Abstract
The foundations of mammalian development lie in a cluster of embryonic epiblast stem cells. In response to extracellular matrix signalling, these cells undergo epithelialization and create an apical surface in contact with a cavity1,2, a fundamental event for all subsequent development. Concomitantly, epiblast cells transit through distinct pluripotent states3,4, before lineage commitment at gastrulation. These pluripotent states have been characterized at the molecular level5, but their biological importance remains unclear. Here we show that exit from an unrestricted naive pluripotent state is required for epiblast epithelialization and generation of the pro-amniotic cavity in mouse embryos. Embryonic stem cells locked in the naive state are able to initiate polarization but fail to undergo lumenogenesis. Mechanistically, exit from naive pluripotency activates an Oct4-governed transcriptional program that results in expression of glycosylated sialomucin proteins and the vesicle tethering and fusion events of lumenogenesis. Similarly, exit of epiblasts from naive pluripotency in cultured human post-implantation embryos triggers amniotic cavity formation and developmental progression. Our results add tissue-level architecture as a new criterion for the characterization of different pluripotent states, and show the relevance of transitions between these states during development of the mammalian embryo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Change history
28 February 2018
Please see accompanying Erratum (http://doi.org/10.1038/nature25995). Extended Data Fig. 4 has been replaced, to correct the missing colours in the key to panels c, h, k, n and q, and to correct the missing colours of the graph in panel k.
References
Bedzhov, I. & Zernicka-Goetz, M. Self-organizing properties of mouse pluripotent cells initiate morphogenesis upon implantation. Cell 156, 1032–1044 (2014)
Hertig, A. T., Rock, J. & Adams, E. C. A description of 34 human ova within the first 17 days of development. Am. J. Anat. 98, 435–493 (1956)
De Los Angeles, A. et al. Hallmarks of pluripotency. Nature 525, 469–478 (2015)
Kalkan, T. & Smith, A. Mapping the route from naive pluripotency to lineage specification. Phil. Trans. R. Soc. Lond. B 369, 20130540 (2014)
Weinberger, L., Ayyash, M., Novershtern, N. & Hanna, J. H. Dynamic stem cell states: naive to primed pluripotency in rodents and humans. Nat. Rev. Mol. Cell Biol. 17, 155–169 (2016)
Acampora, D., Di Giovannantonio, L. G. & Simeone, A. Otx2 is an intrinsic determinant of the embryonic stem cell state and is required for transition to a stable epiblast stem cell condition. Development 140, 43–55 (2013)
Nielsen, J. S. & McNagny, K. M. The role of podocalyxin in health and disease. J. Am. Soc. Nephrol. 20, 1669–1676 (2009)
Boroviak, T. et al. Lineage-specific profiling delineates the emergence and progression of naive pluripotency in mammalian embryogenesis. Dev. Cell 35, 366–382 (2015)
Bedzhov, I., Leung, C. Y., Bialecka, M. & Zernicka-Goetz, M. In vitro culture of mouse blastocysts beyond the implantation stages. Nat. Protoc. 9, 2732–2739 (2014)
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008)
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016)
Taniguchi, K. et al. Lumen formation is an intrinsic property of isolated human pluripotent stem cells. Stem Cell Reports 5, 954–962 (2015)
Betschinger, J. et al. Exit from pluripotency is gated by intracellular redistribution of the bHLH transcription factor Tfe3. Cell 153, 335–347 (2013)
Wang, Y., Medvid, R., Melton, C., Jaenisch, R. & Blelloch, R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat. Genet. 39, 380–385 (2007)
Dutta, D. et al. Self-renewal versus lineage commitment of embryonic stem cells: protein kinase C signaling shifts the balance. Stem Cells 29, 618–628 (2011)
Dunn, S. J., Martello, G., Yordanov, B., Emmott, S. & Smith, A. G. Defining an essential transcription factor program for naïve pluripotency. Science 344, 1156–1160 (2014)
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003)
Buecker, C. et al. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 14, 838–853 (2014)
Yang, S. H. et al. Otx2 and Oct4 drive early enhancer activation during embryonic stem cell transition from naive pluripotency. Cell Rep. 7, 1968–1981 (2014)
Bryant, D. M. et al. A molecular network for de novo generation of the apical surface and lumen. Nat. Cell Biol. 12, 1035–1045 (2010)
