Abstract
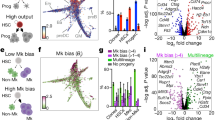
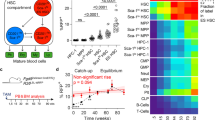
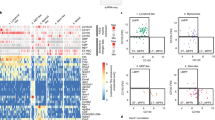
Haematopoiesis, the process of mature blood and immune cell production, is functionally organized as a hierarchy, with self-renewing haematopoietic stem cells and multipotent progenitor cells sitting at the very top1,2. Multiple models have been proposed as to what the earliest lineage choices are in these primitive haematopoietic compartments, the cellular intermediates, and the resulting lineage trees that emerge from them3,4,5,6,7,8,9,10. Given that the bulk of studies addressing lineage outcomes have been performed in the context of haematopoietic transplantation, current models of lineage branching are more likely to represent roadmaps of lineage potential than native fate. Here we use transposon tagging to clonally trace the fates of progenitors and stem cells in unperturbed haematopoiesis. Our results describe a distinct clonal roadmap in which the megakaryocyte lineage arises largely independently of other haematopoietic fates. Our data, combined with single-cell RNA sequencing, identify a functional hierarchy of unilineage- and oligolineage-producing clones within the multipotent progenitor population. Finally, our results demonstrate that traditionally defined long-term haematopoietic stem cells are a significant source of megakaryocyte-restricted progenitors, suggesting that the megakaryocyte lineage is the predominant native fate of long-term haematopoietic stem cells. Our study provides evidence for a substantially revised roadmap for unperturbed haematopoiesis, and highlights unique properties of multipotent progenitors and haematopoietic stem cells in situ.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Morrison, S. J., Wandycz, A. M., Hemmati, H. D., Wright, D. E. & Weissman, I. L. Identification of a lineage of multipotent hematopoietic progenitors. Development 124, 1929–1939 (1997)
Morrison, S. J. & Weissman, I. L. The long-term repopulating subset of hematopoietic stem cells is deterministic and isolatable by phenotype. Immunity 1, 661–673 (1994)
Adolfsson, J. et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential: a revised road map for adult blood lineage commitment. Cell 121, 295–306 (2005)
Akashi, K., Traver, D., Miyamoto, T. & Weissman, I. L. A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature 404, 193–197 (2000)
Ceredig, R., Rolink, A. G. & Brown, G. Models of haematopoiesis: seeing the wood for the trees. Nat. Rev. Immunol. 9, 293–300 (2009)
Forsberg, E. C., Serwold, T., Kogan, S., Weissman, I. L. & Passegué, E. New evidence supporting megakaryocyte–erythrocyte potential of flk2/flt3+ multipotent hematopoietic progenitors. Cell 126, 415–426 (2006)
Notta, F. et al. Distinct routes of lineage development reshape the human blood hierarchy across ontogeny. Science 351, aab2116 (2016)
Perié, L., Duffy, K. R., Kok, L., de Boer, R. J. & Schumacher, T. N. The branching point in erythro-myeloid differentiation. Cell 163, 1655–1662 (2015)
Pietras, E. M. et al. Functionally distinct subsets of lineage-biased multipotent progenitors control blood production in normal and regenerative conditions. Cell Stem Cell 17, 35–46 (2015)
Yamamoto, R. et al. Clonal analysis unveils self-renewing lineage-restricted progenitors generated directly from hematopoietic stem cells. Cell 154, 1112–1126 (2013)
Sun, J. et al. Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014)
Debili, N. et al. Characterization of a bipotent erythro-megakaryocytic progenitor in human bone marrow. Blood 88, 1284–1296 (1996)
Sanada, C. et al. Adult human megakaryocyte–erythroid progenitors are in the CD34+CD38mid fraction. Blood 128, 923–933 (2016)
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015)
Pronk, C. J. et al. Elucidation of the phenotypic, functional, and molecular topography of a myeloerythroid progenitor cell hierarchy. Cell Stem Cell 1, 428–442 (2007)
