Abstract
Cellular senescence is a stress-responsive cell-cycle arrest program that terminates the further expansion of (pre-)malignant cells1,2. Key signalling components of the senescence machinery, such as p16INK4a, p21CIP1 and p53, as well as trimethylation of lysine 9 at histone H3 (H3K9me3), also operate as critical regulators of stem-cell functions (which are collectively termed ‘stemness’)3. In cancer cells, a gain of stemness may have profound implications for tumour aggressiveness and clinical outcome. Here we investigated whether chemotherapy-induced senescence could change stem-cell-related properties of malignant cells. Gene expression and functional analyses comparing senescent and non-senescent B-cell lymphomas from Eμ-Myc transgenic mice revealed substantial upregulation of an adult tissue stem-cell signature, activated Wnt signalling, and distinct stem-cell markers in senescence. Using genetically switchable models of senescence targeting H3K9me3 or p53 to mimic spontaneous escape from the arrested condition, we found that cells released from senescence re-entered the cell cycle with strongly enhanced and Wnt-dependent clonogenic growth potential compared to virtually identical populations that had been equally exposed to chemotherapy but had never been senescent. In vivo, these previously senescent cells presented with a much higher tumour initiation potential. Notably, the temporary enforcement of senescence in p53-regulatable models of acute lymphoblastic leukaemia and acute myeloid leukaemia was found to reprogram non-stem bulk leukaemia cells into self-renewing, leukaemia-initiating stem cells. Our data, which are further supported by consistent results in human cancer cell lines and primary samples of human haematological malignancies, reveal that senescence-associated stemness is an unexpected, cell-autonomous feature that exerts its detrimental, highly aggressive growth potential upon escape from cell-cycle blockade, and is enriched in relapse tumours. These findings have profound implications for cancer therapy, and provide new mechanistic insights into the plasticity of cancer cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Muñoz-Espín, D. & Serrano, M. Cellular senescence: from physiology to pathology. Nat. Rev. Mol. Cell Biol. 15, 482–496 (2014)
Pérez-Mancera, P. A., Young, A. R. & Narita, M. Inside and out: the activities of senescence in cancer. Nat. Rev. Cancer 14, 547–558 (2014)
Zon, L. I. Intrinsic and extrinsic control of haematopoietic stem-cell self-renewal. Nature 453, 306–313 (2008)
Krizhanovsky, V. et al. Implications of cellular senescence in tissue damage response, tumor suppression, and stem cell biology. Cold Spring Harb. Symp. Quant. Biol. 73, 513–522 (2008)
Serrano, M., Lin, A. W., McCurrach, M. E., Beach, D. & Lowe, S. W. Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 88, 593–602 (1997)
Braig, M. et al. Oncogene-induced senescence as an initial barrier in lymphoma development. Nature 436, 660–665 (2005)
Michaloglou, C. et al. BRAFE600-associated senescence-like cell cycle arrest of human naevi. Nature 436, 720–724 (2005)
Schmitt, C. A . et al. A senescence program controlled by p53 and p16INK4a contributes to the outcome of cancer therapy. Cell 109, 335–346 (2002)
Dörr, J. R. et al. Synthetic lethal metabolic targeting of cellular senescence in cancer therapy. Nature 501, 421–425 (2013)
Onder, T. T. et al. Chromatin-modifying enzymes as modulators of reprogramming. Nature 483, 598–602 (2012)
Krizhanovsky, V. & Lowe, S. W. Stem cells: The promises and perils of p53. Nature 460, 1085–1086 (2009)
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000)
Narita, M. et al. Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 113, 703–716 (2003)
Chen, J. et al. H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nat. Genet. 45, 34–42 (2013)
Mosteiro, L. et al. Tissue damage and senescence provide critical signals for cellular reprogramming in vivo. Science 354, aaf4445 (2016)
Ritschka, B. et al. The senescence-associated secretory phenotype induces cellular plasticity and tissue regeneration. Genes Dev. 31, 172–183 (2017)
Wong, D. J. et al. Module map of stem cell genes guides creation of epithelial cancer stem cells. Cell Stem Cell 2, 333–344 (2008)
Reya, T. et al. A role for Wnt signalling in self-renewal of haematopoietic stem cells. Nature 423, 409–414 (2003)
Wang, Y. et al. The Wnt/beta-catenin pathway is required for the development of leukemia stem cells in AML. Science 327, 1650–1653 (2010)
Lin, A. W . et al. Premature senescence involving p53 and p16 is activated in response to constitutive MEK/MAPK mitogenic signaling. Genes Dev. 12, 3008–3019 (1998)
Kelly, P. N., Dakic, A., Adams, J. M., Nutt, S. L. & Strasser, A. Tumor growth need not be driven by rare cancer stem cells. Science 317, 337 (2007)
Chaffer, C. L. et al. Normal and neoplastic nonstem cells can spontaneously convert to a stem-like state. Proc. Natl Acad. Sci. USA 108, 7950–7955 (2011)
Zhao, Z . et al. p53 loss promotes acute myeloid leukemia by enabling aberrant self-renewal. Genes Dev. 24, 1389–1402 (2010)