Ullrich, O., Reinsch, S., Urbé, S., Zerial, M. & Parton, R. G. Rab11 regulates recycling through the pericentriolar recycling endosome. J. Cell Biol. 135, 913–924 (1996)
Meder, D., Shevchenko, A., Simons, K. & Füllekrug, J. Gp135/podocalyxin and NHERF-2 participate in the formation of a preapical domain during polarization of MDCK cells. J. Cell Biol. 168, 303–313 (2005)
Doyonnas, R. et al. Anuria, omphalocele, and perinatal lethality in mice lacking the CD34-related protein podocalyxin. J. Exp. Med. 194, 13–28 (2001)
Strilic´, B. et al. Electrostatic cell-surface repulsion initiates lumen formation in developing blood vessels. Curr. Biol. 20, 2003–2009 (2010)
Blasky, A. J., Mangan, A. & Prekeris, R. Polarized protein transport and lumen formation during epithelial tissue morphogenesis. Annu. Rev. Cell Dev. Biol. 31, 575–591 (2015)
Mangan, A. J. et al. Cingulin and actin mediate midbody-dependent apical lumen formation during polarization of epithelial cells. Nat. Commun. 7, 12426 (2016)
Blakeley, P. et al. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 142, 3151–3165 (2015)
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016)
Theunissen, T. W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014)
Collier, A. J. et al. Comprehensive Cell surface protein profiling identifies specific markers of human naive and primed pluripotent states. Cell Stem Cell 20, 874–890 (2017)
Takashima, Y. et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 158, 1254–1269 (2014)
Vuoristo, S., Jedrusik, A., Shahbazi M. & Zernicka-Goetz, M. Culture of human embryos through implantation stages in vitro. Protoc. Exch. https://doi.org/10.1038/protex.2016.017 (2016)
Rhee, J. M. et al. In vivo imaging and differential localization of lipid-modified GFP-variant fusions in embryonic stem cells and mice. Genesis 44, 202–218 (2006)
Yeom, Y. I. et al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development 122, 881–894 (1996)
Kalkan, T. et al. Tracking the embryonic stem cell transition from ground state pluripotency. Development 144, 1221–1234 (2017)
Panamarova, M. et al. The BAF chromatin remodelling complex is an epigenetic regulator of lineage specification in the early mouse embryo. Development 143, 1271–1283 (2016)
Lee, G. Y., Kenny, P. A., Lee, E. H. & Bissell, M. J. Three-dimensional culture models of normal and malignant breast epithelial cells. Nat. Methods 4, 359–365 (2007)
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012)
Martín-Belmonte, F. et al. Cell-polarity dynamics controls the mechanism of lumen formation in epithelial morphogenesis. Curr. Biol. 18, 507–513 (2008)
Yang, Z. et al. De novo lumen formation and elongation in the developing nephron: a central role for afadin in apical polarity. Development 140, 1774–1784 (2013)
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Acknowledgements
We thank K. McNagny, J. Hanna and R. Jaenisch for reagents and discussions; F. Martin-Belmonte, D. Glover, C. Lynch, M. Serrano, A. Hupalowska, F. Antonica and M. Petruzzelli for feedback; J. N. Skepper for help with electron microscopy; W. Mansfield for help with embryo transfer. This work was supported by Wellcome Trust (098287/Z/12/Z) and ERC (669198) grants to M.Z.-G. Work in the laboratory of T.V. was supported by Wellcome Trust and KU Leuven (SymBioSys PFV/10/016). Work in the laboratory of J.C.M. was supported by EMBL and Cancer Research UK. M.N.S. was supported by Ramon Areces and EMBO postdoctoral fellowships; A.S. by a Wellcome Trust strategic award (105031/D/14/Z) and G.R. by a Newton fellowship.
Author information
Authors and Affiliations
Contributions
M.N.S. designed, performed and analysed most of the experiments. A.S. analysed the sequencing data. N.S. and A.W. performed experiments in Fig. 4g and Extended Data Figs 5–7. M.Z. and G.R. helped with embryo experiments and image analysis. A.J., L.G.D. and L.N. helped with human embryo cultures. I.C.M. prepared cDNA libraries. C.B. generated and analysed ChIP–seq data. D.I. and Y.K. supervised the human embryo experiments. T.V. supervised the cDNA library preparation. J.C.M. supervised the computational analyses of the sequencing data. M.Z.-G. supervised the study. M.N.S. and M.Z.-G. conceived the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J.-L. Maitre and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Epiblast gene expression patterns at peri-implantation stages.
