Abstract
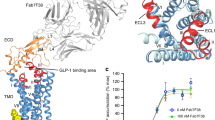
Class B G-protein-coupled receptors (GPCRs), which consist of an extracellular domain (ECD) and a transmembrane domain (TMD), respond to secretin peptides to play a key part in hormonal homeostasis, and are important therapeutic targets for a variety of diseases1,2,3,4,5,6,7,8. Previous work9,10,11 has suggested that peptide ligands bind to class B GPCRs according to a two-domain binding model, in which the C-terminal region of the peptide targets the ECD and the N-terminal region of the peptide binds to the TMD binding pocket. Recently, three structures of class B GPCRs in complex with peptide ligands have been solved12,13,14. These structures provide essential insights into peptide ligand recognition by class B GPCRs. However, owing to resolution limitations, the specific molecular interactions for peptide binding to class B GPCRs remain ambiguous. Moreover, these previously solved structures have different ECD conformations relative to the TMD, which introduces questions regarding inter-domain conformational flexibility and the changes required for receptor activation. Here we report the 3.0 Å-resolution crystal structure of the full-length human glucagon receptor (GCGR) in complex with a glucagon analogue and partial agonist, NNC1702. This structure provides molecular details of the interactions between GCGR and the peptide ligand. It reveals a marked change in the relative orientation between the ECD and TMD of GCGR compared to the previously solved structure of the inactive GCGR–NNC0640–mAb1 complex. Notably, the stalk region and the first extracellular loop undergo major conformational changes in secondary structure during peptide binding, forming key interactions with the peptide. We further propose a dual-binding-site trigger model for GCGR activation—which requires conformational changes of the stalk, first extracellular loop and TMD—that extends our understanding of the previously established two-domain peptide-binding model of class B GPCRs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Drucker, D. J. The biology of incretin hormones. Cell Metab. 3, 153–165 (2006)
Mulder, J. E., Kolatkar, N. S. & LeBoff, M. S. Drug insight: existing and emerging therapies for osteoporosis. Nat. Clin. Pract. Endocrinol. Metab. 2, 670–680 (2006)
Brenneman, D. E. Neuroprotection: a comparative view of vasoactive intestinal peptide and pituitary adenylate cyclase-activating polypeptide. Peptides 28, 1720–1726 (2007)
Sherwood, N. M., Krueckl, S. L. & McRory, J. E. The origin and function of the pituitary adenylate cyclase-activating polypeptide (PACAP)/glucagon superfamily. Endocr. Rev. 21, 619–670 (2000)
Gilligan, P. J. & Li, Y. W. Corticotropin-releasing factor antagonists: recent advances and exciting prospects for the treatment of human diseases. Curr. Opin. Drug Discov. Devel. 7, 487–497 (2004)
Finan, B. et al. Chemical hybridization of glucagon and thyroid hormone optimizes therapeutic impact for metabolic disease. Cell 167, 843–857.e14 (2016)
Longuet, C. et al. The glucagon receptor is required for the adaptive metabolic response to fasting. Cell Metab. 8, 359–371 (2008)
Egerod, K. L. et al. A major lineage of enteroendocrine cells coexpress CCK, secretin, GIP, GLP-1, PYY, and neurotensin but not somatostatin. Endocrinology 153, 5782–5795 (2012)
Hollenstein, K. et al. Insights into the structure of class B GPCRs. Trends Pharmacol. Sci. 35, 12–22 (2014)
Parthier, C., Reedtz-Runge, S., Rudolph, R. & Stubbs, M. T. Passing the baton in class B GPCRs: peptide hormone activation via helix induction? Trends Biochem. Sci. 34, 303–310 (2009)
Mann, R., Wigglesworth, M. J. & Donnelly, D. Ligand–receptor interactions at the parathyroid hormone receptors: subtype binding selectivity is mediated via an interaction between residue 23 on the ligand and residue 41 on the receptor. Mol. Pharmacol. 74, 605–613 (2008)
Liang, Y. L. et al. Phase-plate cryo-EM structure of a class B GPCR–G-protein complex. Nature 546, 118–123 (2017)
Zhang, Y. et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 546, 248–253 (2017)
Jazayeri, A. et al. Crystal structure of the GLP-1 receptor bound to a peptide agonist. Nature 546, 254–258 (2017)
Cho, Y. M., Merchant, C. E. & Kieffer, T. J. Targeting the glucagon receptor family for diabetes and obesity therapy. Pharmacol. Ther. 135, 247–278 (2012)