Zuber, J. et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478, 524–528 (2011)
Acosta, J. C. et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 15, 978–990 (2013)
Baker, D. J. et al. Naturally occurring p16Ink4a-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016)
Baar, M. P. et al. Targeted apoptosis of senescent cells restores tissue homeostasis in response to chemotoxicity and aging. Cell 169, 132–147.e16 (2017)
Pawlikowski, J. S. et al. Wnt signaling potentiates nevogenesis. Proc. Natl Acad. Sci. USA 110, 16009–16014 (2013)
Haugstetter, A. M. et al. Cellular senescence predicts treatment outcome in metastasised colorectal cancer. Br. J. Cancer 103, 505–509 (2010)
Adams, J. M. et al. The c-myc oncogene driven by immunoglobulin enhancers induces lymphoid malignancy in transgenic mice. Nature 318, 533–538 (1985)
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001)
Martins, C. P ., Brown-Swigart, L . & Evan, G. I. Modeling the therapeutic efficacy of p53 restoration in tumors. Cell 127, 1323–1334 (2006)
Reimann, M. et al. Tumor stroma-derived TGF-β limits myc-driven lymphomagenesis via Suv39h1-dependent senescence. Cancer Cell 17, 262–272 (2010)
Schmitt, C. A . et al. Dissecting p53 tumor suppressor functions in vivo. Cancer Cell 1, 289–298 (2002)
Premsrirut, P. K. et al. A rapid and scalable system for studying gene function in mice using conditional RNA interference. Cell 145, 145–158 (2011)
Schmitt, C. A., McCurrach, M. E., de Stanchina, E., Wallace-Brodeur, R. R. & Lowe, S. W . INK4a/ARF mutations accelerate lymphomagenesis and promote chemoresistance by disabling p53. Genes Dev. 13, 2670–2677 (1999)
Banchereau, J., de Paoli, P., Vallé, A., Garcia, E. & Rousset, F. Long-term human B cell lines dependent on interleukin-4 and antibody to CD40. Science 251, 70–72 (1991)
Schmitt, C. A., Rosenthal, C. T. & Lowe, S. W. Genetic analysis of chemoresistance in primary murine lymphomas. Nat. Med. 6, 1029–1035 (2000)
Feil, R., Wagner, J., Metzger, D. & Chambon, P. Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochem. Biophys. Res. Commun. 237, 752–757 (1997)
Jing, H. et al. Opposing roles of NF-κB in anti-cancer treatment outcome unveiled by cross-species investigations. Genes Dev. 25, 2137–2146 (2011)
Yang, D. H. et al. Wnt5a is required for endothelial differentiation of embryonic stem cells and vascularization via pathways involving both Wnt/β-catenin and protein kinase Cα. Circ. Res. 104, 372–379 (2009)
Dickins, R. A. et al. Probing tumor phenotypes using stable and regulated synthetic microRNA precursors. Nat. Genet. 37, 1289–1295 (2005)
Godar, S . et al. Growth-inhibitory and tumor- suppressive functions of p53 depend on its repression of CD44 expression. Cell 134, 62–73 (2008)
Dimri, G. P. et al. A biomarker that identifies senescent human cells in culture and in aging skin in vivo. Proc. Natl Acad. Sci. USA 92, 9363–9367 (1995)
Reimann, M. et al. The Myc-evoked DNA damage response accounts for treatment resistance in primary lymphomas in vivo. Blood 110, 2996–3004 (2007)
Greve, B., Kelsch, R., Spaniol, K., Eich, H. T. & Götte, M. Flow cytometry in cancer stem cell analysis and separation. Cytometry A 81A, 284–293 (2012)
Gupta, P. B. et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 138, 645–659 (2009)
Lu, D. et al. Salinomycin inhibits Wnt signaling and selectively induces apoptosis in chronic lymphocytic leukemia cells. Proc. Natl Acad. Sci. USA 108, 13253–13257 (2011)
Kim, J. et al. A Myc network accounts for similarities between embryonic stem and cancer cell transcription programs. Cell 143, 313–324 (2010)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protocols 2, 1896–1906 (2007)
Boersema, P. J., Raijmakers, R., Lemeer, S., Mohammed, S. & Heck, A. J. Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat. Protocols 4, 484–494 (2009)
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008)
Lee, T. I., Johnstone, S. E. & Young, R. A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat. Protocols 1, 729–748 (2006)
Hu, Y . & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009)
Ivanova, N. B. et al. A stem cell molecular signature. Science 298, 601–604 (2002)
Coppé, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010)
Whitfield, M. L. et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002)
Hu, Z. et al. The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genomics 7, 96 (2006)
Chambers, S. M. et al. Hematopoietic fingerprints: an expression database of stem cells and their progeny. Cell Stem Cell 1, 578–591 (2007)
Acknowledgements
We thank G. Evan, the late A. Harris, T. Jacks and T. Jenuwein for mice, cells and materials; E. Berg, N. Burbach, A. Herrmann, H. Lammert, S. Mende, B. Teichmann and the Berlin-Brandenburg Center for Regenerative Therapies (BCRT) flow cytometry laboratory for technical assistance; and members of the Schmitt laboratory for discussions and editorial advice. This work was supported by a Ph.D. fellowship to J.R.D. from the Boehringer Ingelheim Foundation; by grants from the Deutsche Forschungsgemeinschaft to B.D., M.H. and C.A.S. (SFB/TRR 54) and to A.T. (SFB 873); by the Helmholtz Alliance ‘Preclinical Comprehensive Cancer Center’ grant (HA-305) from the Helmholtz Association to A.T. and C.A.S.; by the Dietmar Hopp Foundation to A.T.; and by the Deutsche Krebshilfe (grant 110678) to C.A.S. This interdisciplinary work was further made possible by the Berlin School of Integrative Oncology (BSIO) graduate program funded within the German Excellence Initiative, and the German Cancer Consortium (GCC).