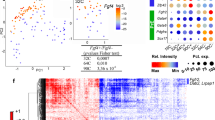
a, CAG–GFP+ embryos recovered at peri-implantation stages, before and after dissection of the epiblast. n = 4 E4.5, 4 E4.75 and 4 E5.0 embryos. b–e, For all the collected samples we computed the total number of reads (b), the fraction of reads mapped to endogenous genes (c), the number of genes with more than 10 reads per million (RPM) (d) and the fraction of reads mapped to mitochondrial genes (e). f, A PCA of the metrics shown in b–e. The sample coloured in grey (E4.5_S1) is characterized by a very low fraction of mapped reads and number of genes detected and was therefore removed from all downstream analysis. g, Hierarchical clustering of the E4.5–E4.75 samples that passed the quality check (black) and samples from the epiblast (red) and primitive endoderm (blue) at E4.5 from the dataset of ref. 8. One sample (E4.5_S3, grey) clusters with the primitive endoderm, whereas all others cluster with the epiblast. E4.5_S3 was therefore excluded from further analysis. h, PCA of our samples and samples from ref. 8. i, log fold change in pluripotency marker genes between E5.0 and E4.5–E4.75 samples from our dataset (x axis) and log fold change between E5.5 and E4.5 epiblast samples from ref. 8 (y axis). Squares, circles and triangles mark genes differentially expressed in both datasets, only in our dataset or only in the dataset of ref. 8 respectively. The naive pluripotency genes significantly downregulated at E5.5 were highly enriched among genes downregulated at E5.0 (P = 7 × 10−8, Fisher’s exact test), whereas the enrichment of upregulated early post-implantation factors was only marginally significant (P = 0.03, Fisher’s exact test). j, Bright field images of cells derived from embryos cultured in IVC1 (0/6 mES cell colonies from IVC1 embryos and 4/6 mES cell colonies from IVC1 +2i/LIF embryos). Scale bars, 50 μm. k, Immunostaining of mES cells derived from embryos cultured in IVC1 +2i/LIF for 24 h. Scale bar, 10 μm. l, Experimental set-up. A, analysis. m, Immunostaining of mouse embryos cultured as indicated in l. Dotted lines indicate the epiblast and the arrow points to the pro-amniotic cavity. Scale bars, 20 μm. n, Lumen formation in embryos from m. n = 12 IVC1 control and 15 IVC1 experimental embryos. χ2 test, *P = 0.0137.
Extended Data Figure 2 Dynamics of polarization, lumenogenesis and naive pluripotency exit in mES cells.
a, Experimental set-up. A, analysis. b, c, Immunostaining of mES cells in 3D Matrigel (b, top, c). Red long arrows (b) indicate the position used to plot intensity profiles (b, bottom). Yellow arrows indicate polarization and white arrows lumen formation. Scale bars, 10 μm. d, Quantification of lumen formation in cells from b and c. n = 21 (2 cells), 22 (4 cells), 20 (8 cells) and 21 (>10 cells) spheroids. χ2 test, ***P < 0.0001. e, Fluorescence intensity in cells from f. Raw fluorescence intensity values were normalized to the fluorescence intensity of reporter cells cultured in gelatin +2i/LIF throughout the time-lapse video. n = 55 Nanog–YFP, 92 ΔPE–Oct4–GFP and 63 Rex1::GFPd2 spheroids. AU, arbitrary units. f, Time-lapse frames of naive reporter mES cells cultured in Matrigel without 2i/LIF. mT: membrane Tomato. Time is indicated as h:min. Dotted lines indicate the outline of the spheroids at the end of the experiment. Scale bars, 20 μm. g–o, Expression of pluripotency genes in mES cells cultured in gelatin or Matrigel, with or without 2i/LIF. n = 5 (24 h), 3 (36 h), 3 (48 h) independent samples for Nanog, Esrrb and Otx2; n = 5 (24 h), 3 (36 h), 4 (48 h) independent samples for Fgf5; n = 4 (24 h), 3 (36 h), 3 (48 h) independent samples for Klf4 and Klf2; n = 4 (24 h), 3 (36 h), 4 (48 h) independent samples for Zfp42; and n = 3 (24 h), 3 (36 h), 3 (48 h) independent samples for Tbx3 and Pou5f1. Unpaired Student’s t-test, NS, not significant. ND, not detected. p, q, Immunostaining of mES cells cultured in 3D Matrigel. Scale bars, 10 μm. M, Matrigel.
Extended Data Figure 3 mES cells initiate polarization in naive conditions in a 3D Matrigel culture.