Velten, L. et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 19, 271–281 (2017)
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015)
Calaminus, S. D. et al. Lineage tracing of Pf4-Cre marks hematopoietic stem cells and their progeny. PLoS ONE 7, e51361 (2012)
Gekas, C. & Graf, T. CD41 expression marks myeloid-biased adult hematopoietic stem cells and increases with age. Blood 121, 4463–4472 (2013)
Haas, S. et al. Inflammation-induced emergency megakaryopoiesis driven by hematopoietic stem cell-like megakaryocyte progenitors. Cell Stem Cell 17, 422–434 (2015)
Nishikii, H. et al. Unipotent megakaryopoietic pathway bridging hematopoietic stem cells and mature megakaryocytes. Stem Cells 33, 2196–2207 (2015)
Roch, A., Trachsel, V. & Lutolf, M. P. Single-cell analysis reveals cell division-independent emergence of megakaryocytes from phenotypic hematopoietic stem cells. Stem Cells 33, 3152–3157 (2015)
Sanjuan-Pla, A. et al. Platelet-biased stem cells reside at the apex of the haematopoietic stem-cell hierarchy. Nature 502, 232–236 (2013)
Vannucchi, A. M. et al. Identification and characterization of a bipotent (erythroid and megakaryocytic) cell precursor from the spleen of phenylhydrazine-treated mice. Blood 95, 2559–2568 (2000)
Busch, K. et al. Fundamental properties of unperturbed haematopoiesis from stem cells in vivo. Nature 518, 542–546 (2015)
Sawai, C. M. et al. Hematopoietic stem cells are the major source of multilineage hematopoiesis in adult animals. Immunity 45, 597–609 (2016)
Junker, J. P. et al. Massively parallel clonal analysis using CRISPR/Cas9 induced genetic scars. Preprint at https://doi.org/10.1101/056499 (2017)
Raj, B . et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain by scGESTALT. Preprint at https://doi.org/10.1101/205534 (2017)
Foudi, A. et al. Analysis of histone 2B-GFP retention reveals slowly cycling hematopoietic stem cells. Nat. Biotechnol. 27, 84–90 (2009)
Oguro, H., Ding, L. & Morrison, S. J. SLAM family markers resolve functionally distinct subpopulations of hematopoietic stem cells and multipotent progenitors. Cell Stem Cell 13, 102–116 (2013)
Wilson, A. et al. Hematopoietic stem cells reversibly switch from dormancy to self-renewal during homeostasis and repair. Cell 135, 1118–1129 (2008)
Harkey, M. A. et al. Multiarm high-throughput integration site detection: limitations of LAM-PCR technology and optimization for clonal analysis. Stem Cells Dev. 16, 381–392 (2007)
Wang, G. P. et al. DNA bar coding and pyrosequencing to analyze adverse events in therapeutic gene transfer. Nucleic Acids Res. 36, e49 (2008)
Zilionis, R. et al. Single-cell barcoding and sequencing using droplet microfluidics. Nat. Protocols 12, 44–73 (2017)
Ilicic, T. et al. Classification of low quality cells from single-cell RNA-seq data. Genome Biol. 17, 29 (2016)
Weinreb, C., Wolock, S. & Klein, A. SPRING: a kinetic interface for visualizing high dimensional single-cell expression data. Preprint at https://doi.org/10.1101/090332 (2016)
Buettner, F. et al. Computational analysis of cell-to-cell heterogeneity in single-cell RNA-sequencing data reveals hidden subpopulations of cells. Nat. Biotechnol. 33, 155–160 (2015)
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016)
Hennig, C. Cluster validation by measurement of clustering characteristics relevant to the user. Preprint at http://arXiv:1703.09282 (2017)
Acknowledgements
We are grateful to members of the Camargo and Klein laboratory for comments. A.R.F. is a Merck Fellow of the Life Sciences Research Foundation and a non-stipendiary European Molecular Biology Organization postdoctoral fellow. This work was supported by National Institutes of Health grants HL128850-01A1 and P01HL13147 to F.D.C. F.D.C. is a Leukemia and Lymphoma Society and a Howard Hughes Medical Institute Scholar. A.M.K. is supported by a Burroughs-Wellcome Fund CASI award, and by the Edward J. Mallinckrodt Fellowship.