Author information
Authors and Affiliations
Contributions
M.Mi. performed mouse lymphoma and leukaemia work, stem-cell and senescence assays, and gene set enrichment analyses. J.H.M.D. and M.R. conducted analyses with human cancer cell lines and primary human material. D.N.Y.F. and D.B. carried out flow cytometric analyses. Z.Z. generated leukaemias in the p53-regulatable mouse T-ALL model, I.A.M.B. and J.Z. in the p53-regulatable mouse AML model. Y.Y. carried out biochemical analyses. J.R.D. provided transcriptome analyses and long-term outcome data from senescence-capable mouse lymphomas. L.D. and M.A.M.-P. performed chromatin immunoprecipitations and analysed related datasets. D.L. conducted Affymetrix gene expression profiling and analyses. T.K. and G.D. carried out proteome analyses. M.Me. generated β-catenin/TCF-reporter cancer cell lines and performed luciferase reporter assays. K.P. generated qPCR data. A.T., B.D., H.G. and S.L. contributed to study design, data interpretation and preparation of the manuscript. M.H. provided immunohistochemical analyses. C.A.S. designed experiments, analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. P. Medema, J. Vormoor and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Senescent cells of mouse and human origin present with enhanced stem-cell markers and functionalities.
a, 5,401 probe sets (corresponding to 3,867 genes) differentially expressed in TIS were determined from the transcriptome data comparing untreated and ADR-senescent primary control;Bcl2 lymphomas by two-way ANOVA adjusted for multiple testing (cut-off q < 0.05, n = 12 biologically independent samples). 181 out of 737 genes belonging to an ATSC17 or 43 out of 337 genes of core embryonic stem-cell (ESC) signature49 were detected and marked orange and blue, respectively, in the fold-change-ranked gene list. Whereas the expression of core embryonic stem-cell genes was not correlated with senescence, ATSC transcripts exhibit a strong association with TIS. b, Senescence-selective gene set enrichment pattern of proliferation- and stem-cell-related gene modules (including haematopoietic stem cell (HSC) and long-term HSC (LT-HSC) signatures)56,58,59,60 in control;Bcl2 and Suv39h−;Bcl2 lymphoma cells as in Fig. 1a. GSEA based on the Kolmogorov–Smirnov test, with negative NES indicating enrichment in untreated lymphomas, and positive NES reflecting enrichment in TIS. n = 12 biologically independent control;Bcl2 samples and n = 5 Suv39h−;Bcl2 samples. NES of P < 0.05 are considered statistically significant and are shown in red. c, Senescence induction by ADR treatment in various human cell lines consisting of haematological malignancies, colorectal cancers, melanomas, or in primary samples from patients with B-CLL as determined by SA-β-gal staining (mean percentage of positive cells ± s.d., n = 3 independent experiments for cell lines; n = 4 individual B-CLL samples). TIS-competent cells are defined by a greater than fourfold induction of SA-β-gal-positive cells (with the exception of B-CLL samples, in which SA-β-gal-positive cells were at least threefold induced), and depicted as a blue box symbol in Fig. 1c. d, ABC transporter activity in cells as in Fig. 1a, measured by the efflux of a fluorescent substrate with and without the ABC transporter inhibitor verapamil. Representative plots of four independent lymphomas tested per genotype. e, Enhanced expression of the stem-cell marker CD34 in the RCK8 cell line or primary human B-cell leukaemia samples exposed to ADR treatment in vitro. Mean fluorescence intensity ± s.d. from three independent experiments (RCK8 cells) and five individual leukaemia cases determined by flow cytometry. Two-tailed, unpaired t-test with Welch’s correction, *P < 0.05. f, TIS-mediated increase and verapamil-dependent blockage of ABC transporter activity in ADR-senescent RCK8 cells and primary human B-cell leukaemia samples as in e. One representative out of three independent experiments shown. g, SAS occurring in non-malignant senescence scenarios: GSEA of proliferation- or stem-cell-related gene sets (as in b) in publicly available transcriptome data representing different models of replicative senescence: primary human mammary epithelial cells in stasis or agonescence (GSE16058, 12 prestasis, 9 stasis and 4 agonescence individual biological samples), high-passage BJ human skin fibroblasts (GSE13330, n = 6 pairs of proliferating/senescent cells from individual donors) or high-passage primary human mesenchymal stem cells (GSE9593, n = 3 pairs of proliferating/senescent cells from individual donors).