a, Experimental set-up. A, analysis. b, Immunostaining of mES cells cultured as indicated in a. Arrows indicate lumens. Scale bars, 5 μm. c, Lumen formation in cells from b. n = 31 (2 cells, gelatin +2i/LIF 48 h, Matrigel −2i/LIF 24 h), 30 (4 cells, gelatin +2i/LIF 48 h, Matrigel −2i/LIF 24 h), 31 (2 cells, gelatin −2i/LIF 48 h, Matrigel −2i/LIF 24 h) and 31 (4 cells, gelatin −2i/LIF 48 h, Matrigel −2i/LIF 24 h) spheroids. χ2 test, ***P < 0.0001. d, Internuclear distance in cells from Fig. 2b. n = 31 (+2i/LIF) and 30 (−2i/LIF) spheroids. Mann–Whitney U-test; NS, not significant. e–g, Immunostaining of mES cells cultured in Matrigel with or without 2i/LIF. Squares indicate magnified regions (e). Scale bars, 10 μm (e and f) and 5 μm (g). h–j, Representative frames from time-lapse experiments using different naive reporter mES cell lines, cultured in either gelatin or Matrigel in the presence of 2i/LIF. Time is indicated as h:min. Scale bars, 50 μm. k–m, Intensity of Nanog–YFP, ΔPE–Oct4–GFP and Rex1::GFPd2 mES cells analysed by time-lapse microscopy from h–j. AU, arbitrary units. n = 65 Nanog–YFP, 55 ΔPE–Oct4–GFP and 43 Rex1::GFPd2 mES cell colonies (gelatin); and n = 61 Nanog–YFP, 84 ΔPE–Oct4–GFP and 61 Rex1::GFPd2 mES cell spheroids (Matrigel). n, Experimental set-up. o, Immunostaining of mES cells cultured as indicated in n. Squares indicate magnified regions. Dotted lines demarcate the border of the colonies. The nucleus–centrosome angle (α) measured in p is indicated. Scale bars, 10 μm. p, Histogram of the angle between the centrosome–nucleus axis and the basal–apical axis of cells in the border of the colonies shown in o. Data are shown as relative frequencies (%). n = 76 (Matrigel +2i/LIF cultured in gelatin), 61 (Matrigel −2i/LIF cultured in gelatin), 62 (Matrigel +2i/LIF) and 60 (Matrigel −2i/LIF) centrosomes. Kruskal–Wallis test; ***P < 0.0001; NS, not significant. q, Immunostaining of control and Dgcr8 knockout mES cells (with or without 2i/LIF). Scale bars, 5 μm. r, Angle between the nucleus–centrosome axis and the nuclear–nuclear axis in cells from q. Each dot represents an individual centrosome. n = 60 control +2i/LIF, 62 control −2i/LIF, 56 Dgcr8 knockout +2i/LIF and 58 Dgcr8 knockout −2i/LIF centrosomes. Kruskal–Wallis test; NS, not significant. s, Internuclear distance in cells from q. n = 30 control +2i/LIF, 31 control −2i/LIF, 28 Dgcr8 knockout +2i/LIF and 29 Dgcr8 knockout −2i/LIF spheroids. Kruskal–Wallis test. t, Nanog intensity in cells from q. n = 30 control +2i/LIF, 31 control −2i/LIF, 28 Dgcr8 knockout +2i/LIF and 30 Dgcr8 knockout −2i/LIF spheroids. Kruskal–Wallis test; ***P < 0.0001. u, Immunostaining of mES cells cultured in Matrigel with or without Gö6983. Scale bars, 10 μm. v, Internuclear distance in cells from Fig. 2i. n = 26 (+Gö6983) and 28 (−Gö6983) spheroids. Mann–Whitney U-test; NS, not significant. G, gelatin; M, Matrigel.
Extended Data Figure 4 2i/LIF inhibits lumen formation in mES cells.
a, Immunostaining of mES cells cultured in Matrigel. Scale bars, 10 μm. b, Immunostaining of Rex1::GFPd2 mES cells cultured in Matrigel. Scale bars, 10 μm. c, Oct4, Nanog and Rex1::GFPd2 fluorescence intensity in cells from a and b. n = 30 spheroids per condition (Nanog); n = 21 (+2i/LIF) and 20 (−2i/LIF) spheroids (Oct4); and 21 (+2i/LIF) and 20 (−2i/LIF) spheroids (Rex1::GFPd2). Unpaired Student’s t-test; ***P < 0.0001; NS, not significant. d, e, Electron microscopy images of mES cells cultured in Matrigel with or without 2i/LIF. Arrows indicate lumens. a, Golgi apparatus; b, basolateral cell–cell adhesion sites; c, tight junctions. Scale bars, 2 μm (d) and 500 nm (e). f, g, Immunostaining of mES cells cultured in Matrigel. Squares indicate magnified regions (g). Arrows indicate inner non-polarized cells. Scale bars, 10 μm. h, Polarization in cells from g and Fig. 3c. n = 27 (+2i/LIF 48 h), 26 (−2i/LIF 48 h), 24 (+2i/LIF 72 h) and 24 (−2i/LIF 72 h) spheroids. χ2 test; ***P < 0.0001. i, Experimental set-up. A, analysis. j, Immunostaining of mES cells cultured as indicated in i. Arrow points to lumen. Scale bars, 10 μm. k, Lumen formation in cells from j. n = 20 spheroids per condition. χ2 test; NS, not significant. l, Nanog intensity in cells from j. n = 20 spheroids per condition. ANOVA; ***P < 0.0001. m, Immunostaining of mES cells cultured in Matrigel with different combinations of inhibitors. Arrow points to lumen. Scale bars, 10 μm. n, Lumen formation in cells from m. n = 20 spheroids per condition. χ2 test; ***P < 0.0001. o, Nanog intensity in cells from m. n = 20 spheroids per condition. ANOVA; ***P < 0.0001. p, Immunostaining of mES cells cultured in Matrigel with a single inhibitor or supplement. 2i/LIF was replaced by a single inhibitor or supplement 48 h before plating the cells in Matrigel. Arrows indicate lumens. Scale bars, 10 μm. q, Nanog intensity in cells from p as a function of their ability to undergo lumenogenesis. n = 30 (+2i/LIF control), n = 50 (+LIF), n = 48 (+GSK3i), n = 51 (+MEKi) and n = 39 (−2i/LIF) spheroids. Mann–Whitney U-test; **P = 0.0055; ***P < 0.0001; NS, not significant. M, Matrigel.
