Abstract
Transfer-RNA-derived small RNAs (tsRNAs; also called tRNA-derived fragments) are an abundant class of small non-coding RNAs whose biological roles are not well understood. Here we show that inhibition of a specific tsRNA, LeuCAG3′tsRNA, induces apoptosis in rapidly dividing cells in vitro and in a patient-derived orthotopic hepatocellular carcinoma model in mice. This tsRNA binds at least two ribosomal protein mRNAs (RPS28 and RPS15) to enhance their translation. A decrease in translation of RPS28 mRNA blocks pre-18S ribosomal RNA processing, resulting in a reduction in the number of 40S ribosomal subunits. These data establish a post-transcriptional mechanism that can fine-tune gene expression during different physiological states and provide a potential new target for treating cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Gebetsberger, J. & Polacek, N. Slicing tRNAs to boost functional ncRNA diversity. RNA Biol. 10, 1798–1806 (2013)
Thompson, D. M. & Parker, R. Stressing out over tRNA cleavage. Cell 138, 215–219 (2009)
Yamasaki, S., Ivanov, P., Hu, G.-F. & Anderson, P. Angiogenin cleaves tRNA and promotes stress-induced translational repression. J. Cell Biol. 185, 35–42 (2009)
Honda, S. et al. Sex hormone-dependent tRNA halves enhance cell proliferation in breast and prostate cancers. Proc. Natl Acad. Sci. USA 112, E3816–E3825 (2015)
Chen, Q. et al. Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science 351, 397–400 (2016)
Sharma, U. et al. Biogenesis and function of tRNA fragments during sperm maturation and fertilization in mammals. Science 351, 391–396 (2016)
Kumar, P., Kuscu, C. & Dutta, A. Biogenesis and function of transfer RNA-related fragments (tRFs). Trends Biochem. Sci. 41, 679–689 (2016)
Blanco, S. et al. Stem cell function and stress response are controlled by protein synthesis. Nature 534, 335–340 (2016)
Ivanov, P., Emara, M. M., Villen, J., Gygi, S. P. & Anderson, P. Angiogenin-induced tRNA fragments inhibit translation initiation. Mol. Cell 43, 613–623 (2011)
Schorn, A. J., Gutbrod, M. J., LeBlanc, C. & Martienssen, R. LTR-retrotransposon control by tRNA-derived small RNAs. Cell 170, 61–71 (2017)
Martinez, G., Choudury, S. G. & Slotkin, R. K. tRNA-derived small RNAs target transposable element transcripts. Nucleic Acids Res. 45, 5142–5152 (2017)
Haussecker, D. et al. Human tRNA-derived small RNAs in the global regulation of RNA silencing. RNA 16, 673–695 (2010)
Maute, R. L. et al. tRNA-derived microRNA modulates proliferation and the DNA damage response and is down-regulated in B cell lymphoma. Proc. Natl Acad. Sci. USA 110, 1404–1409 (2013)
Kumar, P., Anaya, J., Mudunuri, S. B. & Dutta, A. Meta-analysis of tRNA derived RNA fragments reveals that they are evolutionarily conserved and associate with AGO proteins to recognize specific RNA targets. BMC Biol. 12, 78 (2014)
Jepsen, J. S., Sørensen, M. D. & Wengel, J. Locked nucleic acid: a potent nucleic acid analog in therapeutics and biotechnology. Oligonucleotides 14, 130–146 (2004)
Elmén, J. et al. LNA-mediated microRNA silencing in non-human primates. Nature 452, 896–899 (2008)
Berridge, M. V., Herst, P. M. & Tan, A. S. Tetrazolium dyes as tools in cell biology: new insights into their cellular reduction. Biotechnol. Annu. Rev. 11, 127–152 (2005)
Cozen, A. E. et al. ARM-seq: AlkB-facilitated RNA methylation sequencing reveals a complex landscape of modified tRNA fragments. Nat. Methods 12, 879–884 (2015)
Zheng, G. et al. Efficient and quantitative high-throughput tRNA sequencing. Nat. Methods 12, 835–837 (2015)