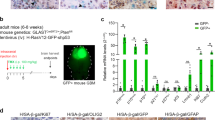
Extended Data Figure 2 Genetic, biochemical and functional properties of regulatable senescence models.
a, Graphic illustration of the model system engineered to stably express a regulatable senescence-essential gene moiety, such as Suv39h1− proficient and -deficient Eμ-Myc transgenic and Bcl2-infected lymphoma variants of which only Suv39h1−;Bcl2;Suv39h1-ERT2 cells regain conditional TIS capability if exposed to 4-OHT. b, Relative transcript levels of the indicated stem-cell-related and Wnt target (asterisk) genes by qPCR in Suv39h1−;Bcl2;Suv39h1-ERT2 lymphoma cells exposed to the indicated treatments for five days. Results represent mean fold induction relative to the untreated condition ± s.d. (n = 3 biologically independent samples). c, Global proteome analysis of total Suv39h1−;Bcl2;Suv39h1-ERT2 cell lysates after five days of ADR ± 4-OHT treatment, showing mean protein expression changes relative to untreated condition (x axis) and their statistical significance (y axis), n = 3 biologically independent samples analysed by Wilcoxon test. All identifications with a −log10 transformed P value greater than 1 were considered significant. Dots representing ATSC factors are highlighted in orange. d, Immunoblot of H3K9me3 expression in Suv39h1−;Bcl2;Suv39h1-ERT2 lymphoma cells treated as in b (‘treatment’), and monitored at the indicated passages in 4-OHT/ADR-free medium (‘post-treatment’; p1–3, each passage reflects 7 days in culture). Never senescent, ADR-only- and previously senescent ADR+4-OHT-pretreated lymphoma cells are analysed, α-tubulin is used as a loading control. One out of two independent experiments shown. For gel source data, see Supplementary Fig. 1. e, f, Growth curve analysis (e) and SA-β-gal reactivity time course (f) of cells treated as in d. Results represent mean cell numbers or percentages of positive cells, respectively ± s.d., from three biologically independent samples. g, Kinetics of the proliferation marker EdU and the fluorescent SA-β-gal marker in Suv39h1−;Bcl2;Suv39h1-ERT2 lymphoma cells after five days of ADR ± 4-OHT treatment (‘treatment’), and subsequent passages in 4-OHT/ADR-free medium (‘post-treatment’, p1–3, each passage reflecting seven days in culture), demonstrating outgrowth of senescent (SA-β-gal+) cells after terminating the 4-OHT/ADR treatment. Mean percentages of EdU+/SA-β-gal+ and EdU+/SA-β-gal− cells ± s.d., n = 4 biologically independent samples. Representative photomicrographs from cell populations marked by red circles are shown in Fig. 2a. h, Competition assays of matched passage 2 previously senescent (GFP-labelled) and never senescent (DsRed-labelled) lymphomas plated at an equal ratio (top) and evaluated by fluorescence microscopy-scored colony formation in vitro (bottom left), and by flow cytometric analysis of lymphoma cells isolated from manifest tumours after transplantation (bottom right). Numbers reflect the ratio of red- to green-fluorescent colonies or cells, respectively. One representative out of four independent experiments shown, including colour reversal.
