 1–7 (1990)
Hagerty, M. J. Han Yen-Chih’s Chü lu (monograph on the oranges of Wên-chou, Chekiang). Toung Pao 22, 63–96 (1923)
Miller, A. J. & Gross, B. L. From forest to field: perennial fruit crop domestication. Am. J. Bot. 98, 1389–1414 (2011)
Gaut, B. S., Díez, C. M. & Morrell, P. L. Genomics and the contrasting dynamics of annual and perennial domestication. Trends Genet. 31, 709–719 (2015)
Hamblin, M. T., Buckler, E. S. & Jannink, J. L. Population genetics of genomics-based crop improvement methods. Trends Genet. 27, 98–106 (2011)
Trabut, J. L’hybridation des Citrus: une nouvelle tangérine ‘la Clémentine’. Revue Horticole 10, 232–234 (1902)
Meléndez-Hevia, E., Waddell, T. G. & Cascante, M. The puzzle of the Krebs citric acid cycle: assembling the pieces of chemically feasible reactions, and opportunism in the design of metabolic pathways during evolution. J. Mol. Evol. 43, 293–303 (1996)
Gross, B. L. & Zhao, Z. Archaeological and genetic insights into the origins of domesticated rice. Proc. Natl Acad. Sci. USA 111, 6190–6197 (2014)
Hufford, M. B. et al. The genomic signature of crop-wild introgression in maize. PLoS Genet. 9, e1003477 (2013)
Cornille, A. et al. New insight into the history of domesticated apple: secondary contribution of the European wild apple to the genome of cultivated varieties. PLoS Genet. 8, e1002703 (2012)
Myles, S. et al. Genetic structure and domestication history of the grape. Proc. Natl Acad. Sci. USA 108, 3530–3535 (2011)
Nesom, G. L. Citrus trifoliata (Rutaceae): review of biology and distribution in the USA. Phytoneuron 46, 1–14 (2014)
Bausher, M. G., Singh, N. D., Lee, S. B., Jansen, R. K. & Daniell, H. The complete chloroplast genome sequence of Citrus sinensis (L.) Osbeck var ‘Ridge Pineapple’: organization and phylogenetic relationships to other angiosperms. BMC Plant Biol. 6, 21 (2006)
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrow–Wheeler transform. Bioinformatics 26, 589–595 (2010)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012)
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010)
Sanderson, M. J. Estimating absolute rates of molecular evolution and divergence times: a penalized likelihood approach. Mol. Biol. Evol. 19, 101–109 (2002)
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004)
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824 (2012)
Frost, H. B. & Soost, R. K. in The Citrus Industry Vol. 2 (eds Reuther, W. et al.) 290–324 (1968)
Acknowledgements
Please see Supplementary Note 11 for funding information.
Author information
Authors and Affiliations
Contributions
M.T., D.S.R. and G.A.W. developed the project and acted as project coordinators and provided scientific leadership; G.A.W. developed methods for admixture analysis and interspecific phasing, and performed comparative genome analysis. J.T., V.I., A.L.-G., E.P.-R., C.B., C.D., F.R.T., J.C.-C., R.A., J.D. and M.T. contributed 26 genomes; J.T., J.C.-C., R.A. and J.D. provided bioinformatics support; J.T. and E.P.-R. contributed to the study of the IDH gene; V.I., E.P.-R. and C.B. contributed to the variant analysis of candidate genes using genome-wide association studies; A.L.-G. and C.B. assisted in the biogeographic study; A.L.-G. and F.G.G. contributed to the description of citrus accessions and discriminatory characteristics; P.O. and F.C. contributed to germplasm, admixture analysis and hypothesis on the origin of cultivated citrus species; D.D. and F.G.G. contributed one citrus genome; M.L.R. contributed seven citrus genomes; F.G.G. contributed perspective garnered from more than 35 years of experience working on the genetic improvement of citrus; G.A.W., M.T., D.S.R. and F.G.G. wrote the manuscript; G.A.W. and M.T. contributed the hypothesis on the origin and dispersal of citrus.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. Ross-Ibarra, P. Wincker and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Citrus biogeography and phylogeny.