Zhang, H. et al. Structure of the full-length glucagon class B G-protein-coupled receptor. Nature 546, 259–264 (2017)
Yang, L. et al. Conformational states of the full-length glucagon receptor. Nat. Commun. 6, 7859 (2015)
Siu, F. Y. et al. Structure of the human glucagon class B G-protein-coupled receptor. Nature 499, 444–449 (2013)
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure–function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995)
Wootten, D., Simms, J., Miller, L. J., Christopoulos, A. & Sexton, P. M. Polar transmembrane interactions drive formation of ligand-specific and signal pathway-biased family B G protein-coupled receptor conformations. Proc. Natl Acad. Sci. USA 110, 5211–5216 (2013)
Yang, D. et al. Structural determinants of binding the seven-transmembrane domain of the glucagon-like peptide-1 receptor (GLP-1R). J. Biol. Chem. 291, 12991–13004 (2016)
Runge, S. et al. Three distinct epitopes on the extracellular face of the glucagon receptor determine specificity for the glucagon amino terminus. J. Biol. Chem. 278, 28005–28010 (2003)
Ahn, J. M., Medeiros, M., Trivedi, D. & Hruby, V. J. Development of potent truncated glucagon antagonists. J. Med. Chem. 44, 1372–1379 (2001)
Unson, C. G., Andreu, D., Gurzenda, E. M. & Merrifield, R. B. Synthetic peptide antagonists of glucagon. Proc. Natl Acad. Sci. USA 84, 4083–4087 (1987)
Moon, M. J. et al. Ligand binding pocket formed by evolutionarily conserved residues in the glucagon-like peptide-1 (GLP-1) receptor core domain. J. Biol. Chem. 290, 5696–5706 (2015)
Unson, C. G. et al. Roles of specific extracellular domains of the glucagon receptor in ligand binding and signaling. Biochemistry 41, 11795–11803 (2002)
Yin, Y. et al. An intrinsic agonist mechanism for activation of glucagon-like peptide-1 receptor by its extracellular domain. Cell Discov. 2, 16042 (2016)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protocols 4, 706–731 (2009)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Smart, O. S. et al. Exploiting structure similarity in refinement: automated NCS and target-structure restraints in BUSTER. Acta Crystallogr. D 68, 368–380 (2012)
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013)
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010)
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007)
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981)
Miyamoto, S. & Kollman, P. A. Settle: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992)
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997)
Essmann, U. et al. A smooth particle mesh Ewald Method. J. Chem. Phys. 103, 8577–8593 (1995)
Van Eps, N., Caro, L. N., Morizumi, T. & Ernst, O. P. Characterizing rhodopsin signaling by EPR spectroscopy: from structure to dynamics. Photochem. Photobiol. Sci. 14, 1586–1597 (2015)
Fleissner, M. R., Cascio, D. & Hubbell, W. L. Structural origin of weakly ordered nitroxide motion in spin-labeled proteins. Protein Sci. 18, 893–908 (2009)
Laskowski, R. A. & Swindells, M. B. LigPlot+: multiple ligand–protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51, 2778–2786 (2011)
Polyhach, Y., Bordignon, E. & Jeschke, G. Rotamer libraries of spin labelled cysteines for protein studies. Phys. Chem. Chem. Phys. 13, 2356–2366 (2011)
Acknowledgements
This work was supported by CAS Strategic Priority Research Program XDB08020000, CAS grants QYZDB-SSW-SMC024 (B.W.) and QYZDB-SSW-SMC054 (Q.Z.), the National Science Foundation of China grants 31422017 (B.W.) and 81525024 (Q.Z.), the Shanghai Science and Technology Development Fund 15DZ2291600 (M.-W.W.), the E-Institutes of Shanghai Municipal Education Commission (E09013), the Special Program for Applied Research on Super Computation of the NSFC-Guangdong Joint Fund (second phase) under Grant No. U1501501, and the Canada Excellence Research Chairs program and the Canadian Institute for Advanced Research (O.P.E.). O.P.E. holds the Anne and Max Tanenbaum Chair in Neuroscience. We also thank the computer centre of East China Normal University for computational resources. The synchrotron radiation experiments were performed at the BL41XU of SPring-8 with the approval of the Japan Synchrotron Radiation Research Institute (proposal numbers 2016B2517, 2016B2518, 2017A2505 and 2017A2506). We thank the beamline staff members K. Hasegawa, N. Mizuno, T. Kawamura and H. Murakami of the BL41XU for help with X-ray data collection.