Elmore, S. Apoptosis: a review of programmed cell death. Toxicol. Pathol. 35, 495–516 (2007)
Blobel, G. & Sabatini, D. Dissociation of mammalian polyribosomes into subunits by puromycin. Proc. Natl Acad. Sci. USA 68, 390–394 (1971)
Robledo, S. et al. The role of human ribosomal proteins in the maturation of rRNA and ribosome production. RNA 14, 1918–1929 (2008)
Oskarsson, T. & Trumpp, A. The Myc trilogy: lord of RNA polymerases. Nat. Cell Biol. 7, 215–217 (2005)
Dianzani, I. & Loreni, F. Diamond-Blackfan anemia: a ribosomal puzzle. Haematologica 93, 1601–1604 (2008)
Donati, G., Montanaro, L. & Derenzini, M. Ribosome biogenesis and control of cell proliferation: p53 is not alone. Cancer Res. 72, 1602–1607 (2012)
Chu, C., Quinn, J. & Chang, H. Y. Chromatin isolation by RNA purification (ChIRP). J. Vis. Exp. 2012, 3912 (2012)
Roy-Chaudhuri, B. et al. Regulation of microRNA-mediated gene silencing by microRNA precursors. Nat. Struct. Mol. Biol. 21, 825–832 (2014)
Rehmsmeier, M., Steffen, P., Hochsmann, M. & Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 10, 1507–1517 (2004)
Hofacker, I. L. & Stadler, P. F. Memory efficient folding algorithms for circular RNA secondary structures. Bioinformatics 22, 1172–1176 (2006)
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003)
Reuter, J. S. & Mathews, D. H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010)
Yan, K. et al. Structure prediction: new insights into decrypting long noncoding RNAs. Int. J. Mol. Sci. 17, 132 (2016)
Lu, Z. et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165, 1267–1279 (2016)
Janas, M. M. et al. Reduced expression of ribosomal proteins relieves microRNA-mediated repression. Mol. Cell 46, 171–186 (2012)
Volarevic, S. et al. Proliferation, but not growth, blocked by conditional deletion of 40S ribosomal protein S6. Science 288, 2045–2047 (2000)
Wei, W. et al. Novel celastrol derivatives inhibit the growth of hepatocellular carcinoma patient-derived xenografts. Oncotarget 5, 5819–5831 (2014)
Liu, F., Song, Y. & Liu, D. Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Ther. 6, 1258–1266 (1999)
Zhang, G., Budker, V. & Wolff, J. A. High levels of foreign gene expression in hepatocytes after tail vein injections of naked plasmid DNA. Hum. Gene Ther. 10, 1735–1737 (1999)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010)
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protocols 7, 562–578 (2012)
Fuchs, G., Diges, C., Kohlstaedt, L. A., Wehner, K. A. & Sarnow, P. Proteomic analysis of ribosomes: translational control of mRNA populations by glycogen synthase GYS1. J. Mol. Biol. 410, 118–130 (2011)
Viatour, P. et al. Hematopoietic stem cell quiescence is maintained by compound contributions of the retinoblastoma gene family. Cell Stem Cell 3, 416–428 (2008)
Hadjiolova, K. V., Nicoloso, M., Mazan, S., Hadjiolov, A. A. & Bachellerie, J. P. Alternative pre-rRNA processing pathways in human cells and their alteration by cycloheximide inhibition of protein synthesis. Eur. J. Biochem. 212, 211–215 (1993)
Choesmel, V. et al. Mutation of ribosomal protein RPS24 in Diamond-Blackfan anemia results in a ribosome biogenesis disorder. Hum. Mol. Genet. 17, 1253–1263 (2008)
Acknowledgements
We thank J. Sage for HCC tissues from conditional TKO (Rblox/lox; p130lox/lox; p107−/−) adult mice and liver tissues from p107−/− mice. This work was supported by grants to M.A.K. from the National Institutes of Health (R01AI071068 and R01DK114483). M.A.K. received support from the Stanford Cancer Institute, and S.S. from the CJ Huang Foundation and the TS Kwok Liver Cancer Foundation.