Extended Data Figure 5 Exit from naive pluripotency is required for lumenogenesis in mES cells.
a, Immunostaining of control and Dgcr8 knockout mES cells (with or without 2i/LIF). b, Lumen formation in cells from a. n = 20 (control +2i/LIF), 20 (control −2i/LIF), 20 (Dgcr8 knockout +2i/LIF) and 27 (Dgcr8 knockout −2i/LIF) spheroids. χ2 test; *P = 0.024. c, Nanog intensity in cells from a. n = 20 (control +2i/LIF), 20 (control −2i/LIF), 20 (Dgcr8 knockout +2i/LIF) and 27 (Dgcr8 knockout −2i/LIF) spheroids. ANOVA; ***P < 0.0001. d, Nanog intensity in Dgcr8 knockout mES cells after 72 h of 3D Matrigel culture (a), as a function of their ability to undergo lumenogenesis. n = 6 (lumen) and 21 (no lumen) spheroids. Unpaired Student’s t-test; *P = 0.0208. e, Immunostaining of mES cells cultured in Matrigel with or without Gö6983. f, Lumen formation in cells from e. n = 30 spheroids per condition. χ2 test; ***P < 0.0001. g, Immunostaining of Rex1::GFPd2 mES cells cultured in Matrigel with or without Gö6983. h, Nanog, Rex1::GFPd2 and Oct4 intensity in cells from e and g. n = 30 spheroids per condition (Nanog); 41 (+Gö6983) and 40 (−Gö6983) spheroids (Oct4 and Rex1::GFPd2). Mann–Whitney U-test; **P = 0.0002; ***P < 0.0001. i, Experimental set-up. A, analysis. j, Immunostaining of mES cells cultured as indicated in i. The percentage of structures showing the representative phenotype is indicated. k, Nanog intensity in cells from j as a function of their ability to undergo lumenogenesis. n = 13 (no lumen) and 20 (lumen) spheroids. Unpaired Student’s t-test; ***P < 0.0001. l, Immunostaining of mES cells cultured as indicated in i. The percentage of structures showing the representative phenotype is indicated. m, Nanog intensity in cells from l as a function of their ability to undergo lumenogenesis. n = 14 (no lumen) and 17 (lumen) spheroids. Unpaired Student’s t-test; ***P < 0.0001. n, Immunostaining of DOX-inducible Nanog mES cells. o, Lumen formation in cells from n. n = 30 (+2i/LIF/DOX), 30 (+2i/LIF), 31 (+DOX) and 28 (−2i/LIF) spheroids. χ2 test, NS, not significant. p, Nanog intensity in cells from n. n = 30 (+2i/LIF/DOX), 30 (+2i/LIF), 31 (+DOX) and 28 (−2i/LIF) spheroids. Kruskal–Wallis test; ***P < 0.0001. q, mRNA levels of Nanog and Otx2 in DOX-inducible Nanog mES cells. n = 5 (Nanog) and 3 (Otx2) independent samples per group. Unpaired Student’s t-test; **P = 0.0044; NS, not significant. r, mRNA levels of naive pluripotency genes in DOX-inducible Nanog mES cells. n = 4 independent samples per group. Unpaired Student’s t-test; **P = 0.0075; NS, not significant. s, Immunostaining of control and Nanog knockdown mES cells. t, Lumen formation in cells from s. n = 20 (control siRNA) and 42 (Nanog siRNA) spheroids per condition. χ2 test; NS, not significant. u, Nanog and Rex1::GFPd2 intensity in cells from s. n = 20 (control siRNA) and 42 (Nanog siRNA) spheroids. v, Correlation between Nanog and Rex1::GFPd2 levels in cells from s. n = 20 (control siRNA) and 42 (Nanog siRNA) spheroids. w, Otx2 and Oct4 intensity in Oct4 knockdown mES cells. n = 19 (lumen) and 28 (no lumen) spheroids. x, Immunostaining of control and Oct4 knockdown mES cells. Scale bars, 10 μm (a, e, g, l, n, s, x) and 5 μm (j); arrows indicate lumens.