Author information
Authors and Affiliations
Contributions
Ha.Z. optimized the construct, developed the purification procedure and purified the GCGR proteins for crystallization, performed crystallization trials and optimized crystallization conditions. A.Q. helped with construct optimization and crystallization trials. L.Y. performed and analysed molecular dynamics simulations. N.V.E. performed and analysed DEER spectroscopy. K.S.F. performed and analysed binding and potency assays of glucagon and NNC1702. D.Y., A.D. and X.C. designed, performed and analysed the whole-cell glucagon binding assay. Hu.Z. collected the X-ray diffraction data. C.Y. expressed the GCGR proteins. C.C. and L.H. helped to analyse the conformational variety of GCGR. J.L., O.P.E., M.A.H., R.C.S, M.-W.W. and S.R.-R. helped with structure analysis and interpretation, and edited the manuscript. O.P.E. oversaw DEER spectroscopy. M.-W.W. oversaw the whole-cell glucagon binding assay. H.Y. and H.J. oversaw molecular dynamics simulations and commented on the manuscript. S.R.-R. designed the peptide and oversaw ligand characterization of NNC1702. Q.Z. and B.W. initiated the project, planned and analysed experiments, solved the structures, supervised the research and wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
K.S.F., J.L. and S.R.-R. are employees of Novo Nordisk, a pharmaceutical company focused on class B GPCRs for type 2 diabetes. R.C.S. is a founder and board member of Bird Rock Bio, a company focused on GPCR therapeutic antibodies. The remaining authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks G. Schertler, D. Wootten and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
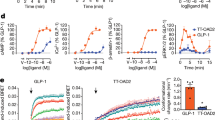
Extended Data Figure 1 Binding affinity and potency of NNC1702.
a, Sequences of glucagon and NNC1702. b, Binding assay of NNC1702. Competitive binding of human glucagon (red dots) and NNC1702 (green squares) to membranes from BHK cells that stably express human GCGR, on WGA-coated SPA beads. Glucagon labelled with 125I (60 pM), and increasing concentrations of human glucagon and NNC1702, were used to generate the binding curves (representative example shown) and calculate IC50 values (glucagon: 1.2 ± 0.5 nM, NNC1702: 12.8 ± 6.6 nM). At least three independent experiments were performed with technical duplicates. c, Potency of NNC1702. The potencies of human glucagon (red dots) and NNC1702 (green squares) were determined by luciferase assays using BHK cells stably transfected with the human GCGR and CRE luciferase. Serial dilutions were prepared in medium (with 1 μM as the highest final concentration). Plate luminescence was read and EC50 values (glucagon: 22.8 ± 18.2 pM, NNC1702: 16.2 ± 8.4 nM) were calculated from the activation curves. At least three independent experiments were performed with technical duplicates (representative example shown). d, e, Inhibition of 125I-labelled glucagon binding to CHO-K1 cells expressing wild-type (WT) and the engineered GCGR used for crystallization by glucagon and NNC1702. Data are shown as mean ± s.e.m. from three independent experiments performed in duplicate. ‘Construct’ indicates the GCGR construct used for crystallization. The IC50 values are listed in e.
