Abstract
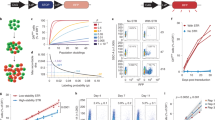
Tissue-resident memory CD8+ T (TRM) cells are found at common sites of pathogen exposure, where they elicit rapid and robust protective immune responses1,2. However, the molecular signals that control TRM cell differentiation and homeostasis are not fully understood. Here we show that mouse TRM precursor cells represent a unique CD8+ T cell subset that is distinct from the precursors of circulating memory cell populations at the levels of gene expression and chromatin accessibility. Using computational and pooled in vivo RNA interference screens, we identify the transcription factor Runx3 as a key regulator of TRM cell differentiation and homeostasis. Runx3 was required to establish TRM cell populations in diverse tissue environments, and supported the expression of crucial tissue-residency genes while suppressing genes associated with tissue egress and recirculation. Furthermore, we show that human and mouse tumour-infiltrating lymphocytes share a core tissue-residency gene-expression signature with TRM cells that is associated with Runx3 activity. In a mouse model of adoptive T cell therapy for melanoma, Runx3-deficient CD8+ tumour-infiltrating lymphocytes failed to accumulate in tumours, resulting in greater rates of tumour growth and mortality. Conversely, overexpression of Runx3 enhanced tumour-specific CD8+ T cell abundance, delayed tumour growth, and prolonged survival. In addition to establishing Runx3 as a central regulator of TRM cell differentiation, these results provide insight into the signals that promote T cell residency in non-lymphoid sites, which could be used to enhance vaccine efficacy or adoptive cell therapy treatments that target cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
10 January 2018
Please see accompanying Erratum (http://doi.org/10.1038/nature25445). The words ‘infection with’ were missing from the sentence ‘Furthermore, Runx3 RNAi also impaired TRM cell differentiation in the context of a localized infection with enteric Listeria monocytogenes expressing GP33–41 (LM–GP33–41) (Fig. 2b).’ In Fig. 1a, in the x-axis label of bottom right graph ‘D35 kid/D35 TCM’ was changed to ‘D7 kid/D7 TCM’. In Fig. 1e, the arrow pointing from the spleen to TCM should have moved to the left, and in Fig. 1f ‘Irf4’ was wrongly set in italic font. These errors have been corrected online. See Supplementary Information to the Erratum for the original Fig. 1
References
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483, 227–231 (2012)
Schenkel, J. M. et al. T cell memory. Resident memory CD8 T cells trigger protective innate and adaptive immune responses. Science 346, 98–101 (2014)
Chang, J. T., Wherry, E. J. & Goldrath, A. W. Molecular regulation of effector and memory T cell differentiation. Nat. Immunol. 15, 1104–1115 (2014)
Mueller, S. N., Gebhardt, T., Carbone, F. R. & Heath, W. R. Memory T cell subsets, migration patterns, and tissue residence. Annu. Rev. Immunol. 31, 137–161 (2013)
Mackay, L. K. et al. The developmental pathway for CD103+CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013)
Mackay, L. K. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 352, 459–463 (2016)
Wakim, L. M. et al. The molecular signature of tissue resident memory CD8 T cells isolated from the brain. J. Immunol. 189, 3462–3471 (2012)
Masopust, D., Vezys, V., Wherry, E. J., Barber, D. L. & Ahmed, R. Cutting edge: gut microenvironment promotes differentiation of a unique memory CD8 T cell population. J. Immunol. 176, 2079–2083 (2006)
Sheridan, B. S. et al. Oral infection drives a distinct population of intestinal resident memory CD8+ T cells with enhanced protective function. Immunity 40, 747–757 (2014)
Masopust, D. et al. Dynamic T cell migration program provides resident memory within intestinal epithelium. J. Exp. Med. 207, 553–564 (2010)
Boddupalli, C. S. et al. ABC transporters and NR4A1 identify a quiescent subset of tissue-resident memory T cells. J. Clin. Invest. 126, 3905–3916 (2016)