Author information
Authors and Affiliations
Contributions
H.K.K. contributed to experimental design, interpretation, execution, and manuscript writing and editing. G.F. performed experiments in Fig. 3a, b and Extended Data Fig. 1g, h, and assisted with interpretation, discussion and manuscript editing. S.W. performed experiments in Figs 1c, 2b and Extended Data Figs 1i, 2c. W.W. designed experiments with the PDX model and conducted experiments in Fig. 2c and Extended Data Fig. 2h, i, k. Y.Z. performed computational analyses (RNA-seq and predictions of tsRNA binding sites on pre-45S rRNA) in Extended Data Figs 3d and 5a. H.P. performed multiple experiments including Figs 2a, d, 4a, e, and 5b. B.R.-C. conducted the experiment in Extended Data Fig. 8a. P.L. analysed icSHAPE data in Extended Data Fig. 9d. J.X. performed LeuCAG3′tsRNA target site prediction in Extended Data Figs 8b and 9b. F.Z. performed RNA extraction and mouse experiments. K.C. conducted protein extraction. M.-S.C. designed experiments with the PDX model, interpreted animal data and assisted in manuscript editing. S.S. provided discussion regarding the xenograft model. Q.C.Z. analysed, interpreted and discussed icSHAPE data. P.S. interpreted and discussed experimental results and assisted in manuscript editing. M.A.K. contributed to the experimental design, data interpretation, and manuscript writing and editing.
Corresponding author
Ethics declarations
Competing interests
H.K.K., S.W. and M.A.K. are inventors on relevant patents filed by Stanford University.
Additional information
Reviewer Information Nature thanks N. Polacek and L. Zender for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 22-nt LeuCAG3′tsRNA is involved in cell viability.
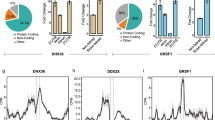
a, b, Inhibition of LeuCAG3′tsRNA impairs HCT-116 cell viability. Three days post-transfection, an MTS assay was performed (n = 3 independent experiments). Each LNA is perfectly complementary to the coloured region on the tRNA diagram above each bar. Blue and red asterisks mark different 2-nt mismatches. c, d, Inhibition of LeuCAG3′tsRNA decreased the number of viable HeLa (c) and HCT-116 cells (d) (n = 3 independent experiments). The cell number on each day was normalized to the day zero value. e, Cleavage of LeuCAG3′tsRNA impairs HeLa cell viability. Three days post-transfection, an MTS assay was performed as in a (n = 3 independent experiments). f, Northern analysis of LeuCAG3′tsRNA and mature Leu-tRNA following transfection. To quantify the tsRNA and mature tRNA levels correctly, transferred blots were cut at the 40–50-nt position and hybridized with the same probe to detect tRNA (top) and tsRNA (bottom) separately (n = 2 independent experiments). U6 snRNA, loading control. g, Twenty-four hours post-transfection, a global protein synthesis assay was carried out using [35S]-methionine metabolic labelling in HeLa cells (n = 3 independent experiments). h, Twenty-four hours post-transfection, global protein synthesis detection was carried out on gels using [35S]-methionine metabolic labelling in HeLa cells. Coomassie brilliant blue (left) was loading control; gels were scanned to measure incorporated radioactivity (right) (n = 2 independent experiments). Each number multiplied by 105 is the number of cells on 6-well culture dishes 24 h before transfection. i, Global protein synthesis assay using a Click-iT AHA Alexa Fluor 488 assay in HeLa cells was performed 24 h post-transfection. The nucleus was stained with DAPI, blue. Protein synthesis was measured with AHA, green. Merge represents the DAPI and AHA merged images (n = 2 independent experiments). Un, untreated cells; mock, transfection without LNA; CHX, cycloheximide-treated positive control. j, The abundance of sequencing reads aligned to LeuCAG-tRNA from HeLa cells. This analysis was generated from tRFdb (http://genome.bioch.virginia.edu/trfdb/search.php). x axis, position on LeuCAG mature tRNA. Blue lines, 18- and 22-nt of LeuCAG 3′tsRNAs. k, The LeuCAG3′tsRNA major isoform is 22 nt. A northern hybridization was performed. The 18-nt isoform was not detected (n = 2 independent experiments). Data show mean ± s.d.; indicated P value by two-tailed t-test (a–e).