Extended Data Figure 2 Structural comparison between the GCGR–NNC1702 crystal structure and previously solved class B GPCR structures.
a, b, Comparison between the GCGR–NNC1702 crystal structure and the cryo-electron microscopy structure of GLP-1–GLP-1R–Gs complex in side (a) and extracellular (b) views. The GCGR–NNC1702 structure is shown in cartoon representation and coloured blue (GCGR) and red (NNC1702). The GLP-1–GLP-1R–Gs electron microscopy structure (PDB ID: 5VAI) is shown in cartoon representation and coloured grey (GLP-1R) and green (GLP-1). c, d, Comparison between the crystal structures of the GCGR–NNC1702 and GLP-1R–peptide 5 complexes in side (c) and extracellular (d) views. The receptor in the GLP-1R–peptide 5 structure (PDB ID: 5NX2) is shown in cartoon representation and coloured pink. The ligand peptide 5 is shown as yellow sticks. The red arrow (in d) indicates the rotation of the ECD in the GLP-1R–peptide 5 structure compared to the GCGR–NNC1702 structure. e, Comparison between the GCGR–NNC1702 structure and the GCGR–NNC0640–mAb1 structure. Only the GCGR TMD in both structures and the peptide ligand NNC1702 are shown as cartoons. The TMD in the GCGR–NNC1702 structure is in blue; the TMD in the NNC0640-bound structure is in yellow; and NNC1702 is in red. A close inspection of the two full-length GCGR structures revealed a spatial hindrance caused by the residue S2 of NNC1702 and its contact with D3857.42b in the peptide-bound structure, pushing the residue F3656.56b on helix VI away from the ligand-binding pocket and subsequently leading to the outward shift of the extracellular portion of helix VI (red arrow). The residues F3656.65b and D3857.42b in both structures are displayed as sticks. The hydrogen bond between S2 and D3857.42b in the peptide-bound structure is shown as a green dashed line.
Extended Data Figure 3 Ligand-binding pocket of NNC1702 and interactions between GCGR and NNC1702.
a, Extracellular view of the binding pocket of NNC1702 N-terminal region within the GCGR TMD. The receptor and the peptide ligand are shown as cartoons, and coloured green (stalk), magenta (ECL1), cyan (ECL2), blue (TMD) and red (NNC1702). b, Binding site of NNC1702 C-terminal region in the stalk (blue), ECL1 (magenta) and ECD (orange) of GCGR. c–e, Schematic representation of interactions between GCGR and NNC1702 analysed by LigPlot+ (ref. 43). c, Interactions between GCGR and the N-terminal region of NNC1702 (residues S2–Y10). d, Interactions between GCGR and the middle region of NNC1702 (residues S11–Q20). e, Interactions between GCGR and the C-terminal region of NNC1702 (residues D21–T29). The stick drawings of GCGR residues and NNC1702 are coloured grey and red, respectively. The labels of GCGR residues are coloured orange (ECD), green (stalk), blue (TMD), magenta (ECL1) and cyan (ECL2). The labels of NNC1702 residues are red.
Extended Data Figure 4 Electron densities of the structure of the GCGR–NNC1702 complex.
a, Electron densities of NNC1702. The peptide NNC1702 is shown in red cartoon representation and as brown sticks. Electron densities are contoured at 1.0σ from a |2Fo| − |Fc| map and coloured blue. b–d, Electron densities of key GCGR residues involved in NNC1702 binding. The receptor is shown in grey cartoon representation. The key residues are shown as sticks and coloured yellow (ECD), green (stalk), magenta (ECL1) and blue (TMD).
Extended Data Figure 5 DEER spectroscopy of assembly of GCGR–ligand complex.