Extended Data Figure 6 Sialomucins are required for mES cell lumenogenesis.
a, mRNA levels of Podxl in mES cells. n = 5 (24 h), 3 (36 h) and 4 (48 h) independent samples. Unpaired Student’s t-test; *P = 0.0395 (gelatin 24 h); *P = 0.0481 (Matrigel 24 h); **P = 0.0038 (gelatin 36 h); *P = 0.0126 (Matrigel 36 h); **P = 0.0075 (gelatin 48 h); *P = 0.0139 (Matrigel 48 h). b, Immunostaining of mES cells cultured in 3D Matrigel. Arrow indicates lumen. Scale bars, 10 μm. c, Immunostaining of control and Dgcr8 knockout mES cells cultured in Matrigel with or without 2i/LIF. Arrow indicates lumen. Scale bars, 10 μm. d, Immunostaining of mES cells cultured in Matrigel with or without Gö6983. Arrow indicates lumen. Scale bars, 10 μm. e, Immunostaining of control and Podxl knockdown mES cells. A binarized image of the Podxl channel was used to determine presence or absence of Podxl. The percentages indicate the proportion of spheroids with or without Podxl in the Podxl siRNA group. Arrows indicate lumens. Scale bars, 10 μm. f, Lumen formation in cells from e. n = 43 (control), 45 (Podxl siRNA with Podxl (Podxl siRNA-treated cells that did not show a reduction in Podxl expression)) and 35 (Podxl siRNA without Podxl (Podxl siRNA-treated cells that showed a reduction in Podxl expression)) spheroids. χ2 test; ***P < 0.0001. g, mRNA levels of early post-implantation factors in control and Podxl knockdown mES cells 24 h after removal of 2i/LIF. n = 3 independent samples per group. Student’s t-test; NS, not significant. h, mRNA levels of naive pluripotency genes in control and Podxl knockdown mES cells 24 h after removal of 2i/LIF. n = 3 independent samples per group. Student’s t-test; NS, not significant. i, Immunostaining of control and Podxl knockdown mES cells cultured in Matrigel. Arrows indicate polarization. Scale bars, 5 μm. j, Centrosome positions in cells from i. Each dot represents an individual centrosome. n = 40 (control siRNA) and 36 (Podxl siRNA) centrosomes. Mann–Whitney U-test; NS, not significant. k, Internuclear distance in cells from i. n = 20 (control siRNA) and 18 (Podxl siRNA) spheroids. Unpaired Student’s t-test; NS, not significant. l, Immunostaining of Podxl heterozygous (HET) and knockout mES cells. One knockout clone is shown as an example. Arrows indicate lumens. Scale bars, 10 μm. m, Lumen formation in cells from l. n = 30 (HET), 33 (clone 1) and 20 (clone 2) spheroids. χ2 test; NS, not significant. n, Immunostaining of control and Podxl knockdown mES cells with or without GFP–Cd34 overexpression. Arrows indicate lumens. Scale bars, 10 μm. o, Lumen formation in cells from n. n = 37 (control siRNA), 29 (Podxl siRNA), 33 (control siRNA +GFP–Cd34) and 31 (Podxl siRNA +GFP–Cd34) spheroids. χ2 test; ***P < 0.0001; NS, not significant. p, mRNA levels of sialomucin proteins in control and Podxl knockdown mES cells (with or without GFP–Cd34 overexpression) 48 h after removal of 2i/LIF. n = 4 (without GFP–Podxl) and 6 (with GFP–Cd34) independent samples per group. Unpaired Student’s t-test; *P = 0.0145; ***P = 0.0001. G, gelatin; M, Matrigel.
Extended Data Figure 7 Otx2 and Oct4 induce Podxl expression upon naive pluripotency exit.