a, Geographical distribution of the genus Citrus in southeast Asia and Australia. Distribution is based on documented reports on the presence of wild genotypes representative of pure citrus species (blue numbers), admixtures (red numbers) and relevant interspecific hybrids (black numbers), growing freely in non-cultivated areas. Numbers are as in Supplementary Table 1.1, 2, Fortunella spp. ; 3, Australian citrus (E. glauca; Microcitrus australasica; Microcitrus australis); 4, C. micrantha; 5–8, C. ichangensis; 9–15, C. medica; 16–19, C. maxima; 20–22, C. reticulata (Sun Chu Sha Kat); 23–25, C. tachibana; 26, C. mangshanensis; 27–29, Citrus spp. (mandarins); 30–33, C. sinensis; 34, 35, Citrus limon (probably not truly wild genotypes); 36, 37, Citrus limonia; 38, Citrus jambhiri; 39–42, C. aurantium; 43, Citrus aurantifolia (probably not truly wild genotypes); 44, 45, Fortunella and C. reticulata hybrid. The red star indicates the location of the C. linczangensis fossil from the Late Miocene of Lincang16. b, Citrus chloroplast genome phylogeny rooted with Severinia. The analysis was performed on 58 citrus accessions and 2 outgroup genera, Poncirus and Severinia. The maximum likelihood tree as inferred from PhyML is shown. Percentage statistical support for the nodes is based on 200 bootstrap replicates. c, Citrus nuclear genome phylogeny rooted with Severinia. Both Bayesian and maximum likelihood trees yield the same topology with highly supported branches. The maximum likelihood tree reconstructed from PhyML is shown. Branch statistical support is based on 1,000 bootstraps and is shown if it is less than 100%. All branches have posterior probability 1.0 with Bayesian inference using MrBayes (not shown).
Extended Data Figure 2 Segmental ancestry and admixture in citrus.
a, Segmental ancestry of 46 citrus accessions derived from the three progenitor species of C. medica (CI), C. maxima (PU) and C. reticulata (MA). UNK, unknown. Mandarins are divided into three types with type-1 representing pure mandarins. Types 2 and 3 are determined by the pummelo admixture pattern. b, Segmental ancestry of an Australian finger lime. Blue segments denote pure finger lime (genotype: F/F), and orange segments have Australian round lime admixture (genotype: F/R). Genomic regions are coloured in grey if segmental ancestry cannot be determined.
Extended Data Figure 3 Pattern of pummelo introgression in mandarins.
a, Distinct admixed pummelo haplotypes among mandarins, oranges and grapefruit are shown in different colours; the C. reticulata haplotypes are masked. The admixture pattern separates the mandarins into three groups, with type-1 representing pure mandarins. Type-2 mandarins contain a small amount of pummelo admixture derived from two C. maxima haplotypes: P1 (light blue colour) and P2 (dark blue), suggesting as few as one common pummelo ancestor in the distant past. Type-3 mandarins are characterized by both marked pummelo admixture and additional pummelo haplotypes besides P1 and P2. b, Haplotype trees for two chromosome segments where pummelo haplotypes of type-2 mandarins are in green. Left, haplotype tree for chr3:3.2–5.2 Mb. Sweet orange, sour orange, and twelve of the sequenced mandarins are interspecific hybrids, and their phased C. maxima and C. reticulata haplotypes are denoted by prepending, respectively, ‘p’ and ‘m’ to the corresponding accession codes. The nine type-2 mandarins share the same pummelo haplotype (P1). Right, the haplotype tree for chr2:31.4–33.4 Mb. Two pummelo haplotypes (P1, P2) are shared among seven type-2 mandarins, with Ponkan mandarin containing both P1 and P2. Sweet orange also carries two pummelo haplotypes at this locus, denoted by pC.SWO (shared with Clementine) and pA.SWO (alternate haplotype).