Extended Data Figure 2 Inhibition of LeuCAG3′tsRNA induces apoptosis in vitro and inhibits the growth of hepatocellular carcinoma (HCC) patient-derived xenograft.
a, A representative result of the apoptosis assay. Apoptosis in HeLa cells was measured using Annexin V-FITC and propidium iodide (PI) staining at 24, 48 and 72 h post-transfection. The percentage of cells is shown in each gate. Q1 (healthy cells), stained with neither Annexin V nor PI; Q2 (early apoptotic cells), positive with Annexin V; Q3 (late apoptotic cells), positive with Annexin V and PI; Q4 (dead cells), positive with PI. The average cell population of the apoptosis assay is provided in Fig. 2a (n = 3 independent experiments). b, Inhibition of LeuCAG3′tsRNA results in increased apoptosis in HCT-116 cells. The apoptosis assay was done as in a (control at 1 d, n = 2; all other samples, n = 3 independent experiments). c, Inhibition of LeuCAG3′tsRNA causes DNA fragmentation in HCT-116 cells. A TUNEL assay was performed 24 h post-transfection. DNase I is a positive control. TUNEL-positive cells are stained red. Merge is DAPI and TUNEL merged staining (n = 2 independent experiments). d, e, Inhibition (mixmer LNA) (d) and cleavage (gapmer LNA) (e) of LeuCAG3′tsRNA causes PARP protein cleavage. Western blot analysis was performed 24 h post transfection. Anti-PARP antibody detects both full-length (116 kD) and cleaved (89 kD) PARP protein. ACTB, loading control (n = 2 independent experiments). f, For liver toxicity, 125 μg of each LNA (tsRNA sequence is the same in human and mouse) was injected into C57BL/6J mice by hydrodynamic tail vein injection (saline group, n = 2; other group, n = 3 independent mice). g, LeuCAG3′tsRNA is highly expressed from mouse models of HCC generated from conditional TKO (Rblox/lox; p130lox/lox; p107−/−) adult mice43. Normal liver was taken from C57/BL6 mice. Northern hybridization was performed as in Extended Data Fig. 1f (n = 2 independent experiments). h, Over the 4-week study period, mice containing an orthotopic human xenotransplanted hepatocellular carcinoma were given intraperitoneal injections of saline, control, or anti-Leu3′tsLNA. The luciferase signal as a marker of tumour growth was monitored weekly (saline group, n = 8; con group, n = 9; anti-Leu3′ts group, n = 10 independent mice). i, After the 4-week injection period, all mice were killed and tumours removed. j, Anti-Leu3′tsLNA inhibits LeuCAG3′tsRNA in vivo. Northern hybridization was performed with total RNAs from mice injected with saline or anti-Leu3′tsLNA (n = 2 independent experiments). k, Body weights of individual mice bearing HCC xenografts during the 4-week experiment (saline group, n = 8; control group, n = 9; anti-Leu3′ts group, n = 10 independent mice). Data show mean ± s.d.; indicated P value by two-tailed t-test (b, h); P value (b), early apoptosis population. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 3 LeuCAG3′tsRNA does not have transgene silencing activity.
a, LeuCAG3′tsRNA does not repress expression of luciferase gene containing perfect complementary target sites in its 3′ UTR or 5′ UTR. A luciferase plasmid (x axis) was co-transfected with control or anti-Leu3′tsLNA (n = 3 independent experiments). The normalization protocol is described in Methods. Scramble, scrambled sequences in 3′ UTR; LeuCAG3′tsPM in 3′ UTR, two copies of the perfect complementary sequence of the LeuCAG3′tsRNA in 3′ UTR; LeuCAG3′tsPM in 5′ UTR, two copies of the perfect complementary sequence of the LeuCAG3′tsRNA in 5′ UTR; Let-7PM is a positive control, a single copy of perfect complementary sequences of the Let-7 miRNA in 3′ UTR. b, AspGTC3′tsRNA or SerGCT3′tsRNA does not repress luciferase gene expression in a construct that contains two copies of the corresponding perfect complementary target site in its 3′ UTR. A luciferase assay was performed as in a (n = 3 independent experiments). x axis, target sites in the 3′ UTR. c, The tsRNAs are not associated with Ago proteins. Endogenous Ago1, Ago2 and Ago3 were immunoprecipitated by the indicated antibodies and the associated RNAs were subjected to northern blotting. The closed triangle in each northern blot indicates the detected tsRNA. IgG, control. Let-7 is a positive control (n = 2 independent experiments). d, LeuCAG3′tsRNA does not affect global gene expression in HeLa and HCT-116 cells. Scatter plots comparing gene expression (log2[FPKM+1]) of two RNA-seq data sets from samples 24 h post-transfection (Supplementary Table 2). The Pearson correlation coefficient is indicated by the r value in each plot (n = 1). Data show mean ± s.d. For gel source data, see Supplementary Fig. 1.