Extended Data Figure 3 Senescence-released (previously senescent) cancer cells display higher tumour-initiating capacity than their never-senescent counterparts.
a–d, Growth properties of conditionally senescent lymphoma cells analysed as in Fig. 2a, b, but using p53-ERTam;Bcl2 lymphoma cells with ADR ± 4-OHT treatment (a, b), or Suv39h1−;Bcl2;Suv39h1-ERT2 lymphoma cells exposed to a single dose of γ-irradiation (8 Gy) instead of ADR, followed by five days of 4-OHT treatment and subsequent passaging in 4-OHT-free medium (c, d). Results presented as mean positive cells or mean colony numbers ± s.d.; n = 4 (a, c, d) or n = 3 (b) biologically independent samples. Representative photomicrographs from one out of three independent experiments (a, c). Two-tailed, unpaired t-test with Welch’s correction, comparing ADR- and 4-OHT+ADR pretreated lymphomas at p6, or 8 Gy- and 4-OHT+8 Gy at p5. *P < 0.05 (b, d). It is noteworthy that the superior growth and clonogenicity of post-senescent cells can be explained neither by rare cells that may simply have bypassed senescence, because the matching never senescent (that is, senescence bypasser) group presented with inferior clonogenicity, nor by an enhanced death rate of non-stem cells in the Suv39h1-proficient aliquot, because no significant differences in viability were observed between never senescent and previously senescent groups throughout these experiments. Viability determined by flow cytometry as thepercentage of annexin V/PI double-negative cells was typically greater than 80% and comparable between never senescent and previously senescent cells (not shown; the same applies for Figs 2a and 4a). Growth-promoting mutations are also unlikely, as senescent cells stopped replicating their DNA. e, f, Colony formation assay of untreated versus five-day-ADR-senescent human RCK8 lymphoma cells (e) or LT174T colon carcinoma cells (f) that were exposed to a shp53-lentivirus or mock infection on day five of ADR treatment, with p53 knockdown enabling outgrowth out of fully established senescence. As observed for mouse lymphoma cells, post-senescent RCK8 and LT174T cells, after just three passages, outperformed the clonogenic potential of tumor cells that were equally exposed to shRNA against p53 but never experienced senescence. Results represent mean colony numbers at indicated passages (each reflecting seven days in ADR-free methylcellulose medium) ± s.d., n = 3 independent experiments. Two-tailed, unpaired t-test with Welch’s correction, comparing untreated shp53 versus ADR + shp53 at p5 (e) or p4 (f). *P < 0.05. g, TIS re-inducibility in Suv39h1−;Bcl2;Suv39h1-ERT2 previously senescent cells (at passage 2, compare with Fig. 2a) re-exposed to 4-OHT and ADR for five days, as detected by SA-β-gal staining (up) and BrdU/PI incorporation (down). Results represent mean percentages of positive cells ± s.d. (n = 4 independent lymphomas).
Extended Data Figure 4 The senescence-associated secretory phenotype (SASP) is dispensable for senescence-associated stemness (SAS) induction.
a, Expression of a panel of SASP transcripts40,57 by qPCR in Suv39h1-regulatable lymphoma cells after five days of ADR ± 4-OHT exposure, and after two passages in 4-OHT/ADR-free medium (that is, in never senescent and previously senescent cells), showing SASP upregulation in TIS and its downregulation back to baseline levels in senescence-released previously senescent cells. Results represent mean fold induction relative to untreated lymphomas ± s.d. (n = 3 biologically independent samples). b, Blunting SASP production (top) by NF-κB super-repressor IκBαΔN (NF-κB-SR)-mediated genetic inhibition of NF-κB as the major SASP driver in TIS cells (without compromising their ability to enter TIS)9,40 did not prevent acquisition of stemness markers (bottom) by qPCR. Results represent mean fold induction relative to mock-transduced untreated cells ± s.d. (n = 4 biologically independent samples). c, Co-expression of the stem-cell marker Sca1 and the TIS marker H3K9me3 by flow cytometry in NF-κB-SR-expressing control;Bcl2 cells exposed to ADR for five days, indicating uncompromised SAS induction. Percentages indicate mean Sca1/H3K9me3 double-positive cells ± s.d. (n = 4 biologically independent samples). d, ABC transporter activity by flow cytometry in control;Bcl2;NF-κB-SR cells as in c, again demonstrating strong induction of stem-cell-reminiscent ABC transporter activity in TIS cells (compare with Extended Data Fig. 1d) irrespective of their blunted SASP response. Representative plots out of four independent lymphomas shown.