Mackay, L. K. et al. T-box transcription factors combine with the cytokines TGF-β and IL-15 to control tissue-resident memory T cell fate. Immunity 43, 1101–1111 (2015)
Laidlaw, B. J. et al. CD4+ T cell help guides formation of CD103+ lung-resident memory CD8+ T cells during influenza viral infection. Immunity 41, 633–645 (2014)
Yu, B. et al. Epigenetic landscapes reveal transcription factors that regulate CD8+ T cell differentiation. Nat. Immunol. 18, 573–582 (2017)
Page, L., Brin, S., Motwani, R. & Winograd, T. The PageRank citation ranking: Bringing order to the web. Technical Report, Stanford InfoLab (1998)
Chen, R. et al. In vivo RNA interference screens identify regulators of antiviral CD4+ and CD8+ T cell differentiation. Immunity 41, 325–338 (2014)
Skon, C. N. et al. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat. Immunol. 14, 1285–1293 (2013)
Taniuchi, I. et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111, 621–633 (2002)
Cruz-Guilloty, F. et al. Runx3 and T-box proteins cooperate to establish the transcriptional program of effector CTLs. J. Exp. Med. 206, 51–59 (2009)
Shan, Q. et al. The transcription factor Runx3 guards cytotoxic CD8+ effector T cells against deviation towards follicular helper T cell lineage. Nat. Immunol. 18, 931–939 (2017)
Reis, B. S., Rogoz, A., Costa-Pinto, F. A., Taniuchi, I. & Mucida, D. Mutual expression of the transcription factors Runx3 and ThPOK regulates intestinal CD4+ T cell immunity. Nat. Immunol. 14, 271–280 (2013)
Grueter, B. et al. Runx3 regulates integrin α E/CD103 and CD4 expression during development of CD4−/CD8+ T cells. J. Immunol. 175, 1694–1705 (2005)
Lotem, J. et al. Runx3-mediated transcriptional program in cytotoxic lymphocytes. PLoS One 8, e80467 (2013)
Reis, B. S., Hoytema van Konijnenburg, D. P., Grivennikov, S. I. & Mucida, D. Transcription factor T-bet regulates intraepithelial lymphocyte functional maturation. Immunity 41, 244–256 (2014)
Djenidi, F. et al. CD8+CD103+ tumor-infiltrating lymphocytes are tumor-specific tissue-resident memory T cells and a prognostic factor for survival in lung cancer patients. J. Immunol. 194, 3475–3486 (2015)
Ganesan, A.-P. et al. Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat. Immunol. 18, 940–950 (2017)
Doedens, A. L. et al. Molecular programming of tumor-infiltrating CD8+ T cells and IL15 resistance. Cancer Immunol. Res. 4, 799–811 (2016)
Gooden, M. J., de Bock, G. H., Leffers, N., Daemen, T. & Nijman, H. W. The prognostic influence of tumour-infiltrating lymphocytes in cancer: a systematic review with meta-analysis. Br. J. Cancer 105, 93–103 (2011)
Singer, M . et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511.e9 (2016)
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016)
Steinert, E. M. et al. Quantifying memory CD8 T cells reveals regionalization of immunosurveillance. Cell 161, 737–749 (2015)
Benck, C. J., Martinov, T., Fife, B. T. & Chatterjea, D. Isolation of infiltrating leukocytes from mouse skin using enzymatic digest and gradient separation. J. Vis. Exp. 107, e53638 (2016)
Knott, S. R. V. et al. A computational algorithm to predict shRNA potency. Mol. Cell 56, 796–807 (2014)
Pear, W. S. et al. Efficient and rapid induction of a chronic myelogenous leukemia-like myeloproliferative disease in mice receiving P210 bcr/abl-transduced bone marrow. Blood 92, 3780–3792 (1998)
Best, J. A. et al. Transcriptional insights into the CD8+ T cell response to infection and memory T cell formation. Nat. Immunol. 14, 404–412 (2013)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Acknowledgements
We thank all the members of the Goldrath and Pipkin laboratories for their contributions. We also thank the Flow Cytometry Core at the La Jolla Institute for Allergy and Immunology. This study was funded in part by the UCSD Molecular Biology Cancer Fellowship (J.J.M.), the US National Institutes of Health U19AI109976 (S.C., M.E.P., A.W.G) and R01 AI095634 (M.E.P.), California Institute for Regenerative Medicine RB5-07012 (W.W.), the Kimmelman Family Foundation and the San Diego Center for Precision Immunotherapy (A.W.G.).