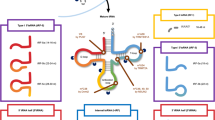
Extended Data Figure 4 LeuCAG3′tsRNA is a non-coding RNA required for ribosome biogenesis.
a, LNAs directed against 5′ end of LeuCAG-tRNA, Ser3′tsRNA (3′ end of the SerGCT-tRNA), and Met3′tsRNA (3′ end of the MetCAT-tRNA) do not change ribosome/polysomal profiles. Twenty-four hours post-transfection, cytoplasmic lysates from HeLa cells were treated with cycloheximide and separated on 10–50% sucrose gradients. The polysomal profile was analysed as in Fig. 3a (n = 2 independent experiments). b, Pre-rRNA processing pathways in human cells based on prior studies44,45. The 45S primary transcript (pre-45S) is processed and categorized as: 5′ external transcribed spacers (5′ETS), mature 18S rRNA, internal transcribed spacer 1 (ITS1), mature 5.8S rRNA, internal transcribed spacer 2 (ITS2), mature 28S rRNA, and 3′external transcribed spacers (3′ETS). There are two alternative processing pathways. Inhibition of LeuCAG3′tsRNA inhibits processing from the 30S intermediate to 21S intermediate form depicted in pathway B. Arrowhead and number indicate cleavage sites. c, Inhibition of the LeuCAG3′tsRNA suppressed 5′ETS processing in 18S rRNA biogenesis in 293T and HCT-116 cells. Northern hybridization was performed with total RNA from HCT-116 and 293T cells 24 h post-transfection. The ITS1 probe detects the 45S primary transcript and intermediate forms of the mature 18S rRNA including 41S, 30S, 21S, and 18S-E pre-rRNAs. The 5′ETS probe detects the 45S primary transcript and 30S intermediate form of the mature 18S rRNA. Each number, multiplied by 104, on top of the image represents the number of cells plated on 6-well culture dishes the day before transfection (n = 2 independent experiments). For gel source data, see Supplementary Fig. 1.
Extended Data Figure 5 LeuCAG3′tsRNA and anti-Leu3′tsLNA do not affect 18S rRNA biogenesis through binding to 45S pre-rRNA.
a, Schematic picture showing putative binding sites of the LeuCAG3′tsRNA and anti-Leu3′tsLNA on the 45S primary transcript (45S pre-rRNA). To identify the tsRNA binding sites in the 45S pre-rRNA, we used the RNAhybrid program and 18- and 22-nt sequences from the 3′ end of LeuCAG-tRNA. The resulting five putative binding sites were positioned in the 5′ETS, 1 site in ITS1, 1 site in ITS2, 3 sites in 28S rRNA, and 1 site in the 3′ETS. The putative LeuCAG3′tsRNA binding site is indicated as a black bar. The putative binding site of anti-Leu3′tsLNA is indicated as a red bar. b, LeuCAG3′tsRNA and anti-Leu3′tsLNA do not bind to 45S pre-rRNA. To inhibit the interaction between LeuCAG3′tsRNA (or anti-Leu3′tsLNA) and 45S pre-rRNA, each LNA design was based on the sequence shown in a and transfected into HeLa cells for 24 h before RNA extraction and northern hybridization (n = 2 independent experiments). The sequences of each LNA are listed on Supplementary Table 4. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 6 Inhibition of LeuCAG3′tsRNA decreases RPS28 protein level, inducing apoptosis.