Extended Data Figure 5 Wnt signalling is upregulated in senescence.
a, GSEA of gene sets probing stem-cell-relevant signalling pathways in ADR-senescent control;Bcl2 or TIS-incompetent Suv39h1−;Bcl2 cells (as in Fig. 1a). Positive NES indicate enrichment in TIS lymphomas. NES of P < 0.05 are considered statistically significant and are presented in red. n = 12 pairs of independent lymphomas. b, GSEA enrichment plots of selected gene sets presented in a; GO term ‘Canonical Wnt receptor signaling’ (top) or subset of proliferation-relevant Wnt target genes (bottom), showing significant enrichment in ADR-senescent control;Bcl2 but not in TIS-incompetent Suv39h1−;Bcl2 cells. c, Immunoblot analysis of Ser37- and Thr41-dephosphorylated (that is, stabilized and nucleus translocation-capable ‘Active β-catenin’) and total β-catenin in three independent pairs of control;Bcl2 and Suv39h1−;Bcl2 lymphoma cells, exposed to ADR for 5 days (+) or left untreated (−). α-Tubulin is used as a loading control. One out of two independent experiments shown. For gel source data, see Supplementary Fig. 1. d, Wnt activity measured by the TOPflash TCF reporter system (with FOPflash as negative control) in human cell lines in correlation with their senescence inducibility by ADR, as indicated by blue box symbols for senescence-competent cell lines (referring to Extended Data Fig. 1c). Results reflect mean relative light units fold change (between untreated and ADR-treated samples) of three independent experiments ± s.d.
Extended Data Figure 6 Cell-intrinsic activation of Wnt signalling cascade in TIS.
a, b, Expression of indicated stem-cell-related transcripts by qPCR (a) and colony formation (b) in control;Bcl2 lymphomas infected with a constitutively active Ctnnb1 mutant (ΔNCtnnb1) or a mock retrovirus. Data represent mean expression fold change normalized to mock-infected cells and mean colony numbers, respectively ± s.d. (n = 3 biologically independent samples). Two-tailed, unpaired t-test with Welch’s correction. *P < 0.05. c, Immunoblot analysis of Ser9-phosphorylated (that is, inactivated) or total GSK3β, active or total β-catenin (as in Extended Data Fig. 5c), Thr202- and Tyr204-phosphorylated or total Erk1/2, and Ser473-phosphorylated or total Akt in control;Bcl2 lymphoma cells treated with ADR for five days, together with pharmacological inhibitors targeting MAPK and PI3K kinase pathways. α-Tubulin was used as a loading control. One out of two independent experiments shown. For gel source data, see Supplementary Fig. 1. d, Expression of the indicated stem-cell-related transcripts by qPCR in never senescent and previously senescent Suv39h1−;Bcl2;Suv39h1-ERT2 cells (passage 2) exposed to Wnt signalling agonists (Wnt3a, Rspo2, or GSK3β inhibitor) for two days. Colour scale represents mean fold change normalized to never senescent cells not exposed to Wnt agonists ± s.d. (n = 3 individual lymphomas). e, Colony formation of never senescent and previously senescent cells (as in d), after seven days in methylcellulose medium supplemented with the indicated Wnt agonists (mean colony numbers ± s.d., n = 3 individual lymphomas). Two-tailed, unpaired t-test with Welch’s correction. *P < 0.05.
Extended Data Figure 7 Wnt signalling is dispensable for senescence induction, but required for senescence-associated stemness.
a, Senescence induction by ADR in control;Bcl2 lymphoma cells with and without parallel application of the indicated pharmacological or genetic Wnt inhibitors (ICG-001, salinomycin or Ctnnb1 knockdown by shRNA (shCtnnb1)). Results reflect mean percentages of SA-β-gal-positive cells ± s.d. (n = 4 independent lymphomas). b, Expression of stemness-related transcripts by qPCR in ADR-treated control;Bcl2 lymphoma cells exposed to Ctnnb1 knockdown by shRNA retroviral infection (shCtnnb1). The colour scale represents mean fold induction normalized to ADR-untreated (ut) and vector-infected controls ± s.d. (n = 3 biologically independent samples). c, Relative viability of Suv39h1−;Bcl2;Suv39h1-ERT2 cells exposed to the indicated Wnt inhibitors either simultaneously with ADR ± 4-OHT treatment (for the last 48 h of treatment), or at passage 2 after terminating ADR ± 4-OHT (never senescent and previously senescent; treated over 48 h with inhibitors). Results show relative viability normalized to sample with no Wnt inhibitor treatment ± s.d. (n = 3 biologically independent samples). d, Colony formation of human LT174T colon carcinoma cells exposed to mock or shp53-lentivirus upon ADR-induced senescence, and further propagated in ADR-free medium (corresponding to passage 3 in Extended Data Fig. 3f). Results show mean colony counts after seven-day exposure to indicated Wnt inhibitors ± s.d. (n = 3 independent experiments per group). Two-tailed, unpaired t-test with Welch’s correction, *P < 0.05. e, Individual survival times of the six matched never senescent and previously senescent lymphoma pairs (shown collectively in Fig. 3d). f, Individual survival times of mice bearing never senescent (left) and previously senescent lymphomas (right) after exposure to Wnt signalling inhibition by Ctnnb1 knockdown (shCtnnb1) or left uninhibited. The line plots represent the same matched never senescent and previously senescent lymphomas as in e.