Author information
Authors and Affiliations
Contributions
J.J.M. designed and performed experiments, analysed the data and wrote the manuscript; C.T. assisted with the RNAi screen, transfections, transductions, tissue processing, and tumour models; B.Y. performed the computational analyses and ATAC–seq experiment; K.Z. assisted with computational analyses; K.O. and T.N. assisted with tissue processing, analysis, and qPCR; A.T.P assisted with tissue processing, inducible deletion experiments, and analysis; D.W. and A.J.G. helped with the inducible deletion experiments, RNA-seq analysis, and tumour models; S.C. provided reagents and advice; W.W. supervised the computational analysis and contributed advice; M.E.P. and A.W.G. supervised the project, designed experiments, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks F. Carbone, D. Mucida, A. Schietinger and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 KLRG1lo cells preferentially give rise to TRM cells.
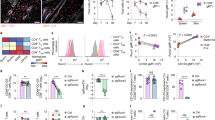
a, Left, representative flow cytometric gating strategy for distinguishing P14 cells located in non-lymphoid tissues after intravenous administration of CD8α in LCMV-infected mice. Right, in vitro activated P14 cells were transferred to recipient mice and infected with LCMV; the frequency of CD69+ and CD103+ P14 cells among KLRG1hi and KLRG1lo on day 7 of infection is indicated. b, Frequency of CCR9, CXCR3 and CD49d in KLRG1lo and KLRG1hi cells in the IEL compartment on day 7 of infection. c, Top, schematic of experimental design. KLRG1lo and KLRG1hi P14 cells were sorted from spleens and lymph nodes on day 5 of LCMV infection and transferred into recipient mice infected 4 days previously with LCMV. Bottom, TCM, TEM and TRM P14 cells were enumerated on days 20 or 25 of infection using flow cytometry. Data are mean ± s.e.m of n = 5 mice (a, b) or n = 3–4 mice (c) from one representative of 2 independent experiments. *P < 0.05, **P < 0.01, ***P < 0.005 (Student’s t-test). Symbols represent an individual mouse (c).
Extended Data Figure 2 Representative ATAC–seq peaks and putative TRM cell regulators identified through PageRank analysis.
a, ATAC–seq analysis of the indicated loci on day 7 of infection (left) and corresponding gene expression (right). b, Personalized PageRank score and gene expression of transcription factors, with selected factors highlighted.
Extended Data Figure 3 Runx3-deficiency impairs IEL TRM cell formation.
a, Runx3 mRNA levels from indicated cells determined by microarray analyses. b, Relative Runx3 mRNA expression of in vitro cultured cells transduced with retroviruses encoding control shRNAmir or Runx3 shRNAmir, measured by qPCR. c, Congenically distinct P14 cells were transduced with retroviruses encoding Runx3 shRNAmir or control shRNAmir, mixed at a 1:1 ratio, and transferred to recipient mice that were subsequently infected with LCMV. Representative flow cytometry plots (bottom, left) and quantification of the ratio of P14 cells transduced with Runx3 shRNAmir or control shRNAmir in indicated tissues on day 12 of infection (bottom, right). d, Representative flow cytometry plots (left) and quantification of the frequency of CD69+ and CD103+ cells of control shRNAmir or Runx3 shRNAmir cells (right) from experimental schematic in c. e, Representative flow cytometry plots and quantification of the frequency of CD69+ and CD103+ cells from Fig. 2c. f, Representative flow cytometry plots and quantification of the frequency of CD69+ and CD103+ cells from Fig. 2e. Data are mean ± s.e.m and representative of two independent experiments (b) with n = 5 (c, d), n = 5 (LM–GP33–41) or n = 6 (LCMV) (e), and n = 5 (vehicle) or n = 3 (tamoxifen) (f). *P < 0.05, **P < 0.01 ***P < 0.005 (Student’s t-test). Symbols represent an individual mouse (c–f).
Extended Data Figure 4 Runx3-deficiency impairs IEL TRM cell formation in a polyclonal setting.
a, Representative flow cytometry plot of H-2Db GP33–41 tetramer staining of lymphocytes from Runx3fl/fl dLck-Cre+YFP and Runx3+/+dLck-Cre+YFP mice on day 12 of LCMV infection (gated on total lymphocytes). b, Quantification of the proportion (left) and absolute number (right) of tetramer+ cells. c, d, Representative flow cytometry plots and quantification of the frequency of CD69+ and CD103+ cells. Data are mean ± s.e.m with n = 4 (Runx3+/+) or n = 5 (Runx3fl/fl) mice pooled from two independent experiments. **P < 0.01, ***P < 0.005 (Student’s t-test). Symbols represent an individual mouse (b, d).