Author information
Authors and Affiliations
Contributions
A.R.F. and F.D.C. designed the study, analysed the data, and wrote the manuscript. A.R.F. performed and analysed the experiments, assisted by M.J., S.P., and J.S. S.W., C.W., R.P., R.A.C., and A.M.K. designed and analysed inDrops experiments and transcriptome data. F.D.C. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks B. Gottgens and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 TARIS.
Illustration of the TARIS procedure. The procedure is described in detail in the Methods.
Extended Data Figure 2 Evaluation of the TARIS method.
a, Design for the detection limit experiment. Spike-ins of a known number of HEK293 cells carrying unique transposon integration tags were used in a mix of 10,000 DsRed+ peripheral blood cells from a freshly induced HSB mouse. b, Detection limit chart. Values represent the read number for each clone and for each number of input cells. Both axes are in log10 scale. Values represent the sum of two independent experiments. c, Comparison of the average read number value between TARIS and the LM-PCR method. Values represent mean ± s.d. of five different transposon tag clones. d, Reproducibility analysis in a non-whole-genome amplified sample with high complexity (2 × 105 bone marrow granulocytes 4 weeks after pulse). e, Reproducibility in a whole-genome amplified sample with low complexity (863 LT-HSCs 4 weeks after pulse). f, Venn diagram showing overlapping transposon tag reads between two TARIS replicates from the same sample high-complexity sample (2 × 105 bone marrow monocytes at 4 weeks after induction). g, Venn diagram showing overlapping transposon tag reads between two TARIS replicates from the same low-complexity sample (863 LT-HSCs at 4 weeks after induction). h, Contamination analysis between samples from two different mice. The plot represents the read numbers of tags from Lin+ populations from mouse 1, and their read number values in Lin+ populations in mouse 2. High-confidence tags are selected as those tags with more than 25 reads, and at least 10 times higher read count compared with any of the samples from a separate mouse.
Extended Data Figure 3 Analysis of residual HSB activity after Dox withdrawal.
a, Experimental design. Residual HSB activity after Dox removal was assayed by transplantation into CD45.1 mice. Sub-lethally irradiated recipients were treated with Dox for 48 h. Dox was removed 12 h before transplantation. Ten million whole bone marrow cells were transplanted and mice were allowed to recover for 2 weeks. As a positive control, mice were continuously treated with Dox until 48 h after transplant. As a negative control, cells were transplanted into non-Dox treated mice. DsRed labelling was analysed as a proxy for HSB activity in granulocytes, erythroblasts, monocytes, and B cells. b, FACS plots showing the negligible labelling of CD45.2 M2/HSB/Tn cells in transplanted recipients 24 h after Dox removal.
Extended Data Figure 4 Additional representations and analyses of Lin+ tags.
a, Lin+ clones of the second and third mice used for quantifications in Fig. 1d–g. b, Scale-adjusted binary (presence/absence) representation of all detected MkP and erythroblast tags in the experiments from Fig. 1d–g. c, Relative quantification of scale-normalized clone sizes for each lineage, comparing unilineage versus oligo/multilineage clones. Values are interquartile range and median from three independent mice at four weeks and eight weeks after induction.
Extended Data Figure 5 Validation of granulocyte/monocyte/erythroblast and megakaryocyte-restricted tags.
a, Three independent transposon tag libraries were prepared and sequenced from 2-week-, 4-week-, and 8-week-chased mice. Reads from the three libraries were then pooled together for each lineage.
Extended Data Figure 6 Lineage fate of myeloid progenitors.
a, Two M2/HSB/Tn mice were induced and chased for one week, and then myeloid progenitors (GMP, MEP, and CMP) and Lin+ cells were isolated from bone marrow and their transposon tag content was analysed. Chart is a binary representation of all Lin+ tags overlapping with any myeloid progenitor tag ranked by lineage. b, Quantification of relative lineage contribution of GMPs, MEPs, and CMPs as a fraction of lineage-specific/total lineage-overlapping clones for each MyP subset. Values are mean of the two analysed mice. c, An additional M2/HSB/Tn mouse was induced and chased for three weeks, and then processed as in a. d, Quantification of relative lineage contribution of GMPs, MEPs, and CMPs at three weeks after labelling.