a, mRNA levels of Fgf5 and Podxl in wild-type and Otx2 knockout mES cells at different time points after removal of 2i/LIF. n = 2 independent samples per group. ANOVA; **P = 0.0092; ***P < 0.0001. b, c, Immunostaining of wild-type and Otx2 knockout mES cells cultured in Matrigel. d, Lumen formation in cells from b and c. n = 46 (wild-type 48 h), 34 (wild-type 72 h), 36 (Otx2 knockout 48 h) and 35 (Otx2 knockout 72 h) spheroids. χ2 test; ***P < 0.0001. e, Immunostaining of Otx2 knockout and Otx2-knockout tetON-Otx2 mES cells in the presence or absence of DOX (DOX addition triggers Otx2 overexpression). f, Lumen formation in cells from e and in wild-type mES cells cultured without 2i/LIF and with or without Otx2 overexpression. n = 19 (wild-type), 20 (wild-type tetON-Otx2 −DOX), 20 (wild-type tetON-Otx2 +DOX), 20 (Otx2 knockout), 20 (Otx2 knockout tetON-Otx2 −DOX) and 23 (Otx2 knockout tetON-Otx2 +DOX) spheroids. χ2 test; ***P < 0.0001. g, Immunostaining of wild-type tetON-Otx2 mES cells overexpressing Otx2 (addition of DOX). h, Lumen formation in cells from g (and their corresponding controls) and in Otx2 knockout mES cells cultured with 2i/LIF and with or without Otx2 overexpression. n = 20 (wild-type), 20 (wild-type tetON-Otx2 −DOX), 50 (wild-type tetON-Otx2 +DOX), 19 (Otx2 knockout), 21 (Otx2 knockout tetON-Otx2 –DOX) and 20 (Otx2 knockout tetON-Otx2 +DOX) spheroids. χ2 test; *P = 0.0196; ***P < 0.0001. i, j, Nanog (i) and Otx2 (j) intensity in cells from g as a function of their ability to undergo lumogenesis. n = 25 spheroids per condition. Mann–Whitney U-test; ***P < 0.0001. k, mRNA levels of naive pluripotency genes in wild-type and Otx2 knockout mES cells at different time points after removal of 2i/LIF. n = 2 independent samples per group. ANOVA; *P = 0.02; **P = 0.0045; ***P = 0.0002 (Klf4); ***P = 0.0001 (Tbx3). l, Immunostaining of wild-type and Otx2 knockout mES cells overexpressing GFP–Podxl. m, Lumen formation in cells from l. n = 66 (wild-type) and 61 (Otx2 knockout) spheroids. χ2 test; NS, not significant. n, ChIP–seq analysis of Oct4, Otx2 and H3K27ac in mES cells (top) and after 48 h of transition into epiblast-like cells (EpiLCs, bottom). Shaded area indicates an enhancer within the first intron of Podxl that is activated during cell fate transition. o, Immunostaining of GFP–Podxl overexpressing mES cells. . p, Lumen formation in cells from o. n = 30 (+2i/LIF clone 1), 32 (−2i/LIF clone 1), 33 (+2i/LIF clone 2) and 34 (−2i/LIF clone 2) spheroids. χ2 test; ***P < 0.0001. Scale bars, 10 μm; arrows indicate lumens; (b, c, e, g, l, o). M, Matrigel.
Extended Data Figure 8 Cgn is induced upon naive pluripotency exit and mediates Rab11 vesicle tethering to the apical membrane.
a, mRNA levels of Cgn in mES cells. n = 5 (24 h), 3 (36 h) and 4 (48 h) independent samples per group. Unpaired Student’s t-test; *P = 0.0431 (gelatin 24 h); *P = 0.0347 (Matrigel 24 h); ***P = 0.0003 (gelatin 36 h), *P = 0.0126 (Matrigel 36 h); **P = 0.0075 (gelatin 48 h); *P = 0.0139 (Matrigel 48 h). b, c, Immunostaining of mES cells cultured in Matrigel with or without 2i/LIF. Squares indicate magnified regions. d, mRNA levels of naive pluripotency genes and early post-implantation factors in control and Cgn knockdown mES cells. n = 3 independent samples per group. Unpaired Student’s t-test; ***P < 0.0001; NS, not significant. e, Immunostaining of Rex1::GFPd2 mES cells in control and Cgn knockdown mES cells. Scale bars, 10 μm (b, c, e). Matrigel (M).
Extended Data Figure 9 Characterization of pluripotency gene expression and epiblast morphogenesis in post-implantation human embryos.
a, Immunostaining of day 9–10 human embryos. Dotted lines indicate the epiblast and the arrow indicates the amniotic cavity. Scale bars, 50 μm and 10 μm (magnified areas). b, NANOG intensity in embryos from a. n = 4 (IVC control with lumen), 3 (IVC control without lumen) and 6 (IVC +5i/LAF) human embryos. Mann–Whitney U-test; *P = 0.0381. c, Three-dimensional reconstruction of the epiblast (based on the KLF17 staining) and the amniotic cavity (based on the PODXL staining) of embryos from Fig. 5b. d, Immunostaining of day 9–10 human embryos. n = 2 embryos per group. Scale bars, 50 μm. e, Immunostaining of day 9–10 human embryos. Scale bars, 50 μm. f, Number of apoptotic cells per embryo in embryos from e. n = 3 (IVC control) and 5 (IVC +5i/LAF) embryos per condition. Mann–Whitney U-test; *P = 0.0179.