a, The GCGR–NNC1702 assembly showing modelled R1 spin labels at the ECD site H89R1 and the TMD site C287R1 on the basis of the GCGR–NNC1702 crystal structure. The nitroxide rotameric models were generated with the MMM software package44. b, Experimental distance distributions between the nitroxide spin-labelled R1 pair of H89R1 and C287R1 in the apo state or in the presence of NNC0640 or NNC1702. The experimental distributions were normalized by area under the curves for comparison purposes. A predicted distance distribution based on the GCGR–NNC1702 structure that was derived from the MMM software (offset blue trace) is also shown. This prediction can be directly compared to the experimentally measured distributions, though rotameric weighting may be different in the prediction. c, Background-corrected dipolar evolution functions (DEFs) and their fits for each of the GCGR samples. The DEF functions were scaled to compare traces. The traces of the apo receptor and the GCGR–NNC0640 and GCGR–NNC1702 complexes are offset in the main plot to show the quality of the fits. The inset shows the overlaid portion of the DEFs. The DEER data demonstrate that all protein samples exhibit multiple peaks, and the addition of the peptide NNC1702 populates longer distances (32–43 Å), which match the distance distribution predicted by the MMM software using the GCGR–NNC1702 structure as a template (b). The main DEER distance that the apo GCGR and the NNC0640-bound receptor showed is around 26 Å. The conformation possibilities of this distance include the inactive conformation observed in the GCGR–NNC0640–mAb1 structure in which the H89R1–C287R1 distance is about 26 Å between nitroxide N–O bonds when using common R1 rotamers42 for modelling, and the different inactive conformational states of the apo receptor that display close contacts between the ECD and TMD, as suggested by previous molecular dynamics simulation studies16,17 , with H89R1–C287R1 distances of 23–29 Å between the nitroxide N–O bonds when R1 side chains are modelled. These results suggest that the ECD in the apo GCGR or the NNC0640-bound receptor may adopt one conformation or multiple conformations, with a H89R1–C287R1 distance of about 26 Å between nitroxide N–O bonds. The longer distance upon binding to the peptide ligand NNC1702 indicates that the receptor ECD undergoes a conformational change to accommodate the peptide. Equilibrium between these conformational states may potentially exist. NNC1702 probably shifts it towards the conformation favourable for peptide binding, in contrast to the small-molecule NAM NNC0640 that has a weak effect on the ECD conformation. This equilibrium between peptide-free and peptide-bound receptors may help explain the fact that more than one peak was observed for the GCGR–NNC1702 complex in this study, although the concentration of NNC1702 used during protein purification and DEER measurements is 50 μM, which is much higher than the binding affinity of the peptide. G-protein binding may further shift the equilibrium to the peptide-bound conformation, although specific experimental data regarding the G-protein-bound receptor are required to validate this point. Our findings support the flexibility of the ECD conformation and further highlight that the conformational change of the ECD is required for peptide binding.
Extended Data Figure 6 Comparison between the GCGR–NNC1702 structure and the GCGR–glucagon model derived from molecular dynamics simulations
. a, Extracellular view of the transmembrane helical bundle. The GCGR–NNC1702 structure is shown in cartoon representation and coloured blue (GCGR) and red (NNC1702). The GCGR–glucagon model derived from molecular dynamics simulations is shown in cartoon representation and coloured orange (GCGR) and yellow (glucagon). The green arrows indicate shifts of helices VI and VII. b, Close-up view of the interaction between H1 of glucagon and D3857.42b of GCGR in the molecular dynamics simulations. The NNC1702 residue S2, the glucagon residues H1 and S2 and the GCGR residue D3857.42b in both the GCGR–NNC1702 structure and the GCGR–glucagon model are shown as sticks. The hydrogen bond formed by H1 and D3857.42b in the molecular dynamics simulations is displayed as a red dashed line. c, Intracellular view of the transmembrane helical bundle. The green arrows indicate shifts of helices V and VI.
Supplementary information
Supplementary Information
This file contains Supplementary Data and Discussion and additional references. (PDF 146 kb)
Source data
Rights and permissions
About this article
Cite this article
Zhang, H., Qiao, A., Yang, L. et al. Structure of the glucagon receptor in complex with a glucagon analogue. Nature 553, 106–110 (2018). https://doi.org/10.1038/nature25153
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25153
This article is cited by
-
Molecular insights into the distinct signaling duration for the peptide-induced PTH1R activation
Nature Communications (2022)
-
Activation of the GLP-1 receptor by a non-peptidic agonist
Nature (2020)
-
Advances in therapeutic peptides targeting G protein-coupled receptors
Nature Reviews Drug Discovery (2020)
-
A unique hormonal recognition feature of the human glucagon-like peptide-2 receptor
Cell Research (2020)
-
Cryo-EM structure of the human PAC1 receptor coupled to an engineered heterotrimeric G protein
Nature Structural & Molecular Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.