Extended Data Figure 4 Haplotype sharing in mandarins.
a, Runs of homozygosity in mandarins. Heterozygosity is plotted in non-overlapping windows of 200 kb along the nine chromosomes of 7 mandarin accessions with the highest degree of inbreeding. Runs of homozygosity correspond to regions with zero heterozygosity as a result of haplotype sharing between the parents. b, Haplotype sharing between sweet orange and mandarins. The two haplotypes of sweet orange are denoted by hapC (transmitted to Clementine) and hapA (alternate), respectively. The hapC haplotype is coloured in red (denoted by MC) or dark green (denoted by PC) if its genetic ancestry is C. reticulata or C. maxima, respectively. Similarly, hapA can take the form of C. reticulata (MA in orange colour) or C. maxima (PA in light green) depending on its genetic ancestry. Shared sweet orange haplotypes in mandarins are coloured accordingly, except when both haplotypes of sweet orange are shared (IBD2) either as two C. reticulata haplotypes (MAC, dark red) or as interspecific hybrid (P + M, blue). Regions coloured in grey denote the absence of haplotype sharing between sweet orange and mandarin.
Extended Data Figure 5 Fruit size and acidity correlated with pummelo introgression.
a, Fruit size strongly correlated with pummelo admixture. The diameters of mandarins, oranges and grapefruit are plotted against the corresponding pummelo admixture proportions. A simple linear regression is shown in red. The strong correlation (Pearson correlation coefficient r = 0.88) between fruit size and pummelo admixture is apparent, especially among the three taxonomic groups of mandarins, oranges and grapefruit. The outliers (Ponkan mandarin and four acidic mandarins) suggest that certain genomic loci could be more important than others in fruit size determination. Accessions without size information are not included. Data are mean ± s.d. from a set of 25 measurements for each of the 15 accessions. b, Fruit size correlation with pummelo allelic proportion with the addition of two pummelos. A polynomial regression provides a better fit than simple linear regression (adjusted R2 = 0.92). Data are mean ± s.d. from a set of 25 measurements for each of the 17 accessions. c, Genome scan of significant loci associated with citrus acidity. Manhattan plot of a case–control analysis of a genome-wide association study (GWAS) of n = 37 citrus accessions with known acidity profile. The horizontal dashed line denotes the conservative Bonferoni-corrected P value of 7.9 × 10−8 for genome-wide significance (α= 0.05). d, Manual inspection of candidate regions identified by GWAS (n = 37 accessions) demonstrates that in addition to the locus at chr1:23512067, single-nucleotide polymorphisms located in chromosome 8 are discriminatory for acidity. Shown are 40-kb zoom windows containing focal single-nucleotide polymorphisms (red vertical lines and green arrows) and gene models depicted by blue boxes in these two regions. TOE1, target of EGR1 protein 1 (Ciclev10007611; https://phytozome.jgi.doe.gov/pz/portal.html); NAD+-IDH, NAD+-dependent isocitrate dehydrogenase (IDH, Ciclev10028714); GH10, glycosyl hydrolase family 10 protein (Ciclev10028121) and MTOR, serine/threonine protein kinase (Ciclev10027661).
Extended Data Figure 6 Species characteristics of citrus.
Reproductive and vegetative characteristics among several species of the genus Citrus and related genera according to refs 1 and 55. The tree topology represented is that of the chronogram shown in Fig. 1c and citrus fruit images are not drawn to scale. Most mandarins in our collection are polyembryonic, though a few are monoembryonic, including Clementine. Other exceptions to the generalized description concerning embryo numbers, in kumquat and other citrus species, can also be found.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-11, Supplementary Tables 1-4 and Supplementary References. (PDF 1426 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) licence. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, G., Terol, J., Ibanez, V. et al. Genomics of the origin and evolution of Citrus. Nature 554, 311–316 (2018). https://doi.org/10.1038/nature25447
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25447
This article is cited by
-
Genome-wide identification of the GRF family in sweet orange (Citrus sinensis) and functional analysis of the CsGRF04 in response to multiple abiotic stresses
BMC Genomics (2024)
-
Identification and diversity analysis of interspecific citrus rootstock hybrids with combination of morphological traits and microsatellite markers
Horticulture, Environment, and Biotechnology (2024)
-
Evaluation of Leaf Morpho-anatomical and Gas Exchange Parameters in Relation to Fruit Yield in Indian Subtropical Grapefruit (Citrus paradisi Macf.) Varieties
Applied Fruit Science (2024)
-
A tetraploid-dominated cytochimera developed from a natural bud mutant of the nonapomictic mandarin variety ‘Orah’
Molecular Breeding (2024)
-
Green solutions and new technologies for sustainable management of fungus and oomycete diseases in the citrus fruit supply chain
Journal of Plant Pathology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.