Extended Data Figure 8 The previously senescent cell population maintains a stable fraction of Wnt-active stem cells over time.
a, Detection of a slowly dividing subpopulation in previously senescent but not in never senescent lymphoma cells (arrow) by the CFSE membrane dye 1, 4 or 8 days after stopping the ADR ± 4-OHT treatment. Experiment performed in triplicates. b, CFSEhigh previously senescent cells exhibited more profound nuclear β-catenin expression, indicating acquired stemness (passage 3 after 4-OHT/ADR removal; compare with c). One out of three independent experiments, each performed in triplicate. c, Co-staining with β-catenin and CFSE as in b in Suv39h1−;Bcl2;Suv39h1-ERT2 cells, untreated or exposed to ADR ± 4-OHT for five days (‘treatment’) and subsequently passaged in 4-OHT/ADR-free medium (p1–2; each passage reflects seven days in culture). The slowly cycling (CFSEhigh) population was positive for β-catenin and persisted over time, although their relative percentage drops owing to outgrowth of their (CFSElow) progeny. Numbers reflect mean percentages from three independent lymphomas ± s.d. d, e, Higher expression of ATSC- or Wnt-related (asterisks) transcripts by qPCR (d) and higher clonogenic capacity, which can be neutralized by indicated pharmacological or genetic Wnt inhibitors (e) in flow-sorted, β-catenin high versus β-catenin low previously senescent cells (passage 3 after 4-OHT/ADR removal). Mean expression levels normalized to untreated cells and mean colony numbers respectively ± s.d., n = 4 biologically independent samples. Two-tailed, unpaired t-test with Welch’s correction, *P < 0.05. f, Immunoblot analysis of β-catenin and H3K9me3 levels in human RCK8 lymphoma cells exposed to ADR for 5 days to induce senescence (‘treatment’), then stably transduced with an shp53- or mock lentivirus, and further propagated in ADR-free medium (‘post-treatment’, p1–5, each reflecting seven days in culture). The senescence-associated high levels of active and total β-catenin achieve a low but stable level at later passages. It is noteworthy that stably senescent ADR-pretreated, mock-infected cells were only blotted in p1. One representative out of three independent experiments shown, with α-tubulin as a loading control. For gel source data, see Supplementary Fig. 1. g, Co-expression of β-catenin and the stem-cell marker CD34 detected by flow cytometry in ADR-pretreated, shp53-infected RCK8 cells as in f, demonstrating a small but stable steady-state fraction of double-positive cells at later passages, explaining the lastingly enhanced colony-forming potential of previously senescent versus never senescent cells. Representative flow cytometry plots from three independent experiments (top) and mean percentages of double-positive cells ± s.d. (bottom) at the indicated passages (n = 3 independent experiments). Two-tailed, unpaired t-test with Welch’s correction. *P < 0.05.
Extended Data Figure 9 Spontaneous escape out of senescence detected in cancer cells without genetic manipulations of senescence-relevant genes.
Flow cytometric analysis of the proliferation marker EdU and a fluorescent SA-β-gal marker in control;Bcl2 cells treated with ADR or left untreated (top), and further cultivated in ADR-free medium (bottom). Co-expression of EdU in a small population of still SA-β-gal-positive cells demonstrates the ability of some ADR-senescent cells to escape the senescence arrest. Numbers represent mean percentages ± s.d. from four independent lymphomas. Photomicrographs depict representative cells from populations marked with red circles (n = 4 independent experiments).
Extended Data Figure 10 Senescence-associated de novo generation of leukaemia stem cells upon depletion of the stem-cell-containing fraction in mouse and human leukaemia samples.