Extended Data Figure 5 Runx3 is required for TRM cell formation in diverse non-lymphoid tissues.
a, Schematic of experimental design. b, Representative flow cytometry plots (left) and quantification (right) of the ratio of Runx3fl/fl to Runx3+/+ P14 cells (gated on YFP-Cre+ cells) in lymphoid and non-lymphoid compartments on days 15/16 of LCMV infection (as in Fig. 2d but including salivary gland (SG) and kidney populations). c, Schematic for experimental design. d, Representative flow cytometry plots (left) and quantification (right) of the ratio of transduced cells in the skin relative to the spleen for control shRNAmir or Runx3 shRNAmir P14 cells on day 12 of an intradermal LCMV infection. e, Frequency of CD69+ and CD103+ cells. f, Schematic for experimental design. g, Representative flow cytometry plots (left) and quantification (right) of the ratio of transduced cells in the lung parenchyma relative to the spleen for control shRNAmir or Runx3 shRNAmir P14 cells on day 12 of an intratracheal LCMV infection. h, Frequency of CD69+ and CD103+ cells. Data are mean ± s.e.m and representative of two independent experiments with n = 6 (b), or data pooled from two individual experiments with n = 6 per group (c–h). *P < 0.05, **P < 0.01, ***P < 0.005 (Student’s t-test). Symbols represent an individual mouse (b, d, e, g, h).
Extended Data Figure 6 Runx3-deficiency enhances TRM cell apoptosis but does affect trafficking or proliferation.
a, Representative flow cytometry histogram of granzyme B (GzB) staining (left) and quantification of frequency of GzB+ cells on day 12 or 14 of infection. b, Representative flow cytometry plots (left) and quantification (right) of the frequency of IFNγ- and TNF-producing control shRNAmir or Runx3 shRNAmir P14 cells on day 6 of LCMV infection, restimulated with GP33–41 peptide. c, d, Representative histograms and quantification of annexin V+ cells from shRNAmir mixed transfers on day 14 of LCMV infection (c) or from day 8 Runx3fl/fl and Runx3+/+ mixed P14 transfers in which tamoxifen was administered on days 2–5 of LCMV infection (d). e, Congenically distinct P14 cells were transduced with control shRNAmir or Runx3 shRNAmir encoding retroviruses, mixed at a 1:1 ratio, and transferred to recipient mice that were subsequently infected with LCMV. On day 6 of infection, splenocytes were collected and retransferred to day 5 infected host mice and 18 h later spleen, mesenteric lymph nodes and small intestine were obtained to assess trafficking. Representative flow cytometry plots (bottom, left) and quantification of the ratio of P14 cells transduced with control shRNAmir to Runx3 shRNAmir (bottom, right) in indicated tissues 18 h after transfer. f, Frequency of Ki-67+ control shRNAmir or Runx3 shRNAmir transduced P14 cells in a mixed transfer setting on days 6 and 12 or 14 of LCMV infection. Data are mean ± s.e.m and representative of two independent experiments with n = 5 (a), n = 3 (b), n = 5 (c), n = 6 (d), n = 4 (e), and n = 3 on day 6 or n = 4 on day 14 (f) except d is pooled from two independent experiments. *P < 0.05, **P < 0.01, ***P < 0.005 (Student’s t-test). n.s., not significant. Symbols represent an individual mouse (a–f).
Extended Data Figure 7 Runx3 overexpression enhances lung TRM differentiation.
a, Runx3 mRNA expression of in vitro cultured cells transduced with GFP-RV or Runx3-RV. b, Schematic for experimental design of intratracheal LCMV infection. c, Representative flow cytometry plots (left) and quantification (right) of the ratio of GFP-RV or Runx3-RV cells in the mediastinal lymph nodes (medLN), lung parenchyma, or CD69+CD103+ lung parenchyma population on day 12 or 13. d, Representative flow cytometry plots (left) and quantification (right) of the frequency of CD69+ and CD103+ P14 cells in the lung parenchyma. Data are mean ± s.e.m and representative of one of two independent experiments (a) and n = 4 per group (c,d). *P < 0.05, ***P < 0.005 (Student’s t-test). Symbols represent an individual mouse (c, d).