a, Inhibition of the LeuCAG3′tsRNA does not change the nuclear–cytoplasmic subcellular localization of RPS6 and RPS28 in HeLa cells. Western blotting was performed 24 h post transfection. (n = 2 independent experiments.) Total, total extracts; C, cytoplasm; N, nucleus. b, c, RPS28 protein levels were downregulated in anti-Leu3′tsLNA-treated HCC samples isolated from the orthotopic PDX. b, Total protein extracts from tumours (Extended Data Fig. 2i) were subjected to western blotting (n = 4 independent experiments). c, Quantification of the RPS28 protein level (n = 4 independent experiments). d, A decrease in RPS28 protein level induces apoptosis in HeLa cells. Western blotting was performed 24 h after transfection of indicated siRNA (n = 2 independent experiments). GAPDH, loading control. e, RPS28 overexpression in HeLa cells for Fig. 4c and Extended Data Fig. 6f–h. The number below the image represents the relative RPS28 protein level normalized to GAPDH (n = 2 independent experiments). f–h, Overexpression of RPS28 restores 18S rRNA processing. After co-transfection with the indicated LNAs and plasmids in HeLa cells for 24 h, northern blots using probes complementary to the 18S rRNA precursor, 18S and 28S rRNA as in Fig. 3e are shown. f, A representative northern blot result (n = 3 independent experiments). g, Relative abundance of 30S pre-rRNA (n = 3 independent experiments). h, Relative abundance of 18S rRNA normalized to 28S rRNA. Each value is normalized to that of control–EGFP transfected cells, which was set at 100 (n = 3 independent experiments). i, RPS28 mRNA levels were unchanged in anti-Leu3′tsLNA-treated HCC samples isolated from an orthotopic PDX. RT–PCR was performed with total RNA from tumours (Fig. 2d). Each mRNA level was normalized to GAPDH mRNA (control and saline groups, n = 4; anti-Leu3′ts LNA group, n = 5). Data show mean ± s.d.; indicated P values by two-tailed t-test (c, g, h). For gel source data, see Supplementary Fig. 1.
Extended Data Figure 7 Inhibition of LeuCAG3′tsRNA specifically alters sedimentation of RPS28 mRNA.
Total RNA from each sucrose gradient fraction in Fig. 3a was extracted (left), and a northern analysis performed. The indicated bp provided to the left of the labelled gene name indicates the size of the coding sequences. The polysome profile is the same as shown in Fig. 3a. Relative distribution of mRNA populations across the gradient (right). Each amount of the specific mRNA for each gradient fraction was normalized using the sum of the mRNA signal across all gradient fractions. x axis is the gradient fraction number; y axis is per cent of mRNA abundance (n = 2 independent experiments). For gel source data, see Supplementary Fig. 1.
Extended Data Figure 8 LeuCAG3′tsRNA regulates RPS28 mRNA translation.
a, LeuCAG3′tsRNA is associated with RPS28 mRNA. RPS28 or GAPDH mRNAs were pulled down with tiling oligos. The enrichment of each mRNA was measured by RT–PCR (left). The associated LeuCAG3′tsRNA was detected by northern hybridization (middle). The relative percentage of associated LeuCAG3′tsRNA is shown. The LeuCAG3′tsRNA was enriched 26 times (right) in the RPS28 versus GAPDH mRNA pulldown after normalization (left). b, The two putative LeuCAG3′tsRNA binding sites in the RPS28 mRNA. Target-site 1 (nt 255–279) in the 3′ UTR. Target-site 2 in the coding sequence (CDS) has two possible predicted configurations (target 2a (nt 108–134) and target2b (nt 117–134). The target2 mutant (labelled C in Fig. 5a and Extended Data Fig. 8e) alters both predicted target 2a and 2b conformations. c, Proposed model of translational regulation. The 3′tsRNA binds to the RPS28 mRNA and disrupts the secondary structure, resulting in enhanced mRNA translation. Protein X, unknown protein(s). d, RPS28 protein levels are affected by the LeuCAG3′tsRNA concentration when the target sites remain unchanged. A representative western result after co-transfection of LNAs (control and anti-Leu3′ts) and the RPS28 mutant plasmids. The relative RPS28 protein level was calculated after normalization to GAPDH (Fig. 5b). Con, control; Anti, anti-Leu3′tsLNA; wt, wild-type construct; other mutant constructs (Fig. 5a and Supplementary Table 8). e, Schematic of C-terminal Flag-tagged RPS28 secondary structure with putative LeuCAG3′tsRNA binding sites. Red, the altered sequences in each mutant; blue, the putative LeuCAG3′tsRNA binding sites in the RPS28 mRNA; black and grey, the coding and non-protein coding sequences, respectively. Black bold, C-terminal Flag tag sequences. f, The C-terminal Flag-tagged RPS28 protein level is affected by LeuCAG3′tsRNA concentrations when the target sites remain unchanged. A representative western result after co-transfection of