Extended Data Figure 7 Additional analyses of MPP clonal outcomes.
a, Quantification of the percentage of MPP clones that produced any Lin+ progeny at different time points. Values are average ± s.d. from three mice. b, Three independent transposon tag libraries were prepared and sequenced for all the populations from one bone marrow at 2, 4, and 8 weeks after labelling. Each column in the charts represents the combined tags detected in any of the three libraries for each population.
Extended Data Figure 8 Single-cell heterogeneity of HSC/MPPs.
a, SPRING plots showing selected differentially expressed markers. Scale represents amount of detected mRNA copies (normalized) of each marker gene. b, Enrichment score analysis for single cells in each FACS-sorted population compared with previously obtained bulk transcriptional signatures of bone marrow populations sorted using traditional markers (from the Immgen database).
Extended Data Figure 9 Differentially expressed markers for clusters C1, C2, C3, and megakaryocyte.
a, FACS plots showing heterogeneity in expression of cluster markers within the analysed HSC/MPP subsets. b, FACS plots showing expression of different megakaryocyte-primed cluster markers (CD41, CD42, and CD9) within the LT-HSC gate. c, The expression value (nTrans) and percentage of expressing cells from each cluster (% Exp). The top ten highest expressed genes that distinguish each cluster are shown.
Extended Data Figure 10 Additional data on clonal origin of MkP.
a, Three independent transposon tag libraries were prepared and sequenced for LT-HSC, MPP, and the five Lin+ populations, from one mouse at four weeks. Each column represents the combined tags detected from three amplicon libraries prepared for each population, to facilitate visualization of the smallest clones. Tags are coloured by frequency in each lineage, and organized by rank. b, Origin of megakaryocytes. Alignment of all MkP clones that had detectable tags in HSC/MPPs from a mixed library combining three independent sequencing reactions. Tags are coloured by frequency in each lineage (except for MkP), and organized by rank. Arrows indicate tags verified by clone-specific PCR. c, Alignment of transposon tags from all Lin+ populations, LT-HSCs, and MPPs collected from 30-week-chased mice. Tags are coloured by frequency in each lineage, and organized by rank. d, Experimental design for testing in vitro myeloid and lymphoid potential from sorted LT-HSCs. e, In vitro myeloid potential of LT-HSCs. Alignment of donor Lin+ tags with transposon tags obtained from myeloid and lymphoid cells derived from donor LT-HSCs after two weeks in culture. f, Clonal output of CD41hi and CD41lo LT-HSCs at four weeks after labelling. g, Quantification of megakaryocyte lineage replacement by CD41hi versus CD41lo LT-HSCs (measured as the percentage of overlapping/total MkP reads) at four weeks after labelling. Values are mean ± s.e.m. of three independent mice.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-3. (PDF 2435 kb)
Supplementary Table 1
Sorted cells, clones identified and % DsRed in each population analysed. This table compiles the information of numbers of barcodes and % of DsRed from each mouse analysed, including the sorting logics for each population in each experiment. (XLSX 47 kb)
Supplementary Table 2
Cluster differential expression analysis results. This set of tables contains the raw results of cluster differential expression analysis for each cluster under different tabs. (XLSX 245 kb)
Rights and permissions
About this article
Cite this article
Rodriguez-Fraticelli, A., Wolock, S., Weinreb, C. et al. Clonal analysis of lineage fate in native haematopoiesis. Nature 553, 212–216 (2018). https://doi.org/10.1038/nature25168
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25168
This article is cited by
-
Mitochondrial genetics through the lens of single-cell multi-omics
Nature Genetics (2024)
-
Increased iron uptake by splenic hematopoietic stem cells promotes TET2-dependent erythroid regeneration
Nature Communications (2024)
-
Made to order: emergency myelopoiesis and demand-adapted innate immune cell production
Nature Reviews Immunology (2024)
-
Trim47 prevents hematopoietic stem cell exhaustion during stress by regulating MAVS-mediated innate immune pathway
Nature Communications (2024)
-
Aging is associated with functional and molecular changes in distinct hematopoietic stem cell subsets
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.