Extended Data Figure 8 Runx3 regulates distinct gene programs in circulating cells versus tissue resident cells and operates upstream of T-bet in programming IEL TRM cell differentiation.
a, Percentage of genes of the core tissue-residency signature, core circulating signature, or background sites that exhibit direct Runx3 binding by ChIP–seq analysis23. b, Left, predicted Runx3 binding network, generated from ATAC–seq analysis, in IEL P14 cells and splenic P14 cells on day 7 of infection. Red indicates genes putatively regulated by Runx3 in IEL cells; grey indicates genes putatively regulated by Runx3 in splenic cells. Right, Gene Ontology (GO) enrichment analysis of gene sets in the predicted Runx3 binding network in each tissue. c, Runx3 ChIP–seq of the Tbx21 locus in naive and activated CD8+ T cells from ref. 23. d, Representative flow cytometry histograms (left) and mean fluorescent intensity (MFI) quantification (right) of T-bet expression in splenic P14 cells on day 8 of infection. e, Schematic for experimental design (left) in which Runx3+/+ Ert2-Cre+ YFP were transduced with control shRNAmir and Runx3+/+Ert2-Cre+ YFP P14 cells were transduced with Tbx21 shRNAmir, mixed 1:1 and transferred into recipient mice subsequently infected with LCMV. Recipient mice were treated with tamoxifen on days 0–4 of infection. Representative flow cytometry plots (middle) and quantification of the ratio of untransduced (ametrine−) Runx3+/+ and Runx3fl/fl P14 cells and the ratio of transduced (ametrine+) Runx3+/+ control shRNAmir to Runx3fl/fl Tbx21 shRNAmir (right) were evaluated on day 12 of LCMV infection. f, Representative flow cytometry plots (left) and quantification (right) of the frequency of CD69+ and CD103+ cells. g, Runx3 ChIP–seq of the Klf2 locus in naive and activated CD8+ T cells23. h, Fold change in gene expression of Klf2, S1pr1 and Ccr7 in Runx3fl/fl and Runx3-RV cells relative to Runx3+/+ wild-type cells, from RNA-seq analysis consisting of two replicates per sample. Data are mean ± s.e.m and representative of one of two independent experiments with n = 6 (Runx3fl/fl) or n = 4 (Runx3 shRNAmir) (d) and n = 4 (e, f) per group. *P < 0.05, **P < 0.01 ***P < 0.005 (Student’s t-test). Symbols represent an individual mouse (d–f).
Extended Data Figure 9 Runx3-deficiency does not impair trafficking to the tumour but does affect the effector phenotype of TIL.
a, Schematic of adoptive therapy experimental design. b, Congenically distinct P14 cells were transduced with retroviruses encoding Runx3 shRNAmir or control shRNAmir, mixed at a 1:1 ratio, and transferred into mice with established B16-GP33 melanoma tumours. Eighteen hours after transfer, tumours were collected to assess the ratio of Runx3 shRNAmir to control shRNAmir P14 cells. c, Representative flow cytometry histograms of control shRNAmir, Runx3 shRNAmir, GFP-RV, or Runx3-RV TILs in mixed transfer settings. Control P14 splenocytes were included in histograms for reference. d, Gene set enrichment analysis of the core tissue-residency and core circulating gene signatures in human lung CD8+ TILs relative to corresponding CD8+ PBMCs25. Data are mean ± s.e.m and combined of two independent experiments with n = 5 mice per group (b) or representative of two independent experiments with n = 3–6 per group (b). Symbols represent an individual mouse (b).
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3. (PDF 351 kb)
Supplementary Data
This file contains tumour source data. (XLSX 56 kb)
Rights and permissions
About this article
Cite this article
Milner, J., Toma, C., Yu, B. et al. Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumours. Nature 552, 253–257 (2017). https://doi.org/10.1038/nature24993
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24993
This article is cited by
-
Tissue-resident memory T cells: decoding intra-organ diversity with a gut perspective
Inflammation and Regeneration (2024)
-
Current challenges and therapeutic advances of CAR-T cell therapy for solid tumors
Cancer Cell International (2024)
-
RUNX3 pathway signature predicts clinical benefits of immune checkpoint inhibition plus tyrosine kinase inhibition in advanced renal cell carcinoma
BMC Urology (2024)
-
The transcriptional cofactor Tle3 reciprocally controls effector and central memory CD8+ T cell fates
Nature Immunology (2024)
-
Multimodal profiling reveals site-specific adaptation and tissue residency hallmarks of γδ T cells across organs in mice
Nature Immunology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.