LNAs (control and anti-Leu3′ts) and plasmids (Flag–RPS28 and Flag–EGFP). The relative RPS28 signal is shown in Fig. 5b and calculated after normalization to co-transfected Flag-tagged EGFP. g, Uncapped firefly luciferase mRNA was translated in RRL with the indicated amounts of synthetic LeuCAG3′tsRNA (Leu). h, Synthetic LeuCAG3′tsRNA increases RPS28 mRNA translation in vitro. Xef1 (control) and RPS28 mRNAs were translated with a synthetic LeuCAG3′tsRNA in vitro. Mock, no mRNA; (-), no control and LeuCAG3′tsRNA; con1, con2, and con3, three different control RNAs. i, The relative RPS28 translation product was normalized to Xef1 from h. j, Leu3′tsRNA does not affect translation of RPS28 in vitro when the putative binding sites are altered. Xef1 and RPS28 wild-type or mutant mRNAs were translated in RRL. The normalized RPS28 protein level is shown in Fig. 5c. k, RPS28 target2 mutant C mRNA translation was enhanced by a compensatory tsRNA mimic (tsRNA(comp)) nearly complementary to sequence-modified target2 site sequences from mutant C. Xef1 and RPS28 wild-type or mutant C mRNAs were translated with the compensatory tsRNA mimic (tsRNA(comp)). Normalized quantification of the RPS28 translation products is shown in Fig. 5d. Each western and IVT figure was cropped from a single image (gel source data, Supplementary Fig. 1). Data show mean ± s.d.; P values by two-tailed t-test (a, i). For a, g–j, n = 3 independent experiments; for d, f, k, n = 4 independent experiments.
Extended Data Figure 9 Double-strandedness of LeuCAG3′tsRNA target sites.
a, Schematic prediction of LeuCAG3′tsRNA binding sites. The target sites of the LeuCAG3′tsRNA in the coding sequences (CDSs) and flanking 30 bp of each mRNA were predicted using RNAhybrid based on the m.f.e. Secondary structures of target sites that were predicted to have a binding site based on the low m.f.e. were analysed by icSHAPE. b, c, Thermodynamics of the putative LeuCAG3′tsRNA binding sites in the RPS15 (b), RPS9 and RPS14 (c) mRNAs. Indicated numbers on each diagram represent the 5′ end and 3′ end position on each mRNA, respectively. d, The icSHAPE data track of LeuCAG3′tsRNA binding sites and 20 nt of flanking regions contained within the RPS28, RPS15, RPS9 and RPS14 mRNAs. The icSHAPE data are scaled from 0 (no reactivity; double-strandedness) to 1 (maximum reactivity; single-strandedness). Red box represents a target site. The complete icSHAPE data for each mRNA are in Supplementary Table 10.
Extended Data Figure 10 The tsRNA affects RPS15 but not RPS9 and RPS14 protein levels.
a, LeuCAG3′tsRNA inhibition decreases the RPS15 protein concentration. Protein levels were determined by western blot (n = 3 independent experiments). The number under the image is the relative RPS15 protein level (anti-Leu3′tsLNA) normalized to control (con). b, RPS15 protein levels were reduced in anti-Leu3′tsLNA-treated HCC orthotopic PDX. Western blotting was performed with PDX tumours (Extended Data Fig. 2i) (n = 4 independent mice). c, Quantification of the RPS15 protein level from b (n = 4 independent mice). d, Inhibition of LeuCAG3′tsRNA does not alter RPS15 mRNA levels. RT–PCR was performed (n = 3 independent experiments). e, RPS15 mRNA levels were unchanged in anti-Leu3′tsLNA treated HCC orthotopic PDX. RT–PCR was performed with total RNA from tumours (Extended Data Fig. 2i) (n = 4 independent mice). f, g, The RPS9 and RPS14 wild-type and target site mutant protein levels are not affected by LeuCAG3′tsRNA concentrations. A representative western result after co-transfection of LNAs (control and anti-Leu3′ts) and RPS9 wild-type or target mutant plasmids (f), and RPS14 wild-type or target mutant plasmids (g) (each n = 2 independent experiments). Target, modified target site mutant. h, The RPS15 protein level is affected by LeuCAG3′tsRNA concentrations when the target site is not altered. Western blotting was performed as in f (n = 4 independent experiments). Non-target, modified non-target site mutant. i, The normalized RPS15 protein level from h was calculated as in Extended Data Fig. 8d (wild-type and target group, n = 6; non-target group, n = 4 independent experiments). j, Leu3′tsRNA does not affect translation of RPS15 mRNA in vitro when the target site is altered. Xef1 and RPS15 wild-type and mutant mRNAs were translated in RRL as in Fig. 5c (n = 4 independent experiments). k, The normalized RPS15 protein level from j (n = 4 independent experiments). Each gel figure was cropped from a single image. Normalization of RT–PCR result is described in the Methods. Data show mean ± s.d.; indicated P value by two-tailed t-test (c, i, k). For gel source data, see Supplementary Fig. 1. The mutant constructs are listed in Supplementary Table 8.