a, Flow cytometry plots of mouse KrasG12D;DOX-on-shp53–GFP-induced T-cell acute lymphoblastic leukaemias (total splenocytes after short-term culture and retroviral Bcl2 infection), stained with a panel of mouse lineage antibodies before and after flow-based sorting of the Lin+GFP+ population. The Lin−GFP+ population (including Kit+Sca1+ leukaemia stem cells) was used as a positive control. Shown are representative plots (n = 3). b, Colony formation of mouse Lin+GFP+ leukaemia cells as in a, pretreated with ADR ± doxycycline (DOX) for five days and subsequently seeded in ADR-free/DOX-supplemented medium, thus producing never senescent and previously senescent cells, respectively. Results represent mean colony counts at passage 2 (each passage reflecting 10 days in culture) ± s.d. (n = 3 biologically independent samples). Two-tailed, unpaired t-test with Welch’s correction. *P < 0.05. c, Nuclear β-catenin expression by immunofluorescence (in red) in equally five-day-ADR-exposed senescent versus non-senescent settings (that is, DOX− versus DOX+). DAPI was used as a nuclear counterstain (in blue). Numbers represent mean percentages of β-catenin-positive cells ± s.d. (n = 3 biologically independent samples). d, Colony formation of never senescent and previously senescent leukaemia cells pretreated as in b (passage 3) with the addition of the indicated pharmacological Wnt inhibitors (mean colony numbers ± s.d., n = 3 biologically independent samples per group). *P < 0.05, two-tailed, unpaired t-test with Welch’s correction. e, Senescence induction by SA-β-gal staining in mouse NrasG12D;MLL-AF9;DOX-on-shp53;Bcl2 bulk AML cells (Lin−Kit+Sca1+-depleted) after five days of the ADR ± DOX treatment. Numbers reflect mean percentages of SA-β-gal-positive cells ± s.d (experiment performed in triplicate). Notably, viability determined as the percentage of annexinV/PI double-negative cells was typically greater than 80% and comparable between treatment groups. f, Stemness-related transcripts by qPCR in conditionally senescent mouse AML cells as in e. Graphs represent mean fold induction ± s.d. (n = 3 independent experiments). g, Colony formation of mouse bulk leukaemia cells pretreated as in e, further propagated in ADR-free DOX-containing medium for 14 days, and plated in methylcellulose medium supplemented with the Wnt inhibitors ICG-001 or salinomycin. Colonies were counted after seven days. Previously senescent AML cells, emerging via DOX-mediated p53 knockdown, presented with the highest, Wnt-dependent clonogenicity, which could be attenuated by pharmacological Wnt inhibition. Results represent mean colonies ± s.d. (n = 3 independent experiments). Two-tailed, unpaired t-test with Welch’s correction. *P < 0.05. h, Colony formation of the CD34+ cell-depleted human AML cell line Molm13 (with constitutive retroviral Bcl2-expression) exposed to senescence-inducing ADR treatment for five days (‘treatment’) and subsequently transduced with the lentiviral shp53 or mock construct (p53-knockdown enabling outgrowth from fully established senescence). Results reflect mean colony numbers ± s.d. (n = 3 independent experiments). Two-tailed, unpaired t-test with Welch’s correction. *P < 0.05. i, Flow cytometric detection of the CD33 myeloid differentiation marker and CD34 stem-cell marker surface expression in samples from patients with AML obtained at diagnosis, before any cell cultivation and after six days of cultivation in vitro. Representative plots are shown (n = 5 individual patient samples). j, Expression of stemness-related transcripts in five-day-ADR-senescent versus untreated, ex vivo CD34+-depleted primary human AML cells as in i (qPCR; average fold induction ± s.d., n = 5 individual patient samples, left). Photomicrographs (right) confirm ADR-inducible senescence by SA-β-gal staining (mean percentages of SA-β-gal positive cells ± s.d., representative photomicrographs from five independent samples). k, Regained CD34 surface expression upon ADR-induced senescence in CD34+-depleted primary human AML cells as presented in j. Numbers reflect mean fluorescence intensity detected by flow cytometry ± s.d. (n = 5 individual patient samples). Two-tailed, paired t-test, *P < 0.05. l, ABC transporter activity in ADR-senescent versus untreated cells as in k. Representative plots are shown (n = 5 individual samples).
Supplementary information
Supplementary Information
This file contains patient information. (PDF 58 kb)
Supplementary Information
This file contains supplementary data figure 1; showing the complete western blot membrane scans corresponding to extended data figures 2, 5, 6 and 8. (PDF 333 kb)
Source data
Rights and permissions
About this article
Cite this article
Milanovic, M., Fan, D., Belenki, D. et al. Senescence-associated reprogramming promotes cancer stemness. Nature 553, 96–100 (2018). https://doi.org/10.1038/nature25167
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25167
This article is cited by
-
Dormancy of cutaneous melanoma
Cancer Cell International (2024)
-
Therapy-induced senescent tumor cell-derived extracellular vesicles promote colorectal cancer progression through SERPINE1-mediated NF-κB p65 nuclear translocation
Molecular Cancer (2024)
-
The senescence journey in cancer immunoediting
Molecular Cancer (2024)
-
Regulation of cancer stem cells by CXCL1, a chemokine whose secretion is controlled by MCM2
BMC Cancer (2024)
-
Blocking methionine catabolism induces senescence and confers vulnerability to GSK3 inhibition in liver cancer
Nature Cancer (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.