Extended Data Figure 10 Exit from naive pluripotency is required for lumenogenesis in hES cells.
a, Immunostaining of WIBR3 ΔPE–OCT4–GFP cells cultured in primed FBS/KSR/bFGF2 medium or N2B27 2i/hLIF with or without DOX. DOX addition induces the expression of NANOG and KLF2. Scale bars, 20 μm. b, c, mRNA levels of pluripotency genes in cells from a. n = 3 independent samples per condition. ANOVA; ***P < 0.0001 (NANOG and KLF2); ***P = 0.0001 (TFCP2L1); **P = 0.0013 (DNMT3L). d, Immunostaining of WIBR3 ΔPE–OCT4–GFP cells cultured in Matrigel in primed or naive conditions. Arrows indicate lumens. Scale bars, 20 μm. e, Lumen formation in cells from d. n = 38 (2i/hLIF +DOX), 33 (2i/hLIF −DOX), 34 (FBS/KSR/bFGF2) and 20 (mTESR) spheroids. χ2 test; ***P < 0.0001. f–h, mRNA levels of pluripotency genes and early post-implantation factors in cells from i. n = 4 (NANOG, KLF2, TFCP2L1, DNMT3L, KLF4 and PODXL) and 3 (KLF17) independent samples per group. Kruskal–Wallis test; ***P = 0.0002 (NANOG); ***P < 0.0001 (KLF2, DNMT3L, KLF4, PODXL); **P = 0.0058 (TFCP2L1); *P = 0.0205 (KLF17). i, Immunostaining of WIBR3 ΔPE–OCT4–GFP cells cultured in primed mTESR medium or naive conditions (2i/hLIF +DOX, 5i/LAF and RSet). Scale bars, 10 μm. j, Immunostaining of hES cells cultured in 3D Matrigel with different naive and primed conditions. Arrow indicates lumen. Scale bars, 10 μm. k, Lumen formation in cells from j. n = 31 (2i/hLIF +DOX), 27 (5i/LAF), 33 (RSet) and 20 (mTESR) spheroids. χ2 test; **P = 0.0019; ***P < 0.0001. l, Immunostaining of hES cells cultured in 3D Matrigel with different naive and primed conditions. Arrows indicate polarized centrosomes. Scale bars, 5 μm. m, Angle between the nucleus–centrosome axis and the nuclear–nuclear axis in cells from l. Each dot represents an individual centrosome. n = 53 (2i/LIF +DOX), 51 (5i/LAF), 40 (RSet) and 42 (mTESR) centrosomes. Kruskal–Wallis test; NS, not significant. n, Immunostaining of naive hES cells cultured in 3D Matrigel with mTESR. The initial naive conditions in which the cells were cultured are indicated. Arrows indicate lumens. Scale bars, 10 μm. o, Lumen formation in cells from n. n = 30 (2i/hLIF +DOX 48 h), 30 (2i/hLIF +DOX 72 h), 30 (5i/LAF 48 h), 29 (5i/LAF 72 h), 30 (RSet 48 h) and 30 (RSet 72 h). M, Matrigel.
Supplementary information
Supplementary Table 1
This file contains genes expressed in the mouse epiblast at peri-implantation stages. (XLS 6565 kb)
Supplementary Table 2
This file contains antibodies used in this study. (PDF 147 kb)
Supplementary Table 3
This file contains a list of RT-PCR primers used in this study. (PDF 153 kb)
ΔPE-Oct4-GFP mESCs cultured in 3D matrigel without 2i/LIF and imaged every 30 minutes.
The arrow points to the cell shown in Extended Data Fig. 2e. The maximum projection is shown throughout the movie. Scale bars, 50 μm. (AVI 11268 kb)
Rex1::GFPd2 mESCs cultured in 3D matrigel without 2i/LIF and imaged every 30 minutes.
The arrow points to the cell shown in Extended Data Fig. 2e. The maximum projection is shown throughout the movie. Scale bars, 50 μm. (AVI 15662 kb)
Nanog-YFP mTmG mESCs cultured in 3D matrigel without 2i/LIF and imaged every 30 minutes.
The arrow points to the cell shown in Extended Data Fig. 2e. The maximum projection is shown throughout the movie. Scale bars, 50 μm. (AVI 7893 kb)
Source data
Rights and permissions
About this article
Cite this article
Shahbazi, M., Scialdone, A., Skorupska, N. et al. Pluripotent state transitions coordinate morphogenesis in mouse and human embryos. Nature 552, 239–243 (2017). https://doi.org/10.1038/nature24675
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24675
This article is cited by
-
Modelling post-implantation human development to yolk sac blood emergence
Nature (2024)
-
Hypoblast from human pluripotent stem cells regulates epiblast development
Nature (2024)
-
Distinct pathways drive anterior hypoblast specification in the implanting human embryo
Nature Cell Biology (2024)
-
A 3D “sandwich” co-culture system with vascular niche supports mouse embryo development from E3.5 to E7.5 in vitro
Stem Cell Research & Therapy (2023)
-
Esrrb guides naive pluripotent cells through the formative transcriptional programme
Nature Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.