Supplementary information
Supplementary Figure 1
This file contains gel source data. (PDF 9288 kb)
Supplementary Table 1
This file contains the DNA sequences of the CUG (original) and CUC/CUU (modified) Renilla genes from the psiCHECK-2 plasmid from Fig. 1c. (PDF 64 kb)
Red colored nucleotides show the LeuCUG codon and blue colored nucleotides are LeuCUU or LeuCUC codons. There are thirteen LeuCUG, four LeuCUU, and five LeuCUC codons in original Renilla gene. Thirteen LeuCUG codons were replaced by CUU or CUC codons in the modified Renilla gene.
Supplementary Table 2
This file contains samples that were sequenced in Extended Data Fig. 3d. 50bp paired-end reads were generated on an Illumina HiSeq 2000 machine yielding a total of 10 to 40 million paired-end reads. Sequences were mapped to the human hg19 genome. (PDF 58 kb)
Supplementary Table 3
This file contains quantification of each ribosomal RNA in Fig. 3d. Each pre-rRNA was normalized by each mature 28S rRNA. Normalized pre-rRNA from Anti-Leu3′ts LNA was again normalized to that of control (con). Normalized pre-rRNA from siRPS6, 10, 13, and 29 were normalized to the siRNA control (sicontrol). (PDF 71 kb)
Supplementary Table 4
This file contains the list of antisense Locked nucleic acid (LNA), synthetic RNA, and northern probe oligonucleotides. LNA bases are upper-case letters and DNA bases are lower –case letters. (PDF 115 kb)
Supplementary Table 5
This file contains DNA oligonucleotides used for the target sequences of tsRNAs and microRNAs in the luciferase vector in Extended Data Fig. 3a. (PDF 57 kb)
Supplementary Table 6
This file contains PCR primers for the generation of the Northern probes. (PDF 56 kb)
Supplementary Table 7
This file contains Biotin labelled oligonuclotides used for the ChIRP studies in Extended Data Fig. 8a. (PDF 54 kb)
Supplementary Table 8
This file contains modified nucleotide sequences of the ribosomal protein mutants. Upper characters are altered sequences and the numbers next to each sequence indicate the sequence position in the each ribosomal protein gene. (PDF 55 kb)
Supplementary Table 9
This file contains primers for site-directed mutageneis. (PDF 64 kb)
Supplementary Table 10
This file contains icSHAPE scores for the full-length studied mRNAs. Each number represents the scores for each nucleotide. The icSHAPE data are scaled from 0 (no reactivity; double-strandedness) to 1 (maximum reactivity; single-strandedness). (PDF 497 kb)
Rights and permissions
About this article
Cite this article
Kim, H., Fuchs, G., Wang, S. et al. A transfer-RNA-derived small RNA regulates ribosome biogenesis. Nature 552, 57–62 (2017). https://doi.org/10.1038/nature25005
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25005
This article is cited by
-
METTL1 mediated tRNA m7G modification promotes leukaemogenesis of AML via tRNA regulated translational control
Experimental Hematology & Oncology (2024)
-
tRNA-derived small RNAs in human cancers: roles, mechanisms, and clinical application
Molecular Cancer (2024)
-
Non-coding RNAs as potential therapeutic targets for receptor tyrosine kinase signaling in solid tumors: current status and future directions
Cancer Cell International (2024)
-
tiRNA-Val-CAC-2 interacts with FUBP1 to promote pancreatic cancer metastasis by activating c‑MYC transcription
Oncogene (2024)
-
Roles and regulation of tRNA-derived small RNAs in animals
Nature Reviews Molecular Cell Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.